Unit 6 classoom notes AI flashcards
Unit 6 Gene Expression & Regulation
6.1 DNA & RNA structure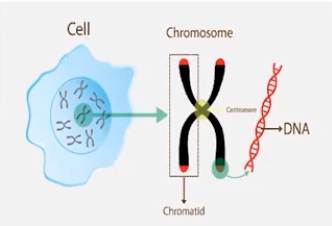
DNA & in some cases RNA, is primary source of heredity material
- Genetic information is stored as the sequence of bases in DNA & RNA
- Genetic information is transmitted from one generation to the next through DNA or, in some cases RNA
- DNA, is packaged into chromosomes before genetic information is passed from parent to daughter cells
- Many viruses use RNA molecules to encode genetic information
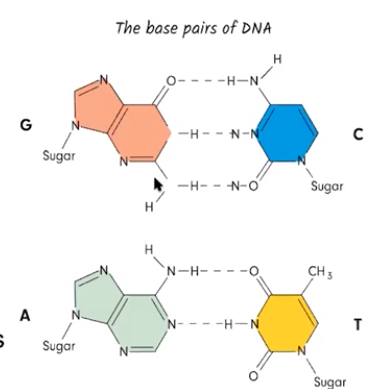
DNA, & sometimes RNA, exhibits specific nucleotide base pairing
- DNA & RNA are structurally similar in the following ways:
- Both are polymers containing nucleotides
- Both are chain-like molecules
- Both follow base pairing rules
- When DNA nucleotides pair with each other, adenine pairs with thymine & guanine pairs with cytosine
- When RNA nucleotides pair with DNA nucleotides, there is one base pair change in which RNA’s uracil nucleotides pair with DNA’s adenine nucleotides
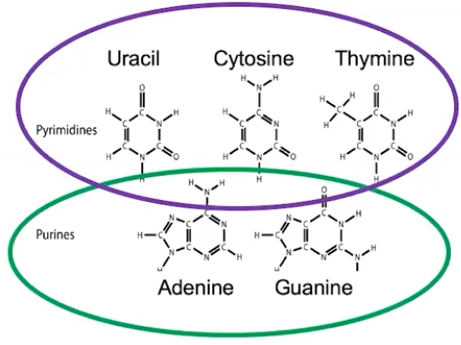
Specific nucleotide base pairing is conserved through evolution
- DNA & RNA both follow conserved base pairing rules in which pyrimidines pair with only specific purines
- Pyrimidines have a single ring structure. Examples are cytosine, uracil, & thymine (remember: “pyramids CUT”)
- Purines have a double ring structure. Examples are adenine & guanine
Prokaryote & eukaryote genomes differ in certain aspects.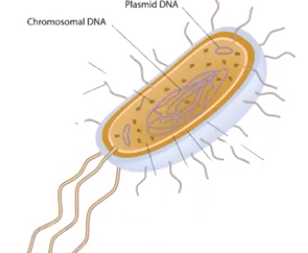
- Prokaryotic organisms typically have circular chromosomes while eukaryotic organisms have multiple, linear chromosomes
- Prokaryotic genomes are typically smaller than eukaryotic genomes
- Prokaryotes & eukaryotes can contain plasmids, which are small extrachromosomal, double-stranded, circular DNA molecules
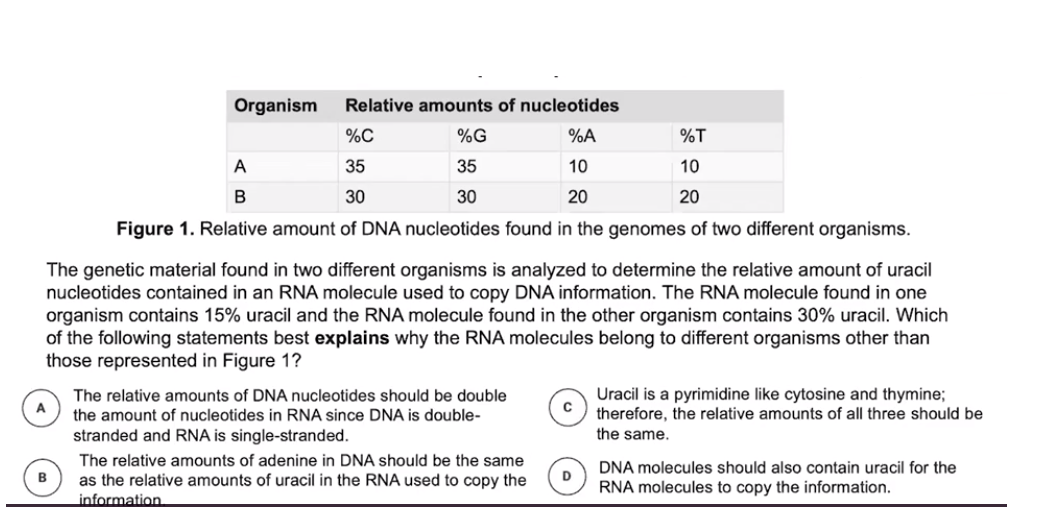
- Prokaryotic plasmids are found in the cytosol while eukaryotic plasmids are found in the nucleus
Ex1
6.2 Replication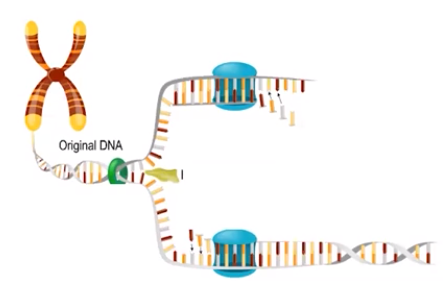
DNA replication ensures continuity of hereditary information
- Genetic information is copied during DNA replication
- Occurs before cell division
- Allows transmission of a complete genome from one generation to the next
- Specific replication mechanisms ensure the continuity of hereditary information
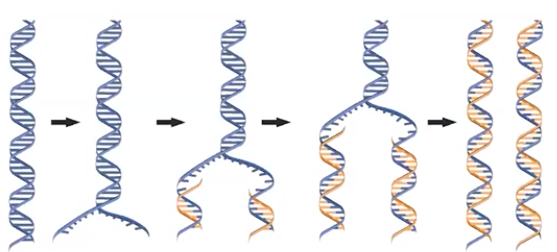
DNA replication is semiconservative.
- Semiconservative replication results in a DNA molecule containing one original strand and a newly synthesized compliment
- Each original strand(blue) serves as a template for the synthesis of a new complementary strand
Directionality in a DNA molecule influences the replication process.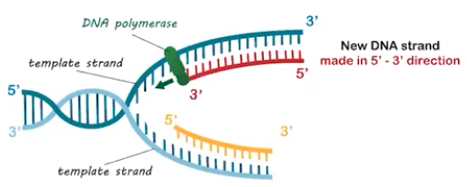
- Each DNA strand has a terminal phosphate group on one end & a terminal hydroxyl group (OH) on the other end
- Phosphate terminus is referred to as the 5-prime end (5’)
- Hydroxyl terminus is referred to as the 3-prime end (3’)
- The two strands of a DNA molecule run antiparallel to each other where the 5’ end of one strand is opposite the 3’ end of the complementary strand
- Nucleotides can only be added to a growing strand in a 5’ → 3’ direction
- One strand will always be synthesized continuously = leading strand
- One strand will always be synthesized discontinuously, in fragments = lagging strand
Enzymes involved in DNA replication
- Helicase unwinds the DNA strands
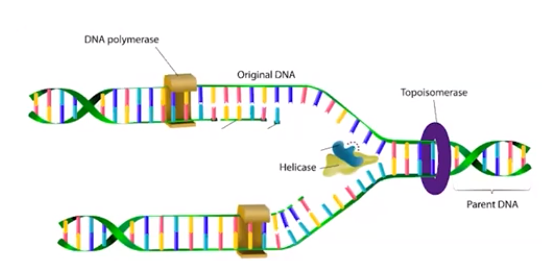
- Topoisomerase relaxes the supercoil at the replication fork
- Replication fork - the location where the two strands are separated
- DNA polymerase synthesizes new strands
- Requires RNA primers to initiate synthesis
- Attaches to 3’ end of the template strand
- Builds strands in a 5’ → 3’ direction
- Ligase joins DNA fragments on the lagging strand
Ex2
6.3 Transcription & RNA Processing
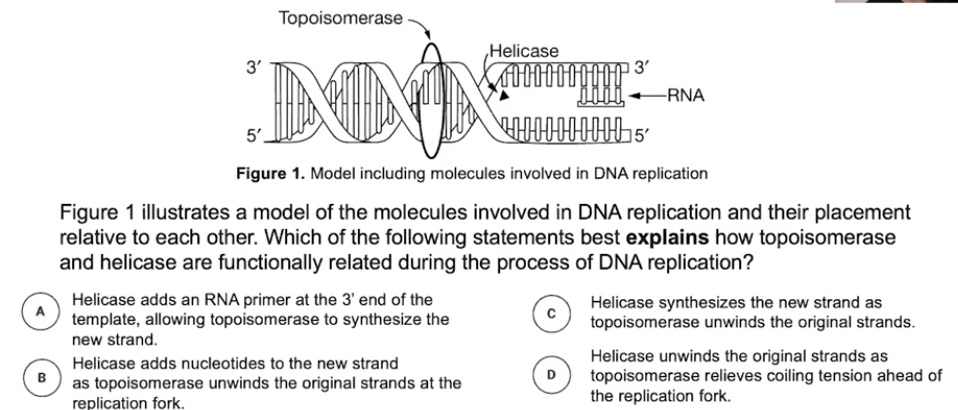
Genetic information flows from DNA to RNA to protein
- Genetic information is typically stored in DNA molecules
- RNA molecules are used to facilitate protein synthesis using DNA information
- Ribosomes contain RNA & assemble proteins
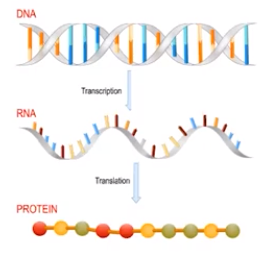
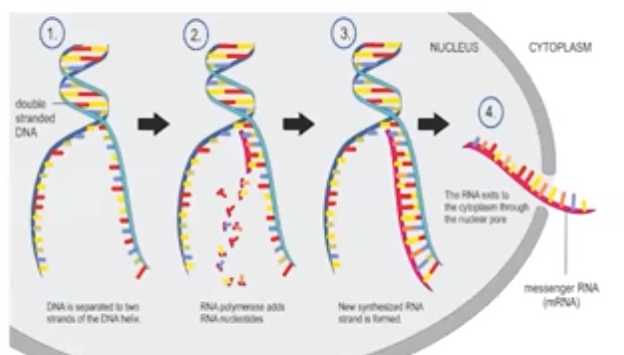
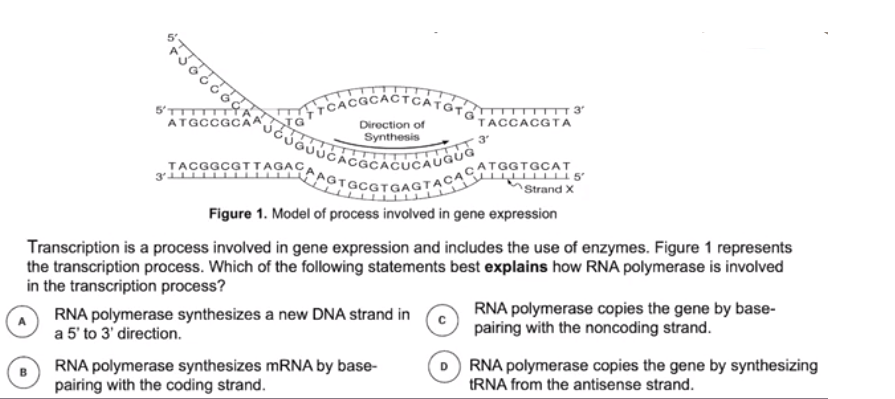
Transcription is the process in which an enzyme directs the formation of an mRNA molecule
- DNA strands are separated during transcription.
- One strand serves as the template strand, also known as the noncoding strand, minus strand, or antisense strand.
- The other strand is the non-template strand, also known as the coding strand.
- The strand serving as the template strand depends on the gene being transcribed.
- The gene targeted for transcription is located on the coding strand
- The enzyme RNA polymerase synthesizes messenger RNA (mRNA) in the 5’ → 3’ direction reading the template in the 3’ - 5’ direction.
- mRNA molecule is the transcribed copy of a particular gene
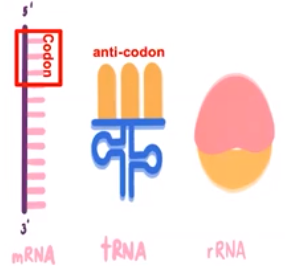
There are 3 types of RNA molecules
- Messenger RNA (mRNA) carries genetic information from DNA to the ribosomes.
- Information is used to direct protein synthesis at the ribosomal site
- A codon is a three-base sequence found on mRNA
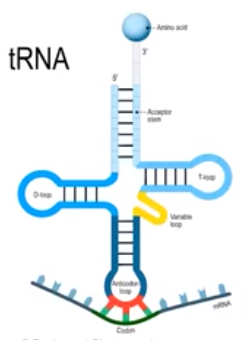
- Transfer RNA (tRNA) is recruited to the ribosome to help create a specific polypeptide sequence as directed by mRNA
- There are various tRNA molecules, each carry a specific amino acid
- An anti-codon is a three-base sequence on tRNA
- Correct base pairing of tRNA anti-codons with mRNA codons will result in the release and addition of an amino acid to a growing polypeptide.
- Ribosomal RNA (rRNA) molecules are functional units of ribosomes responsible for protein assembly.
- Base-pairing of anticodons & codons occurs in the ribosome
- Creates primary polypeptides as tRNA releases amino acids
Ex3.
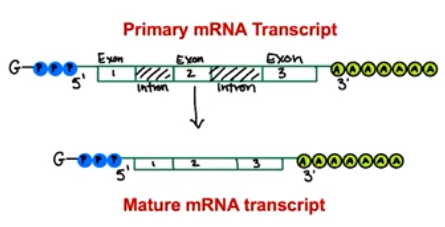
A series of enzyme-regulated modifications occur to the mRNA transcript in eukaryotic cells.
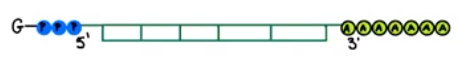
- Addition of a poly-A tail
- 100-200 adenine nucleotides
- Increases stability
- Helps with exporting from nucleus
- Addition of GTP cap
- Modified guanine nucleotide
- Protects the transcript
- Helps ribosomes attach to mRNA
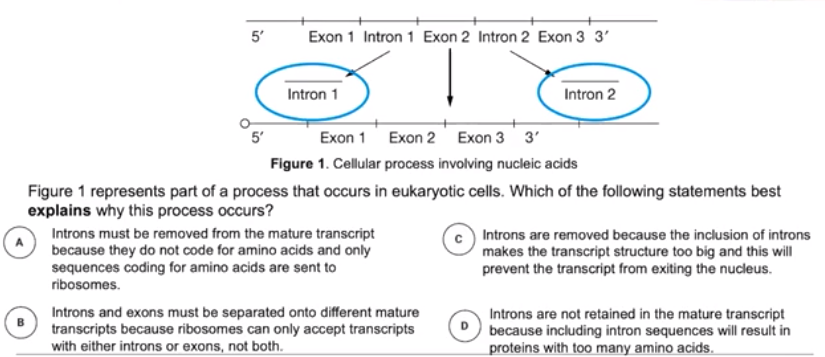
- Introns are sequences of an mRNA transcript that do not code for amino acids (interfering sequences)
- Are excised during RNA processing
- Not included in mature mRNA transcript
- Exons are sequences of an mRNA transcript that code for amino acids. (expressed sequences)
- Sequences are retained during RNA processing
- Different exons are connected in the mature mRNA transcript
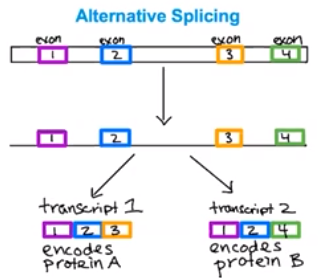
- The process of splicing introns and connecting retained exons in the mature mRNA transcript called alternative splicing.
- There are many different exons on a primary transcript
- Different mRNA transcripts can be produced from one primary transcript
- Exons can be retained in different variations
- Different mRNA transcripts lead to different proteins
- Different mRNA transcripts can be produced from one primary transcript
Ex4.
6.4 Translation
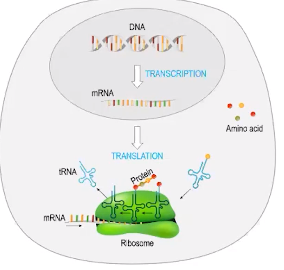
Translation of mRNA generates polypeptides
- Translation is the process by which an mRNA sequence is used to generate a polypeptide
- Translation occurs on ribosomes
- Prokaryotes only have cytosolic ribosomes
- Eukaryotes have cytosolic ribosomes & ribosomes bound to the rough endoplasmic reticulum
Translation involves energy & many sequential steps
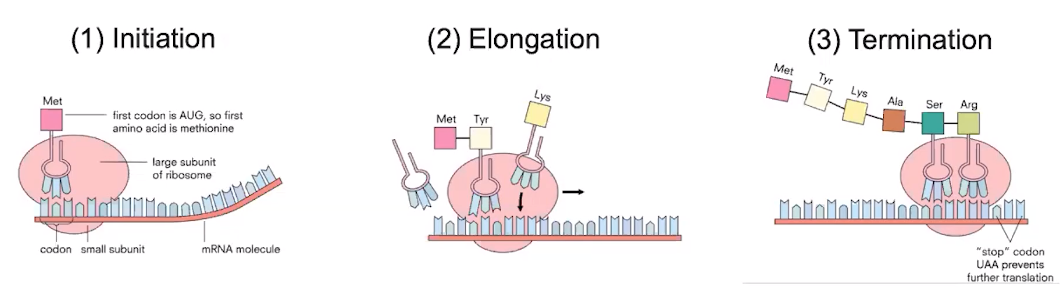
- Translation involves 3 main steps
- In prokaryotes organisms, translation occurs while the mRNA is being transcribed
- In prokaryotes organisms, translation occurs while the mRNA is being transcribed
Retrovirus translation is a special case because of an alternate flow of information
- Retroviruses introduce viral RNA, not DNA, into host cells.
- Reverse transcriptase is an enzyme that copies the viral RNA into viral DNA
- This is a viral enzyme
- This enzyme is introduced into the host cell.
- Once the enzyme converts the viral RNA into viral DNA, the DNA is integrated into the host genome. (*remember HIV replicates through the lysogenic cycle)
- Once integrated, the viral DNA will be transcribed & translated.
- Transcription & translation of the viral DNA results in assembly of new viral progeny
- Reverse transcriptase is an enzyme that copies the viral RNA into viral DNA
Nearly all organisms use the same genetic code.
- Translation mechanisms are similar in nearly all organisms.
- Nucleotides used to construct DNA & RNA molecules are common among organisms
- This conservation is evidence of common ancestry
- Viral DNA & RNA molecules are chemically compatible with host-cell genomes
- This allows host-cell translation mechanisms to work with viral genomes
Ex5.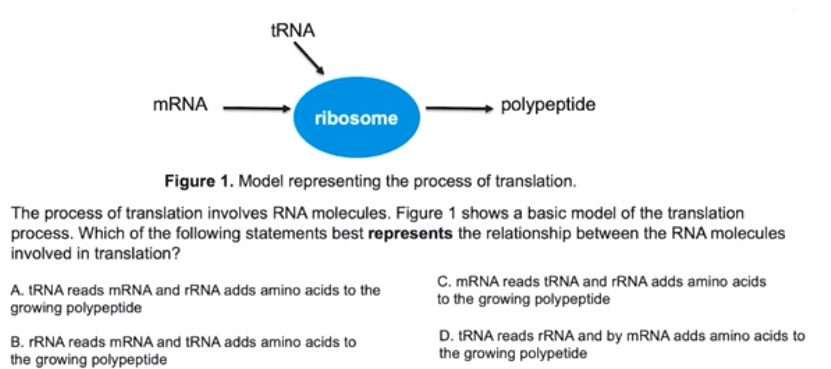
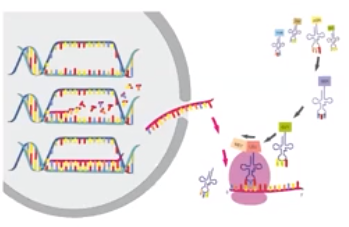
Translation involves many sequential steps
- Translation is the final process in the flow of information from DNA → RNA → Protein
- Involves converting RNA infrastructure into a protein
- The first step of translation is initiation.
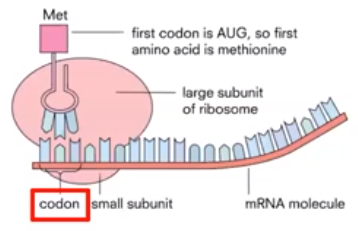
- rRNA in the ribosome interacts with the mRNA at the first start codon.
- mRNA nucleotides are grouped together & read in triplets called codons
- Each codon encodes a specific amino acid
- Can be deduced by using a codon chart
- Many amino acids are encoded for by more than one codon
- Each codon encodes a specific amino acid
mRNA Codon Chart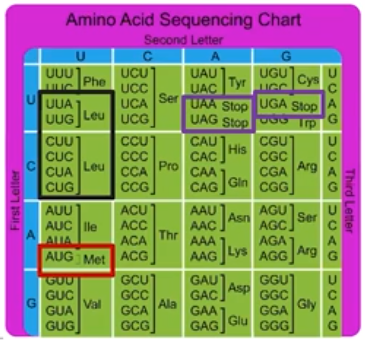
- This chart can be used to determine which codon encodes for each amino acid.
- There are 20 naturally occurring amino acids
- Some amino acids are coded for by more than one codon.
- For example, 6 codons encode leucine
- The starting codon (AUG) only codes for the amino acid methionine
- There are stop codons that do not code for an amino acid (AKA nonsense codons)
- Used to terminate translation
- Used to terminate translation
- Some amino acids are coded for by more than one codon.
- There are 20 naturally occurring amino acids
Translation involves many sequential steps.
- tRNA molecules bring the correct amino acid to the correct place, specified by the codon on the mRNA
- tRNA anti-codons must complement mRNA codons to help ensure the correct amino acid is brought to the ribosome
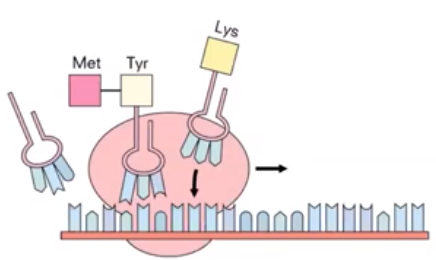
- The second step of translation is elongation
- Each newly arrived tRNA brings another amino acid to be added to a growing polypeptide chain.
- The rRNA adds the amino acids as tRNA brings them
- Each newly arrived tRNA brings another amino acid to be added to a growing polypeptide chain.
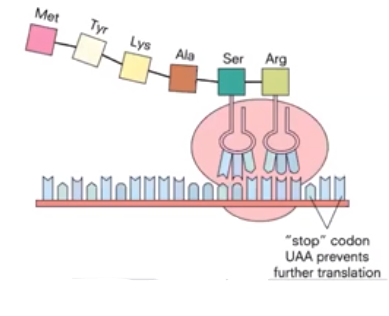
- The last step of translation is termination.
- Amino acids continue to be added to the growing polypeptide chain until a STOP codon is reached.
- Translation ends.
- No more amino acids added.
- Newly synthesized polypeptide is released
- Amino acids continue to be added to the growing polypeptide chain until a STOP codon is reached.
Ex6.

6.5 Regulation of Gene Expression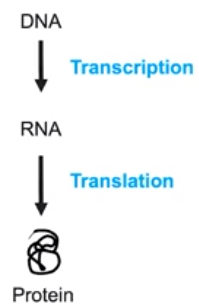
Differences in the expression of genes account for phenotypic differences between organisms.
- Gene expression is the process by which instructions in the DNA are transcribed & translated into a functional protein
- The flow of information is from DNA to RNA to protein
- Different types of chemical reactions regulate gene expression.
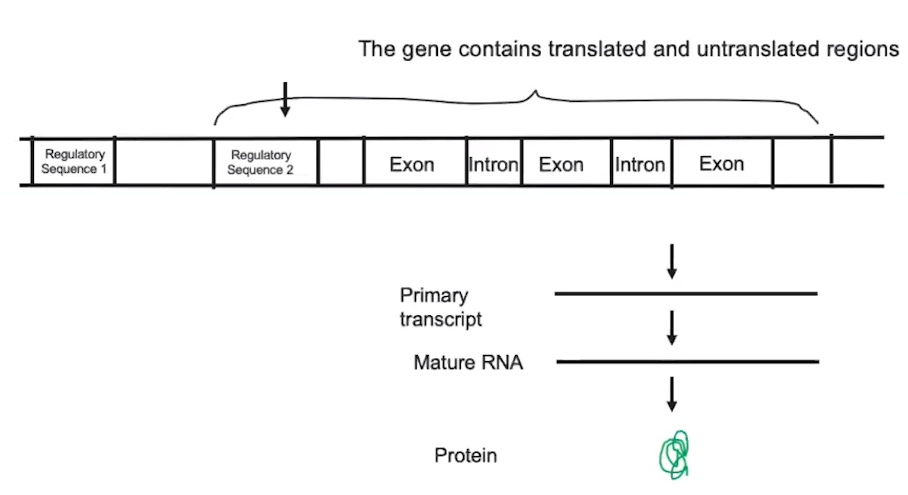
Regulatory sequences are stretches of DNA that interact with proteins
- Regulatory sequences are stretches of DNA that can be used to promote or inhibit protein synthesis.
- Regulatory proteins are used to assist with the promotion or inhibition of protein synthesis.
- The interaction of regulatory sequences with regulatory proteins controls transcription.
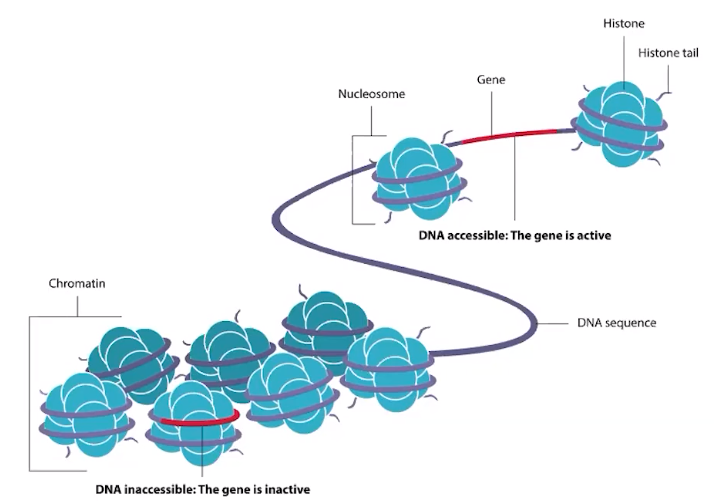
Epigenetic changes can affect gene expression.
- Epigenetic changes involve reversible modifications of DNA or histones
- Histones are proteins used to wrap DNA around
- Slight chemical modifications of DNA & histones cause tight packing or loose packing of DNA
- Packing & unpacking regulates gene expression
- Slight chemical modifications of DNA & histones cause tight packing or loose packing of DNA
- Histones are proteins used to wrap DNA around
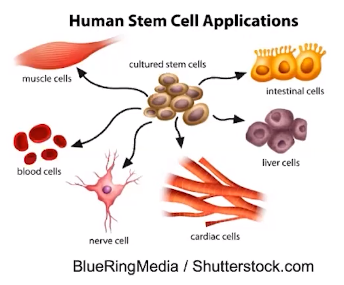
Observable cell differentiation results from the expression of genes for tissue - specific proteins
- The cells within a multicellular organism have the same DNA sequences
- Tissues are groups of cells that have the same function
- The presence of specific proteins within the cell of tissues give the tissue its function.
- The phenotype of a cell or organism is determined by the combination of genes that are expressed.
- Cell differentiation refers to cells within with the same organisms having different phenotypes
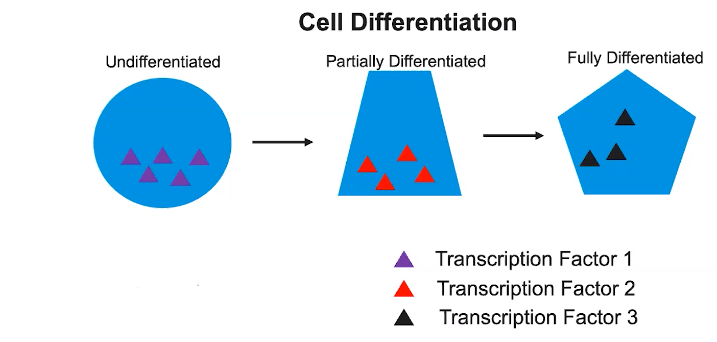
Induction of transcription factors during development results in sequential gene expression.
- Transcription factors are proteins that promote or inhibit transcription of a gene.
- The presence of various transcription factors during development helps determine how a cell differentiates.
- The process of development from an undifferentiated cell to a differentiated cell is a result of sequential gene expression.
Ex7.
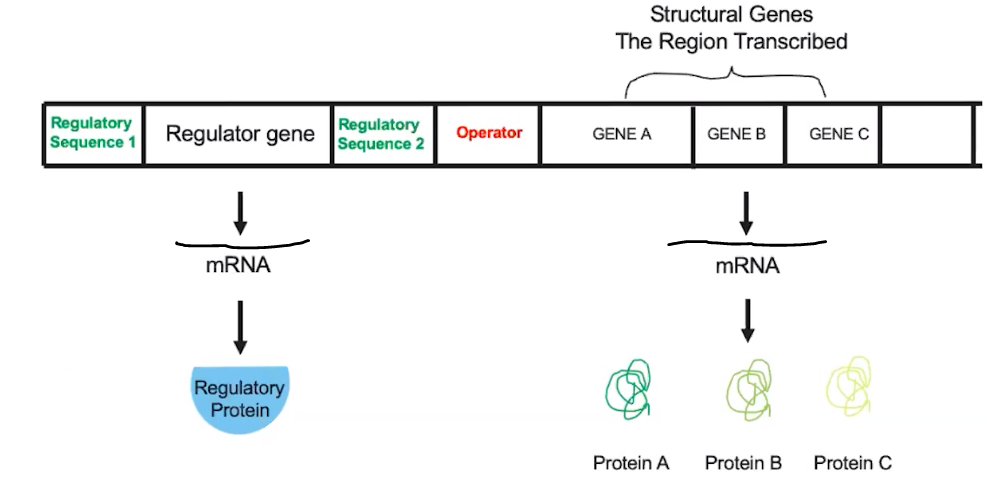
Both eukaryotes and prokaryotes have genes that are coordinately regulated.
- Operons are closely linked genes that produce a single mRNA molecule during transcription.
- They are under the control of the same regulatory sequence.
- Includes the genes to be transcribed, the regulatory sequence, & the operator.
- An operator is a sequence that either inhibits or promotes transcription by binding with regulatory proteins.
In prokaryotes, groups of genes called operons are transcribed in a single mRNA.
- Structural proteins with related functions are typically encoded together in the genome
- They are controlled by a single regulatory protein.
- Regulatory genes can control the expression of all the genes at the same time.
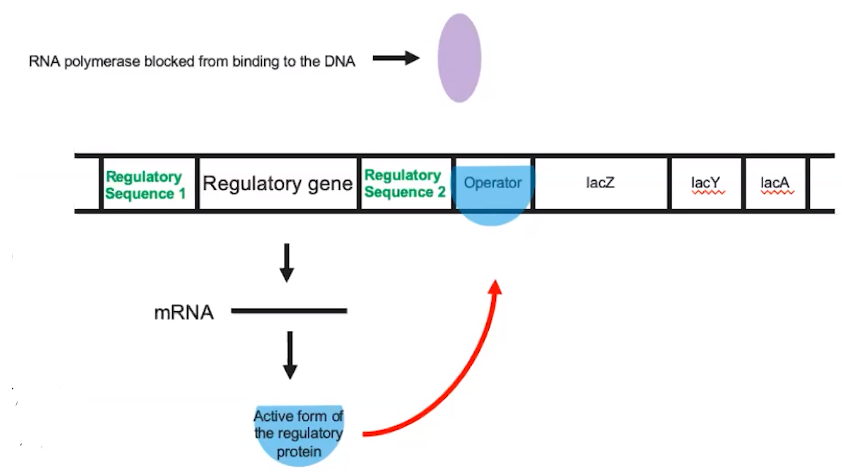
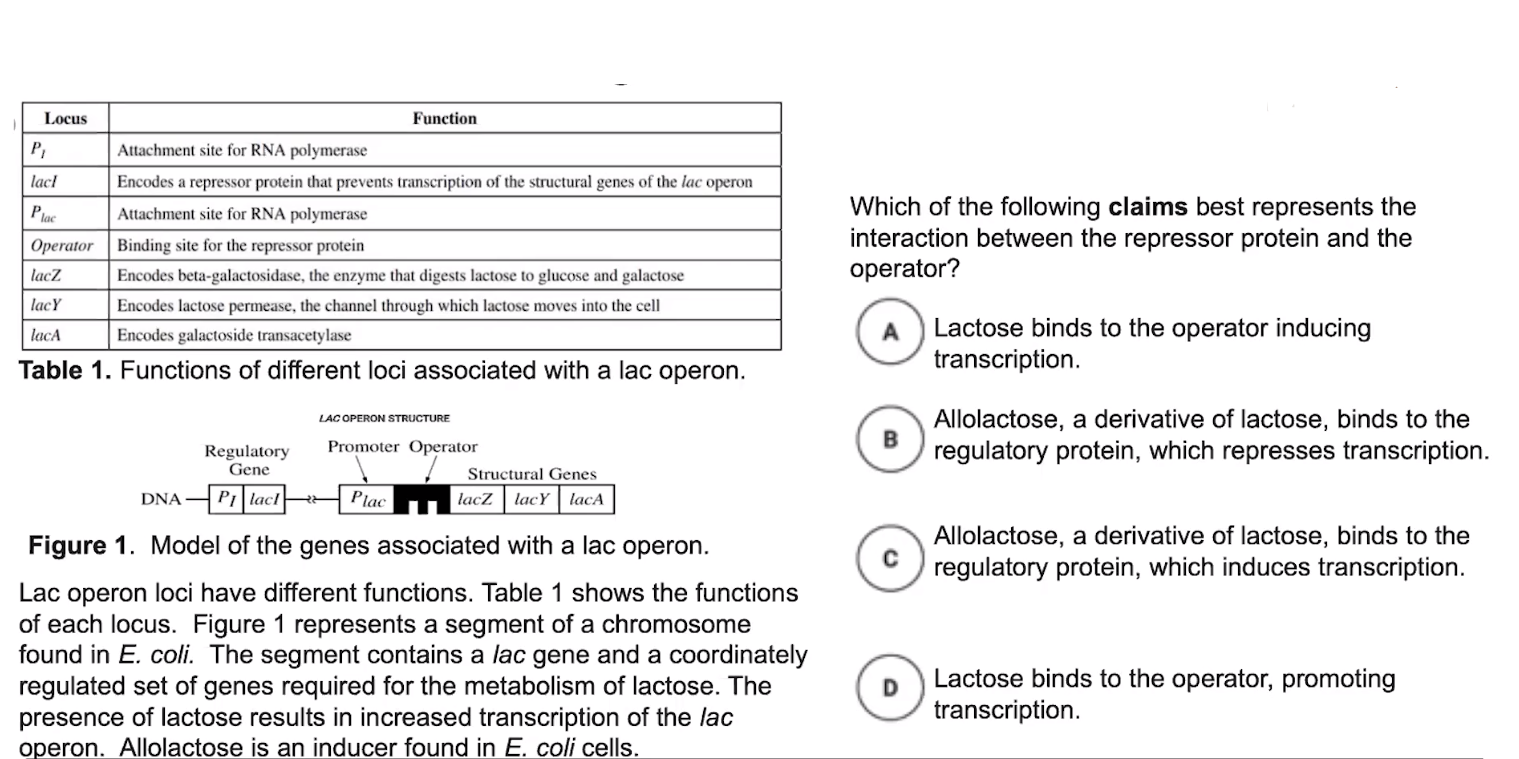
The lac operon is an example of an inducible system.
- The lac operon is considered inducible because it is usually turned off
- When the regulatory protein is bound to the operator, RNA polymerase cannot bind to the regulatory sequence
- This inhibits transcription of the genes that are part of the lac operon
- Inducers are molecules that can bind to the regulatory protein & cause it to change shape.
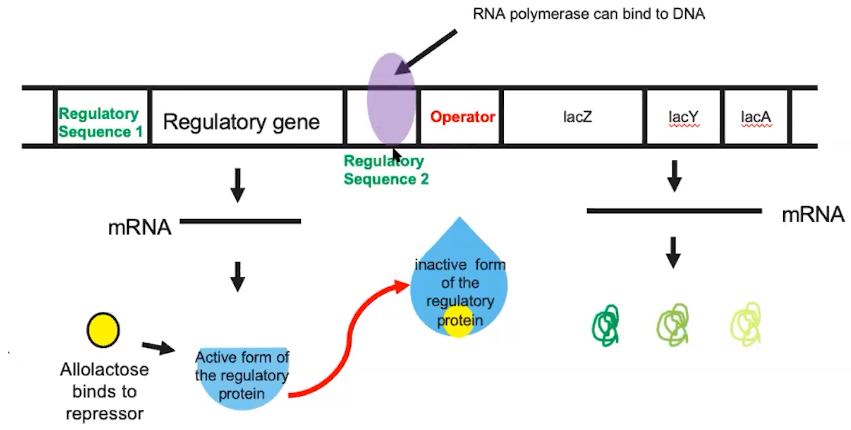
- When the inducer binds to the regulatory protein, the protein changes shape
- For example, allolactose = inducer
- This causes the regulatory protein to release from the operator, & frees up RNA polymerase to transcribe the operon’s genes.
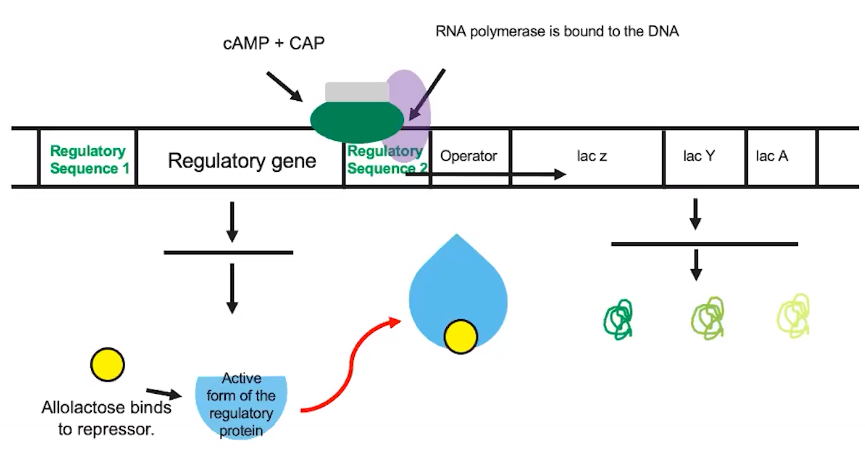
Example: Other transcription factors help regulate the lac operon. →
- The amount of glucose in the cell also helps to regulate gene expression
- When glucose is low, other transcription factors bind to the regulatory sequence to further promote transcription.
- When glucose levels are high, these transcription factors are not present
Ex8.

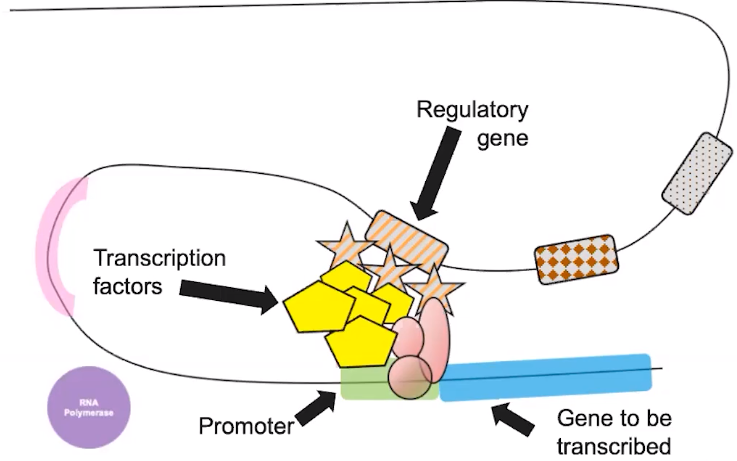
6.6 Gene Expression & Cell Specialization
Differences in the expression of genes account for phenotypic differences between organisms
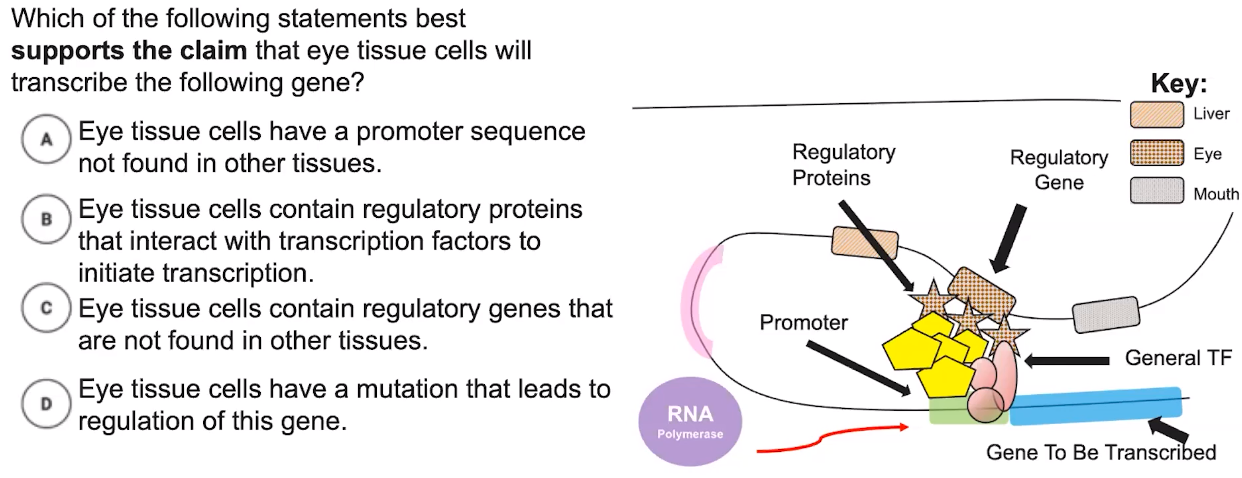
- Promoters are DNA sequences upstream of the transcription start site where RNA polymerase & transcription factors bind to initiate transcription.
- The interaction between regulatory proteins, regulatory genes, & transcription factors initiates transcription.
- The combination of genes expressed determines the phenotype of the organism
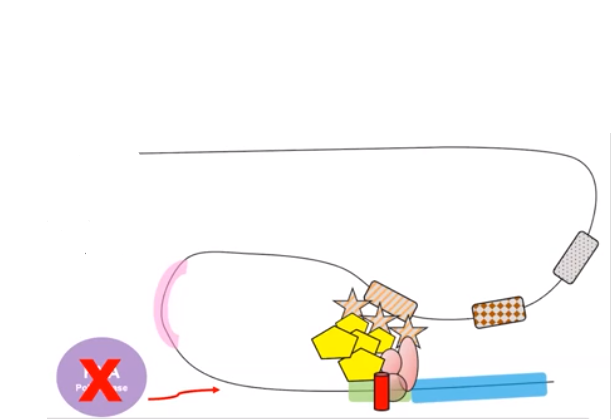
RNA polymerase may transcribe genes in the presence of transcription factors.
- Negative regulatory molecules inhibited gene expression by blocking transcription.
- RNA polymerase is blocked from binding to the promoter.
Gene regulation results in differential gene expression & influences cell products & function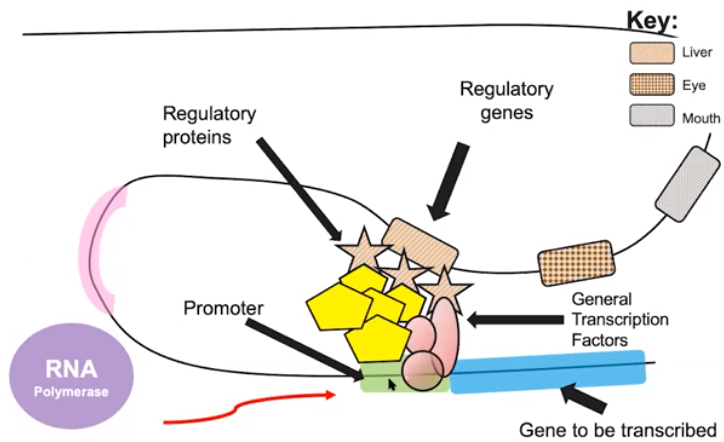
- Cells in from the same organism have the same DNA
- Only certain tissues have activator proteins that turn on the regulatory genes.
- The phenotype of the cell or organism is determined by the combination of genes that are expressed.
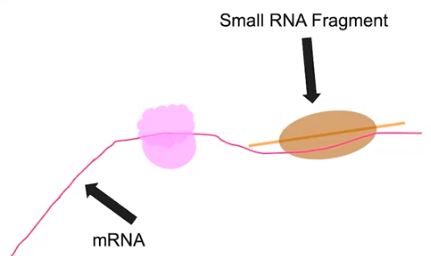
Certain small molecules have roles in regulating gene expression
- Small RNA fragments can have roles in gene expression
- Are coded by DNA
- Can break down mRNA in the cytoplasm
- Can block transcription
Ex9.
6.7 Mutations
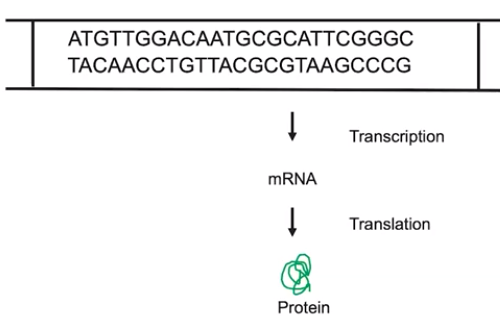
The normal function of the genes & gene products collectively comprises the normal function of organisms.
- When a gene is transcribed & translated, proteins are produced
- The function & amount of gene products determine the phenotype of organisms
Changed in genotype can result in changes in phenotype
- Mutations are changes in the genome of an organism
- DNA mutations can be positive, negative or neutral based on the effect or the lack of effect they have on the resulting nucleic acid or protein & the phenotypes that are conferred by the protein.
- Whether a mutation is detrimental, beneficial, or neutral depends on the environmental context.
- Distributions in genes & gene products can result in new phenotypes
- Mutations are the primary source of genetic variation
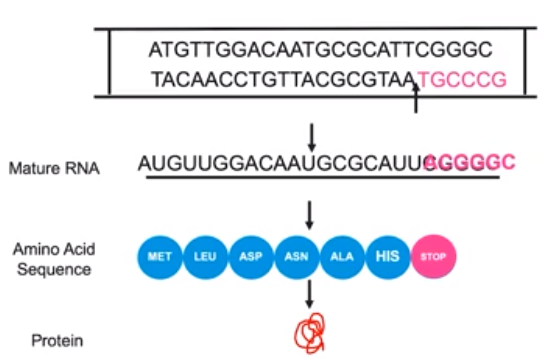
Alterations in a DNA sequence can lead to change in the type or amount of the protein produced.
- Changes in genotype can result in changed in phenotype through base insertions & deletions
- DNA nucleotides can be deleted from a sequence
- Deletion & insertions shift the order of the gene sequence.
- Can lead to no protein product or additional protein products
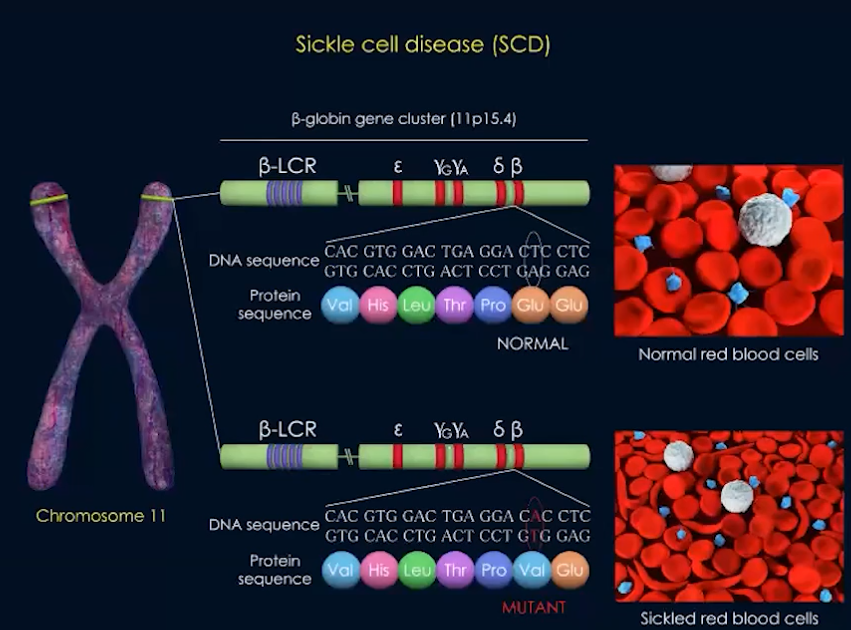
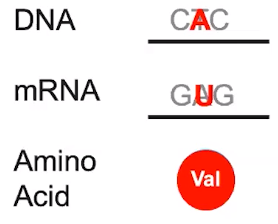
Sickle cell disease results from a change in base sequences
- One base change leads to change in mRNA & ultimately a difference in the sequence of amino acids
- Individuals with this mutated genotype have a different phenotype & their blood may be sickled
- The environment determines whether this phenotype is detrimental, beneficial, or neutral.

Ex. 10
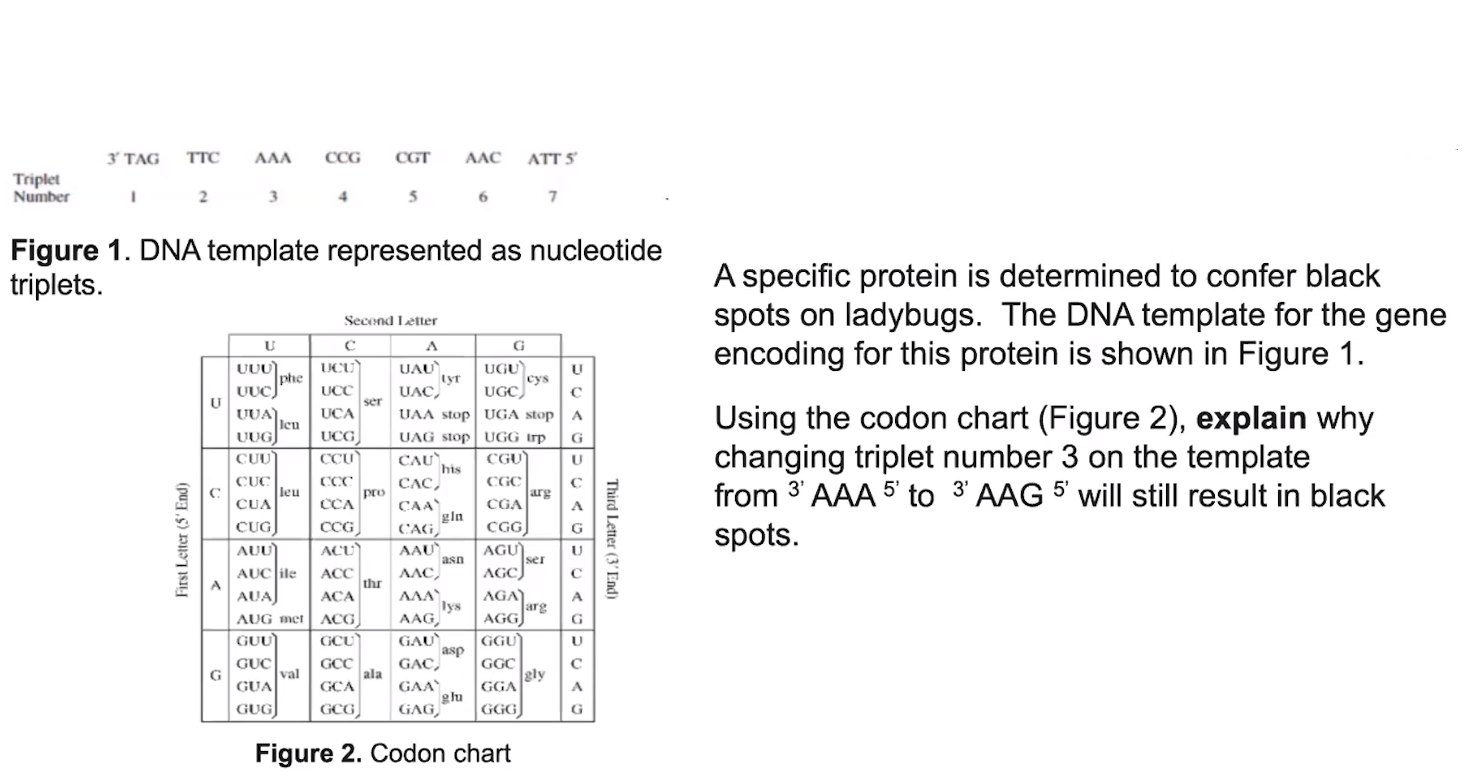

The processing of genetic information is imperfect & is a source of genetic variation.
- Distributions in genes & gene products can cause new phenotypes
- Random mutations can be caused by each of the following:
- Errors in DNA replication
- Errors in DNA repair
- Radiation
- Reactive chemicals
- Mutations are primary source of genetic variation
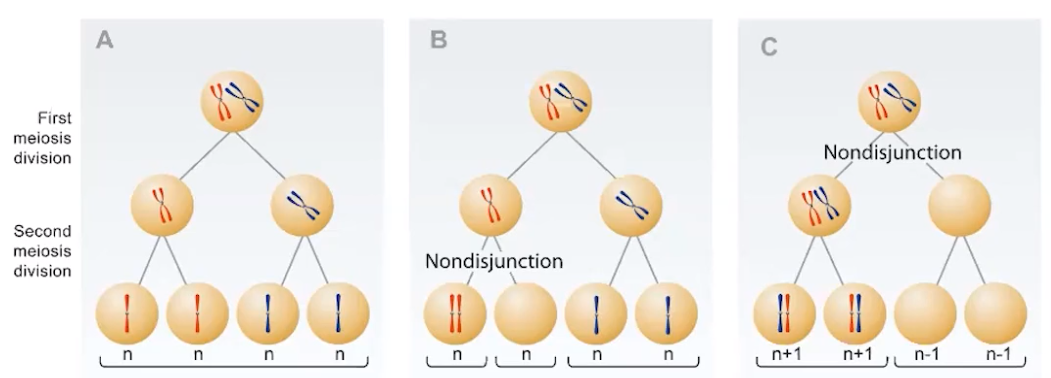
Changes in chromosome numbers often result in new phenotypes.
- Errors in mitosis or meiosis can result in changes in phenotype.
- Chromosomes may not separate properly during cell division
- Triploidy - having three copies of a particular chromosome
- Can result in sterility in some animals
- Polyploidy - having multiple sets of homologous chromosomes.
- Can result in increased vigor in plants
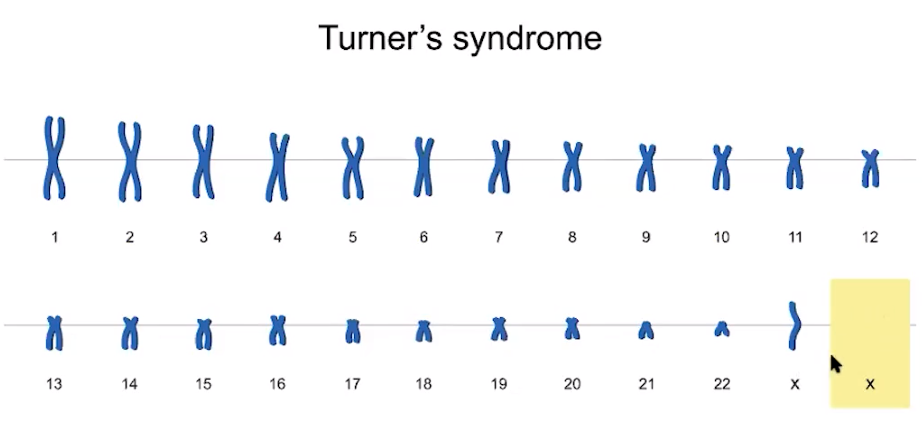
Changes in chromosome number often result in human disorders with developmental limitations.
- Improper separation of chromosomes can lead to gametes with missing or extra chromosomes.
- Trisomy 21 (47XX +21) results when you have an extra chromosome at chromosome 21.
- Turner syndrome (45XO) results when you have a missing x chromosome
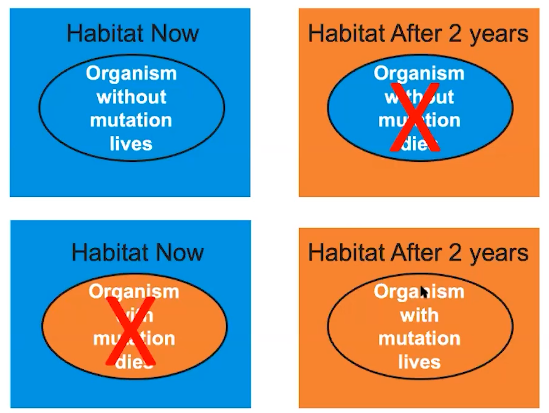
Whether a mutation is detrimental, beneficial or neutral depends on the environmental context
- The environment is not static
- Changes in the environment can impact survival of organisms
- A particular phenotype can be beneficial under certain environmental conditions & detrimental under other conditions
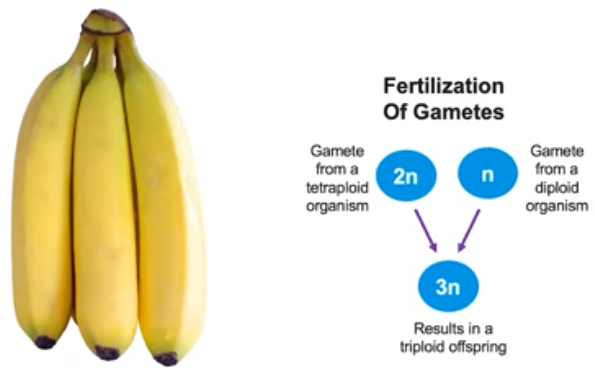
Polyploidy in plants - sterility
- Polyploidy can also result in sterility
- Cavendish bananas are triploids (tetraploid mated with diploid)
- These polyploids are infertile.
- Can’t reproduce or make seeds.

Polyploidy in plants - Increased Vigor
- In plants this often results in increased vigor
- These strawberries are phenotypically larger
than normal
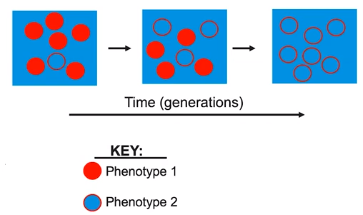
Changes in genotype may affect phenotypes that are subject to natural selection
- Natural Selection is the process in which organisms better adapted to a particular environment have a higher chance of survival & reproduction, thereby passing on those adaptations to the next generation.
- Genetic changes that enhance survival & reproduction can be selected for by environmental conditions.
The horizontal acquisition of genetic information in prokaryotes increases variation.
- Primarily occurs in prokaryotes.
- Contributes to genetic variation in prokaryotic populations
- Includes transformation, transduction, conjugation, & transposition
- Transformation refers to the uptake of naked DNA
- Naked DNA - DNA not protected by proteins or other molecules
- DNA comes from an external environmental source
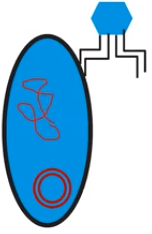
- Primary occurs in prokaryotes
- Transduction is the transmission of foreign DNA into a cell when a viral genome integrates with the host genome
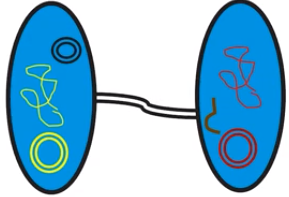
- Conjugation is a cell-to-cell transfer of DNA
- Allows for small segments of DNA to move from one cell to another
- An external cell extension connects the cell & allows transfer
- Primarily occurs in prokaryotes
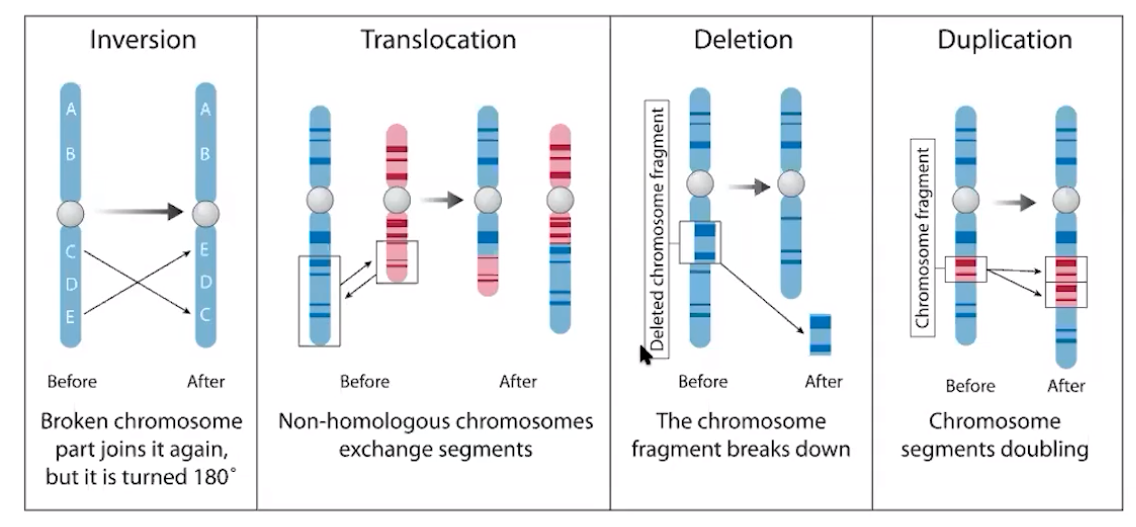
- Transposition is the movement of DNA segments within & between DNA molecules
- Conjugation is a cell-to-cell transfer of DNA
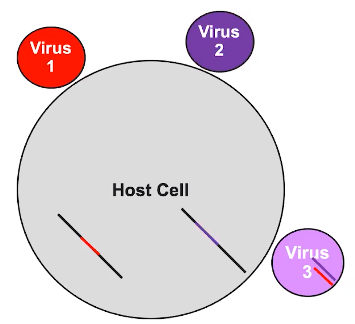
Related viruses can combine genetic information
- More than one virus can infect the same cell
- Viral genomes can combine once integrated in host genome
- New viruses can have the new viral combinations
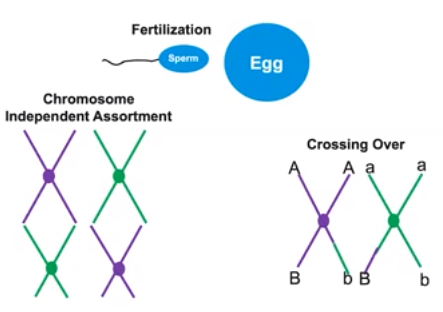
Reproduction process can increase genetic variation
- Certain reproductive processes are evolutionarily conserved & are shared by various organisms
- Sexual reproduction & the combination of random gametes
- Independent assortment of homologous pairs
- Crossing over during meiosis
Ex11.

6.8 Biotechnology
Genetic engineering techniques can be used to analyze & manipulate DNA & RNA
- Common techniques used to analyze DNA & RNA include the following:
- Electrophoresis
- Polymerase chain reaction (PCR)
- DNA sequencing
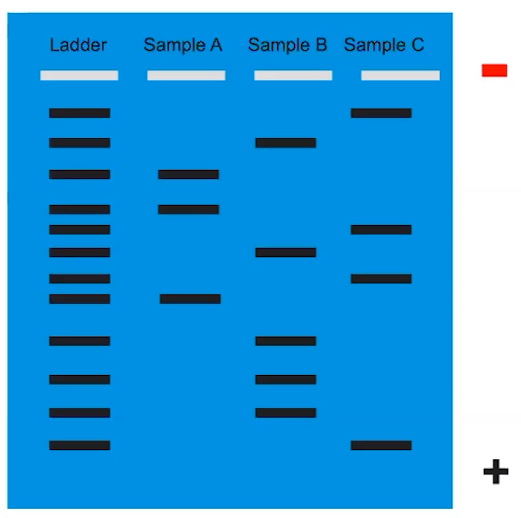
Gel electrophoresis separates molecules according to size & charge
- DNA molecules are negatively charged.
- DNA samples containing fragments are loaded into wells filled with a gel
- Current applied to gel creating positive & negative ends
- DNA moves toward the positive end of the gel
- This separates DNA fragments by size
- Smaller particles will move faster & further through the gel
- Bands indicate molecules that migrated to the location in the gel.
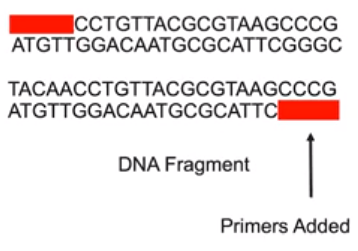
DNA polymerase chain reaction (PCR), DNA fragments are amplified
- PCR techniques allow scientists to create large samples of DNA to analyze when small samples are initially available
- Technique involves several steps.
- DNA is denatured (by heating)
- Primers are added.
- DNA is replicated
- Process is repeated until an ample amount of DNA is obtained
- Technique involves several steps.
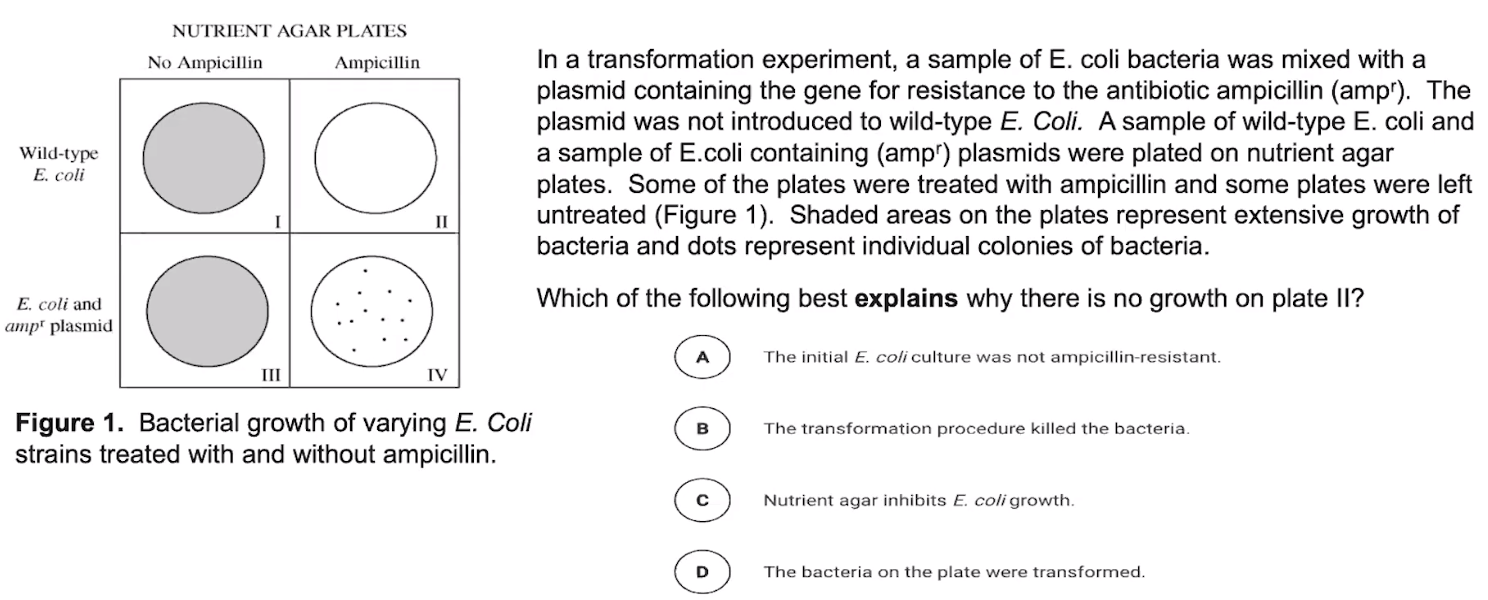
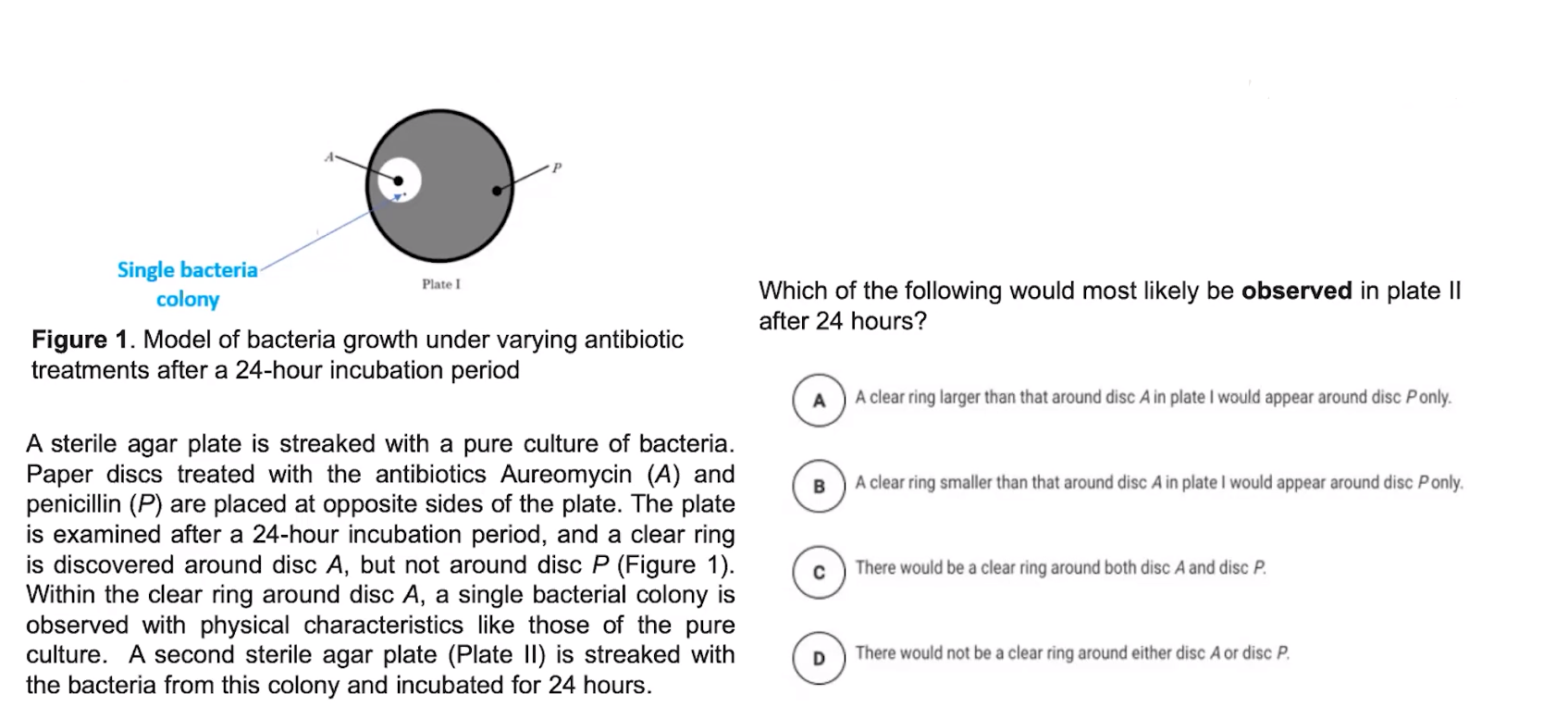
Bacterial transformation introduces DNA into bacterial cells.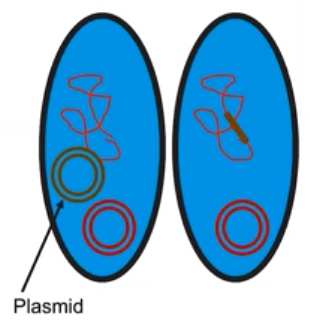
- Transformation is the process by which bacterial cells take up foreign DNA from external sources
- Bacterial cells only uptake foreign DNA at specific times
- This DNA can be incorporated into the bacterial chromosome or exist separately in the cytosol (plasmid)
- Scientists introduced foreign DNA during the time bacterial cells are primed for uptake.
- Used to make medicines, modify food, or amplify DNA
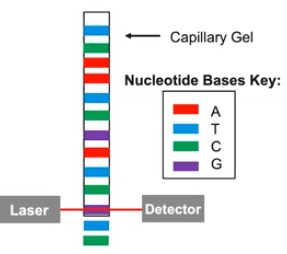
DNA sequencing determines the order of nucleotides in a DNA molecule.
- DNA sequencing techniques establish the order of nucleotides in a strand of DNA molecules
- There are numerous techniques employed to sequence DNA, which include:
- Nucleotides are labeled with fluorescent dye to read & build copies of DNA
- DNA runs through the capillary gel, a detector reads the sequence.
- Nucleotides are labeled with fluorescent dye to read & build copies of DNA
- DNA sequencing is important in evolutionary biology, medicine, & forensics
Ex12.
