Chapter 12 - ER system and Protein Sorting (2/20)
Endomembrane System Components
Rough/Smooth ER
Golgi Apparatus
Lysosomes
Endosomes (helps with sorting within the endomembrane system)
Also includes…
Nuclear Envelope (which the ER is an extension of)
Plasma Membrane
Peroxisome
Transport vesicles
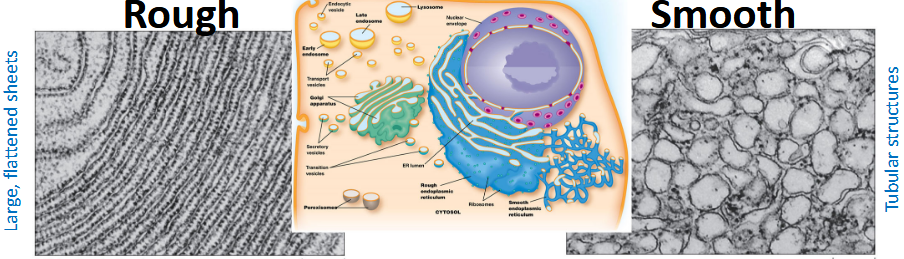
Endoplasmic Reticulum
An extensive network of flattened membrane sacs (cristae) and tubules, that extends outward throughout the cell.
Lumen = inside space of the ER
This is one of the largest membrane system inside the cell, makes up about 50 to 90% of the membrane mass in the cell
Rough ER
Characterized by flat membrane sheets with attached ribosomes that give it a “rough” appearance.
Functions related to the synthesis and processing of proteins. These proteins become…
Secreted from cell
Trans-membrane Proteins
Hydrolytic enzymes used by Lysosomes
Protein Synthesis Pathway
Signal Sequence targets the ribosomes to the ER
Protein enters ER lumen while being made (co-translational)
Protein can embed into membrane or stay in lumen
Other Functions of RER
Molecular chaperones to assist in protein folding and quality control
Misfolded protein detection: The unfolded protein response (UPR) is activated when misfolded or unfolded proteins accumulate, leading to →
Halting protein synthesis
Upping regulation of chaperone production
Transport of misfolded proteins to the cytosol for degradation.
Initial glycosylation (adding sugars to proteins)
Assembly of multi-subunit proteins
Smooth ER
Lacks ribosomes, characterized by tubular membranes.
Functions are NOT related to proteins. Plays roles in…
Lipid synthesis (steroids)
Drug detoxification via cytochrome P450 enzymes
Carbohydrate metabolism (glycogen breakdown, ex. G6P)
Calcium storage for muscle contraction (sarcoplasmic reticulum in muscle cells).
The ER are the Primary Source of Membrane Lipids, with the exceptions of… (a few organelles are specialized and create specific lipids for themselves. )
Mitochondria → phosphatidylethanolamine
PE is a type of phospholipid crucial for the inner workings of the mitochondrion.
Peroxisomes → cholesterol
peroxisomes contain special enzymes that help build cholesterol from scratch, especially from specific fats.
Chloroplasts → chloroplast-specific lipids
fats needed for photosynthesis, The machinery to build these specific types of lipids is located within the chloroplast itself, not the ER.
How Phosolipids are made…
Fatty acids are synthesized in the cytoplasm→
Added to cytosolic leaflet of ER membrane
The “Head” group is then added
Flippases move lipids to the other lumenal leaflet
Flippases are specific
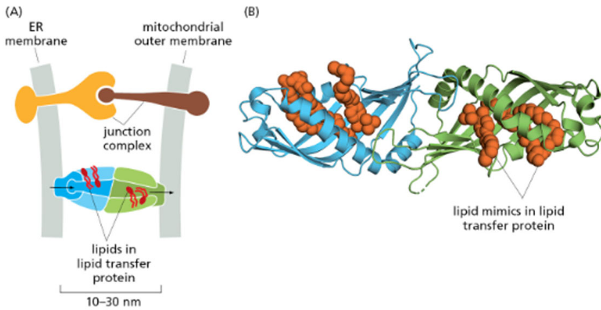
ER forms membrane contact sites with:
ER + Mitochondria
The ER sends lipids needed for the mitochondrial membrane. It also acts as a "manager," helping to divide the mitochondria (fission) and regulating energy production.
ER + Chloroplasts
The ER provides essential lipids to the chloroplast membrane to keep it running.
ER + Peroxisomes
The ER helps build peroxisomes, and the contact sites are used to share fats (fatty acids) that the peroxisome breaks down
Lipid transfer proteins move lipids directly (NOT vesicles), since it’s faster and more efficient. It allows for precise regulation of the lipid composition of the recipient organelle
Membrane Composition Difference
ER Membrane (~5 nm, Low Cholesterol): This is the raw material. It needs to be thin, soft, and very fluid to allow proteins and lipids to be built, inserted, and packaged quickly.
Plasma Membrane (7–8 nm, High Cholesterol): This is the final product. It is the outer edge of the cell, so it needs to be tough, thick, and protective
Membrane | Cholesterol | Thickness |
|---|---|---|
ER | Low | ~5 nm |
Plasma membrane | High | 7–8 nm |
**Cholesterol increases membrane thickness and rigidity.
Golgi Apparatus
Plays central role in membrane and protein trafficking in eukaryotic cells.
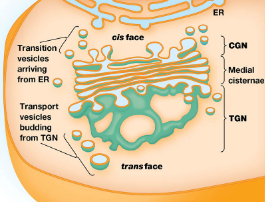
Structure
a series of flattened membrane-bounded cisternae, which form a Golgi stack.
Consist of 3 regions:
Cis-Golgi Network (CGN) → faces ER
Medial cisternae → Middle region with distinct enzymes
Trans-Golgi Network (TGN) → faces plasma membrane; where vesicles bud off for secretion.
Function
Modifies proteins and lipids received from the ER before distribution
Sorting and trafficking proteins and lipids
Transport Mechanisms - Movement through the Golgi takes place by either:
Stationary Cisternae Model: Cisternae remain in place while vesicles transport materials between them.
Cisternal Maturation Model: Cisternae themselves move through the Golgi, changing their functions as they mature. Enzymes are recycled backward
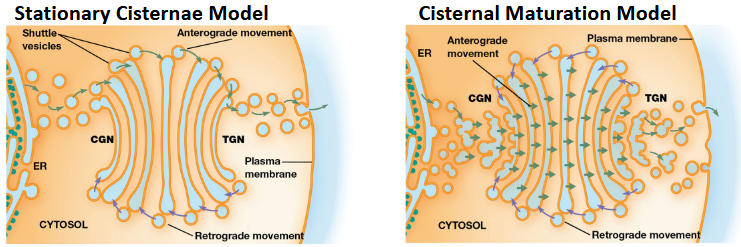
Anterograde and Retrograde Transport:
Anterograde Transport is the movement toward plasma membrane. ER → Golgi → Plasma membrane.
Retrograde Transport is the movement back toward ER. Plasma membrane → inside (endocytosis) → Golgi → ER
Purpose: Prevent membrane buildup; Return misplaced proteins
BIG PICTURE: The ER and the Golgi work together, functioning as one connected system for protein processing.
The 3 BIG protein-processing functions:
Protein folding
Quality control (Unfolded Protein Response, UPR)
Glycosylation (adding carbohydrates)
These processes start in the ER and continue in the Golgi.
Protein Folding in the RER
Molecular chaperones assist in folding proteins correctly. These chaperones bind non-polar regions of proteins to facilitate proper folding into their three-dimensional shapes. Without proper folding, misfolded proteins can accumulate, leading to cellular stress responses known as the unfolded protein response.
ex. BiP clamps onto hydrophobic regions as folding happens
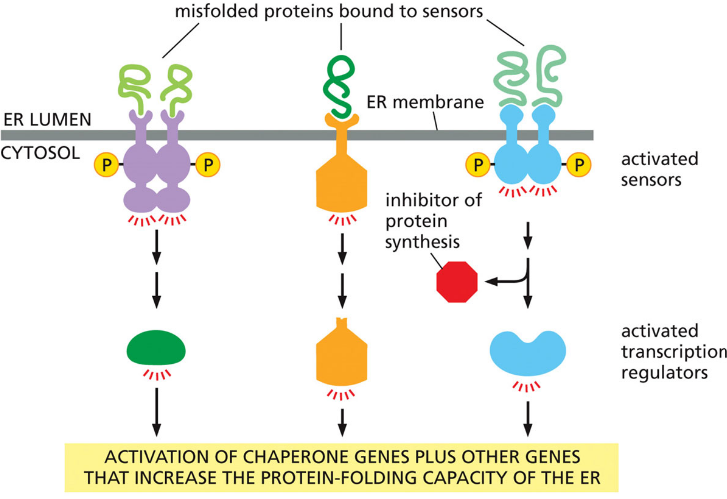
Unfolded Protein Response (UPR) - Triggered when misfolded proteins accumulate in ER. UPR does…
Stops general protein synthesis
Increases chaperone production
Exports misfolded proteins to cytosol
Misfolded proteins destroyed by proteasome
Purpose: protect the cell from toxic protein buildup.
Glycosolation
There are 2 types:
N-linked glycosylation
Sugar attached to Nitrogen (N) of asparagine
Begins in rough ER → Continues in Golgi
Added co-translationally (while protein is being made)
O-linked glycosylation
Sugar attached to Oxygen (O) of serine or threonine
Happens only in Golgi
Added post-translationally
Why Glycosolation Matters: Help protein folding. Act as cell ID markers (glycoproteins, glycolipids). Help protein–protein interactions. Affect stability and function.
O-linked Glycosylation Process
Happens in Golgi
Sugars added one at a time
Attached to the (-OH) group on the side chains of:
Serine
Threonine
Sometimes tyrosine
Unlike N-linked, no big core sugar added at once.
N-linked Glycosylation Process
2 N-acetylglucosamine + 9 mannose + 3 glucose = Total 14 sugars
Build Core Sugar on Dolichol Phosphate
Dolichol phosphate = hydrophobic lipid in ER membrane
Sugars added on cytosolic side:
2 N-acetylglucosamine and 5 mannose
Flip into ER Lumen
Flippase enzyme flips dolichol + sugars into ER lumen
Finish Core in Lumen
4 more mannose and 3 glucose are added
Now full 14-sugar core oligosaccharide exists.
Transfer to Protein
By using the Enzyme Oligosaccharyl transferase →
Transfers core sugar to asparagine residue
Happens while protein is being synthesized (co-translational)
Processing in ER + Golgi
Sugars can be Trimmed, Modified, and Additional sugars added
Final glycoprotein structure varies.
Which Asparagine Gets Glycosylated? Not every Asparagine (N) in a protein gets a sugar attached. The cell’s machinery only adds sugar to an Asparagine that is part of a specific "Recognition Sequence" or code
The Code: N – X – S/T
N = Asparagine (The anchor spot)
X = Any amino acid (literally, any of the 20)
S/T = Serine or Threonine
BUT if If X = proline → NO glycosylation: Proline causes a sharp, rigid bend (a "kink") in the protein chain. This kink makes it physically impossible for the enzyme to fit and attach the sugar to the Asparagine
Why not all asparagine get glycosylated? Factors include…
Protein folding speed
Accessibility of residue
Surrounding amino acids
Enzyme active-site constraints
Protein length (first ~12–14 aa often NOT glycosylated)
Compartmentalization ensures correct modification order.
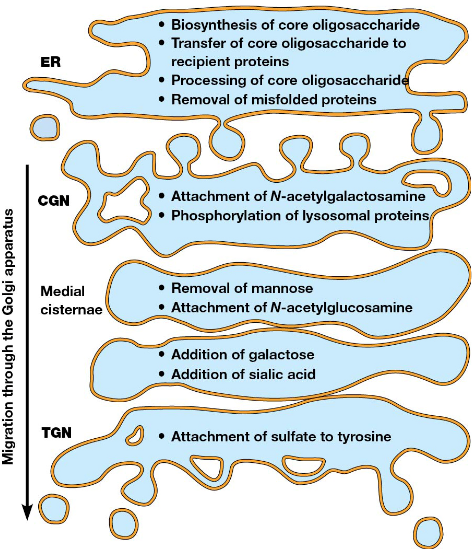
Protein Trafficking from the ER to the Golgi
1. ER Exit Sites (ERES)
What they are: Specific, specialized regions on the Rough ER membrane.
Key Feature: Unlike the rest of the rough ER, these areas have no ribosomes attached.
Function: They act as "loading docks" where mature, properly folded proteins are collected and concentrated to leave the ER.
Action: Transport vesicles, coated with a protein called COPII, bud off from these sites, carrying the cargo
2. Vesicular Tubular Clusters (VTCs) / ERGIC (ER Golgi Intermediate complex)
What they are: Small vesicles that budded off the ER fuse together to form larger, irregular structures.
Intermediate Step: These VTCs act as a "sorting center" or intermediate station before reaching the Golgi.
Alternative Name: These structures are collectively called the Endoplasmic Reticulum-Golgi Intermediate Complex (ERGIC).
Function: They move along microtubule tracks towards the Golgi, serving as the transporter
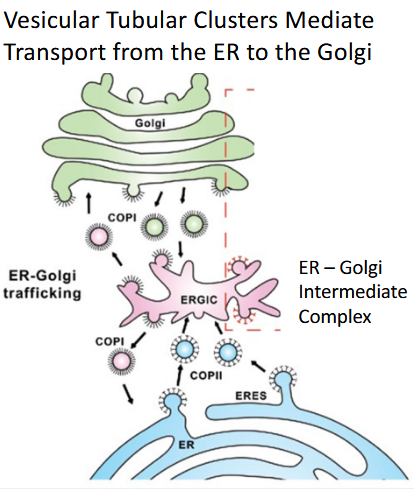
Transport Pathway
The pathway follows a strict, one-way, "anterograde" (forward) route:
ER (Production): Protein is synthesized and folded.
ERES (Loading): Protein is packaged into COPII vesicles.
ERGIC/VTC (Transport): Vesicles fuse to move cargo toward the Golgi.
Cis-Golgi (Receiving): VTCs fuse with the "cis" (entry) face of the Golgi.
Medial-Golgi (Processing): Proteins are modified (e.g., glycosylation).
Trans-Golgi (Sorting/Shipping): Final sorting and packaging.
Final Destination: fuse with endosomes to develop Lysosomes, some of those vesicles are going to be secretory vesicles, go to the plasma membrane, or secretion outside the cell (exocytosis).
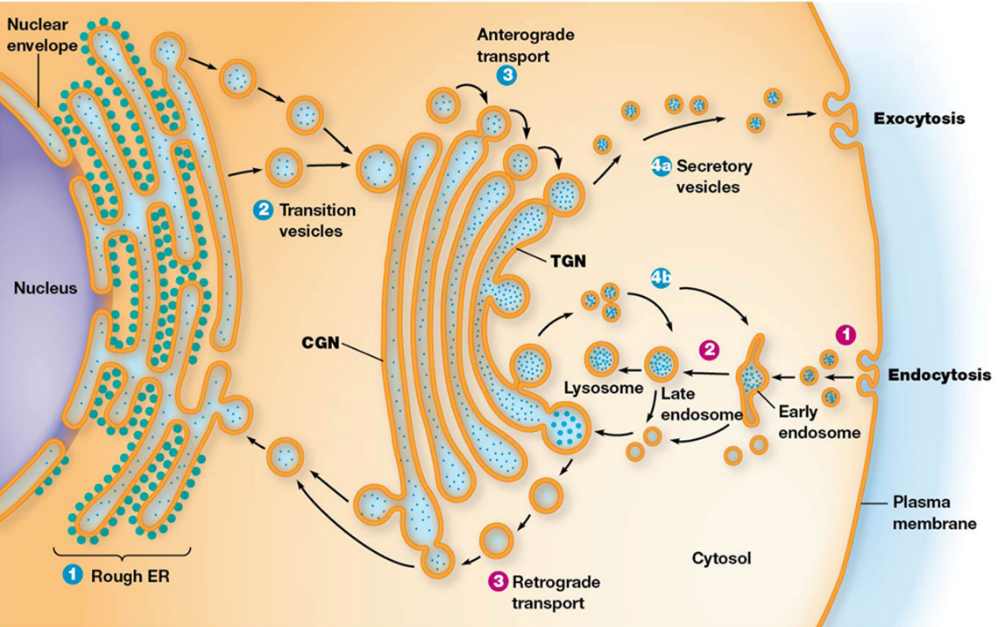
Secretion Pathways
Constitutive Secretion - “The Automatic Stream"
Flows continuously. This process happens 24/7 without any special instructions or triggers.
Default pathway: If a protein doesn't have a specific tag telling it to go elsewhere, it follows this path automatically.
t supplies the "bricks and mortar" for the cell, such as membrane proteins to repair the cell's outer wall or ECM proteins to build the scaffolding outside the cell.
Regulated Secretion - “The On-Call Delivery”
Storage: Proteins are packed into vesicles and sit near the "exit door," just waiting.
The Signal: They are only released when the cell receives a specific "Go!" command (like an electrical impulse or a chemical signal).
High-stakes cargo like hormones (e.g., insulin waiting for blood sugar to rise) or neurotransmitters (waiting for a brain signal) use this pathway.
How Proteins Know Where to Go (Targeting Tags)
ER Retention Tags used for Quality Control
Prevent proteins from leaving ER.
Main retention tag: RXR (Arg-X-Arg)
Example: NMDA receptor (a protein used in brain signaling) is a great way to see this in action. Imagine it as a complex piece of IKEA furniture that comes in several parts.
If folded correctly → tag hidden → Leaves ER: it "tucks" the RXR tag into its center. Because the tag is now hidden inside the protein's folds, the "security guards" can't see it. The protein is cleared for departure and moves to the Golgi.
If misfolded → tag exposed → stays in ER: it can't hide the RXR tag. The tag remains exposed on the surface. The cell sees this "Not Ready" signal and forces the protein to stay in the ER to be fixed or recycled
ER Retrieval Tags used as a search team
These tags are short amino acid sequences at the very tail end of a protein. If the Golgi detects these specific sequences, it knows the protein is out of place.
KDEL (Lys-Asp-Glu-Leu): The most famous retrieval tag. It is used by soluble proteins (proteins floating in the fluid).
KKXX (Lys-Lys-Any-Any): Used by proteins that are actually embedded in the membrane.
HDEL: The version of KDEL in yeast.
Golgi Proteins stay in the Golgi using 3 mechanisms
Retrieval Tags (like ER)
Large complexes that are excluded from transport vesicles
Too big to fit into vesicles → stay in Golgi
Membrane Thickness Mechanism
Transmembrane proteins have hydrophobic regions
Membrane gets thicker from ER → Golgi → plasma membrane
If hydrophobic region is too short → protein stays in cis-Golgi
Targeting of Lysosomal Enzymes
made in: Rough ER → Golgi → vesicles → lysosomes
(The Key Targeting Enzyme: Mannose-6-Phosphate)
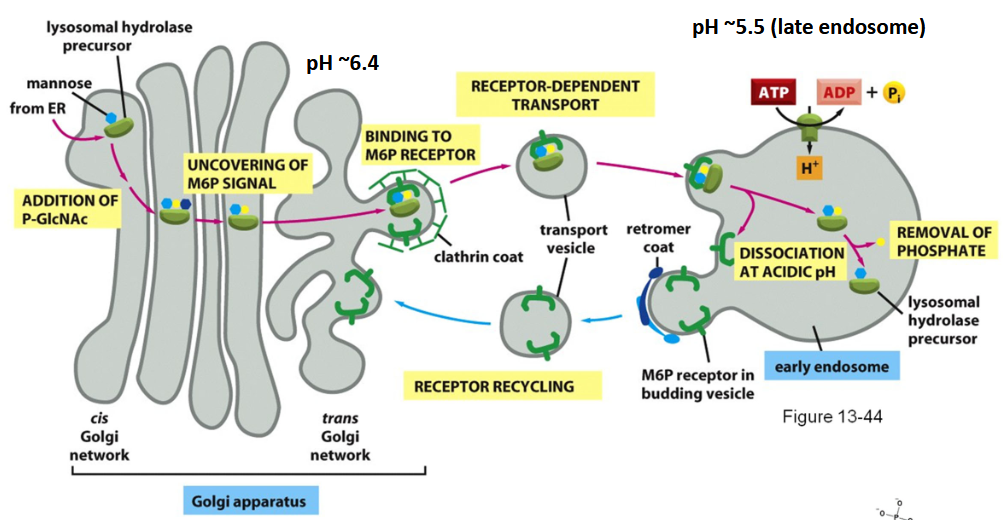
Steps:

Don’t forget: Lysosomes use carbohydrate tags, NOT amino acid tags.
Lysosomes act as the cell's recycling and waste disposal system, breaking down materials within the cell.
These contain digestive enzymes to break down substances.
The cell takes in external materials through processes like phagocytosis and receptor-mediated endocytosis, which are then broken down by lysosomes.
Damaged or unneeded cellular structures are broken down and recycled through a process called autophagy.
Lysosomes develop from endosomes. The enzymes they use are made on the rough endoplasmic reticulum (RER) and delivered via the trans-Golgi network (TGN) to the endosomes in transport vesicles. These endosomes then mature into functional lysosomes.
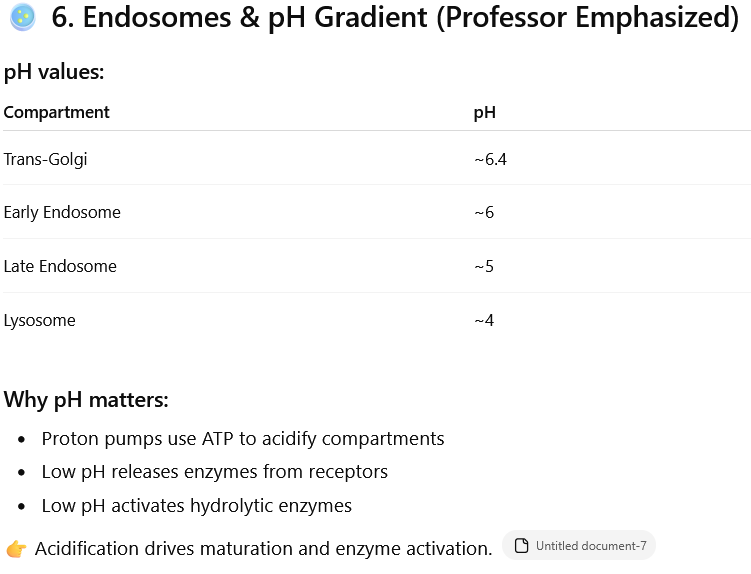
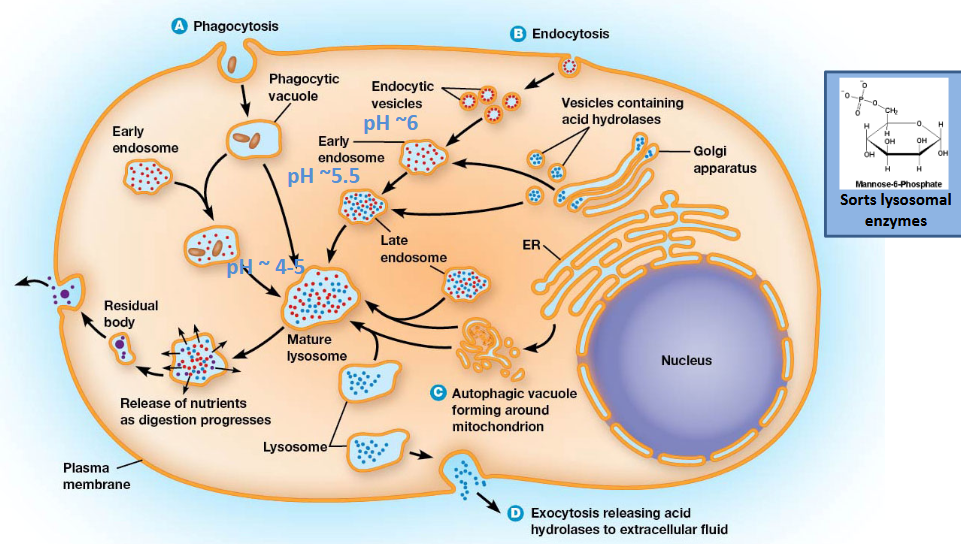

Why pH matters: This increasing acidity is crucial for several reasons:
Acidification: Proton pumps use energy (ATP) to make the compartments more acidic.
Enzyme activation: The low pH activates specific enzymes (hydrolytic enzymes) that break down waste materials.
Enzyme release: Low pH helps release enzymes from their receptors so they can perform their function.
Maturation: This acidification process is essential for the proper maturation of these cellular compartments and the activation of their enzymes.
Lysosomes are like the cell's recycling centers or "stomachs," breaking down waste materials. They get the things they need to do this job from two main places:
From the cell's internal "factory": Small sacs (vesicles) carrying special breakdown chemicals (enzymes with M6P) come from the Golgi apparatus.
From outside the cell: Other sacs (endocytic vesicles) bring in materials from the cell's outer edge (plasma membrane).
The Formation Happens in 3 Steps:
Merging: The sacs from inside and outside the cell join together to form an "early endosome".
Changing the environment: The early endosome starts pumping in protons, making the inside more acidic (lower pH), and turns into a "late endosome".
Final step: The late endosome becomes even more acidic, finally turning into a mature, working lysosome.
Multivesicular bodies are intermediate structures formed when endosomes fold their own membrane inward, creating vesicles inside vesicles, which then become late endosomes and fuse with lysosomes.
Endosomes can internalize their own outer layer.
This forms structures that look like a bubble filled with smaller bubbles inside (multivesicular bodies).
These specific structures eventually mature into late endosomes and join with existing lysosomes to dispose of their internal contents
Membrane Balance
Exocytosis:
Vesicles fuse with plasma membrane
Releases contents
Adds membrane lipids/proteins
Ex. Sperm release enzymes to penetrate egg coat (Lysosomes can digest outside the cell too)
Endocytosis:
Plasma membrane invaginates
Removes membrane and brings material inside
👉 Together they maintain membrane size and composition.
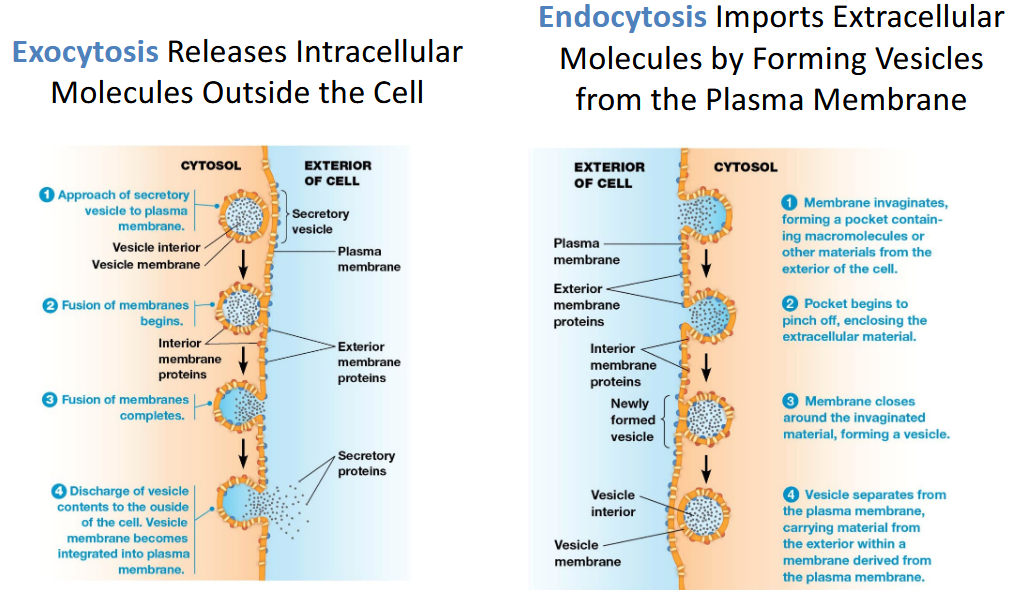
Endocytosis Includes Phagocytosis, Pinocytosis, and Receptor-Mediated Endocytosis.
Phagocytosis (“Cell Eating”)
Large particles (bacteria, debris)
Pseudopods extend around particle
Vesicle called phagosome
Used by macrophages, immune cells
Pinocytosis (“Cell Drinking”)
Small extracellular fluid droplets
Continuous process
Cell samples environment
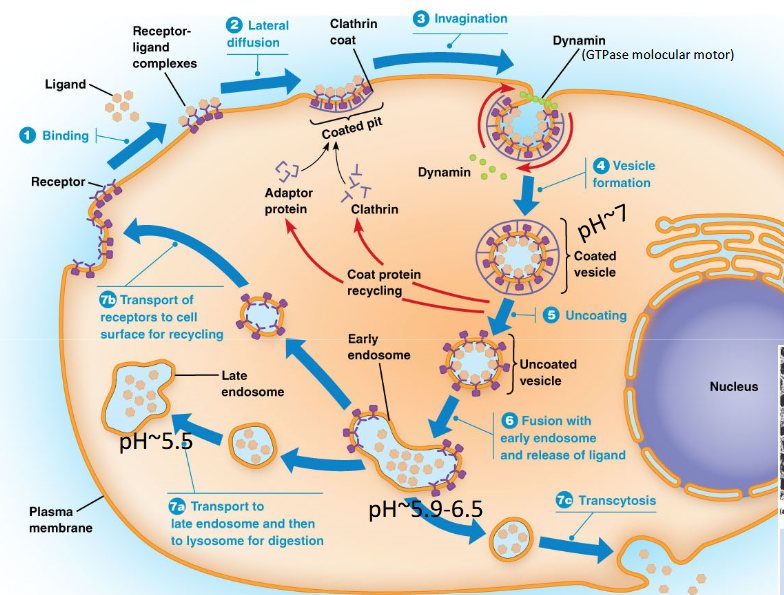
Receptor-Mediated Endocytosis (VERY HIGH-YIELD)
Steps:
Ligand binds receptor
Receptors cluster in clathrin-coated pits
Clathrin lattice forms vesicle
Dynamin pinches vesicle off (GTPase)
Coat removed
Vesicle fuses with early endosome
Uses:
Cholesterol (LDL)
Hormones
Growth factors
Nutrients
👉 Main nutrient uptake pathway
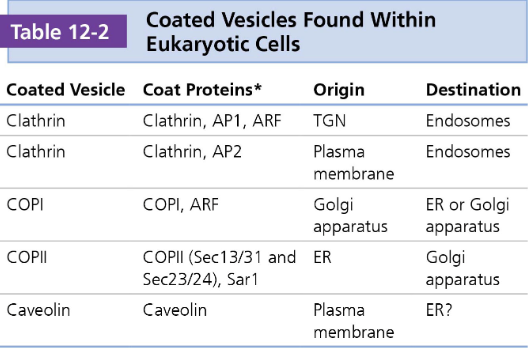
Vesicle Coat Proteins
Vesicles require a "coat" of proteins to form and to select the correct cargo for transport. Different coat proteins are associated with specific pathways and directions of movement.
The type of coat protein on a vesicle helps to determine the destination of the vesicle
COPII: These proteins help move items from the endoplasmic reticulum (ER) to the Golgi apparatus. This is an "anterograde" (forward) direction of transport.
COPI: These proteins manage traffic in the opposite direction, from the Golgi back to the ER ("retrograde" transport), or between different parts of the Golgi itself.
Clathrin: This coat is involved in endocytosis (bringing materials into the cell from the outside) and transport from the trans-Golgi network. \
Dynamin is an accessory protein crucial for the final step of vesicle formation.
It is a GTPase protein, meaning it uses energy from GTP to function.
It wraps around the "neck" or stalk of a budding vesicle.
It then twists the membrane, effectively pinching off the vesicle from the original membrane.
This protein is essential for processes like receptor-mediated endocytosis.
Coat proteins enable vesicle formation, but what mediates vesicle fusion (to the ER, PM, etc.)?
SNARE proteins!
SNARE proteins show the vesicles which targets to bind to.
SNARES attach to the vesicle and the its target
There are complementary proteins: v- (vesicle) and t- (target) SNARES
The vesicle has the v- SNARE, along with Rab GTP-ase
The Vesicle moves to the target
The target has the short t-SNARE, with long tethering complex to grab the vesicle
If the v-SNARE and t-SNARE are complementary to each other, they bind w/ the help of Rab GTPase leading to fusion of the membrane
SNAPS and NSF’s then break off the uneeded SNAREs and Rab GTP-ase for recycling
Perioxsome
Carry out oxidative rxns with a H2O2 byproducts
Distinguisig Periroxsomes from other by looking at crystalline core…
Urate oxidase in animals
Catalase in plants
Perioxsomes form by divison of eachother or peroximal ghost
peroximal ghost: membrane could bud off from ER and forms the shells inwhich peroximaal enezymes can be added
Roles of Peroximsome
Metabolizes harmful hydrogen peroxide
Oxidases generate hydrogen perixide
Catalase detoxifed H2O2
Detoxification of harmful compounds out of substances
Oxidizes long chain FA
Metabolizes nitrogen containing compounds
Breaks down unusual substances