The Complement System
Innate vs Adaptive Immunity
Characteristic | Innate | Adaptive |
Source | Its receptors and mechanisms are encoded in germ-line DNA | Recombination of genes (Somatic recombination) |
Inheritance | Inherited | Non-inherited |
Specificity | Recognition of conserved molecular patterns like PAMPs and DAMPs | Recognition of dynamically changing molecular patterns |
Memory | Epigenetic memory of past pathogen encounters | Survival of clones of cells expressing specific receptors to past antigens |
Structure | Physical barriers, chemical defenses, cellular responses | Cellular responses, antibody production |
Dynamics | Present in all individuals at all times | Formed as antigens are encountered |
Antibodies
Antibodies: Soluble polypeptide molecules manufactured and secreted by B cells and plasma cells that can recognize any substance. They have neutralizing or cytotoxic activities (cell or complemented-mediated)
Antibodies participate in both innate and adaptive immunity
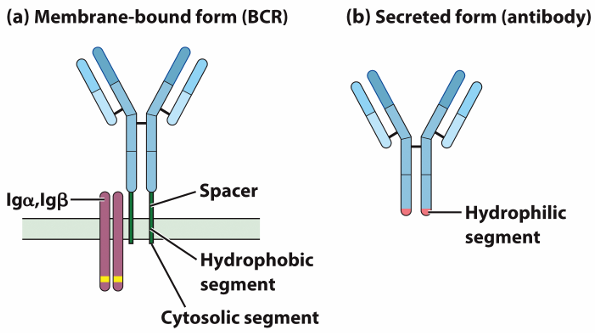
Structure of Antibodies and B cell receptors:
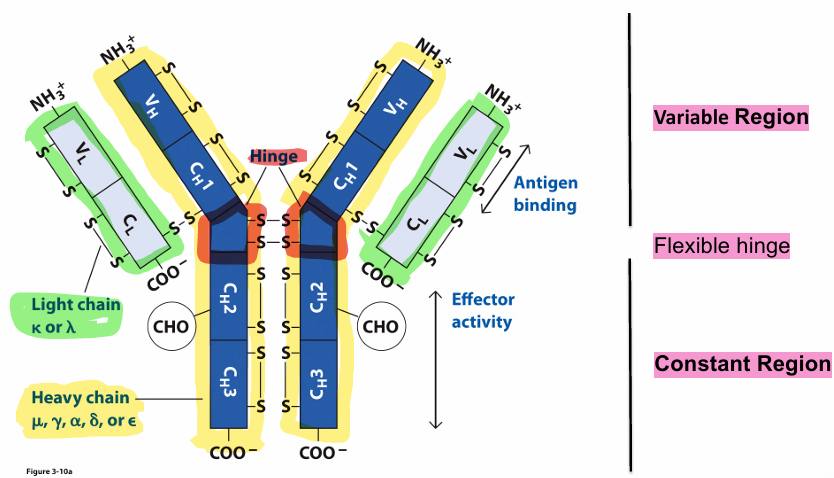
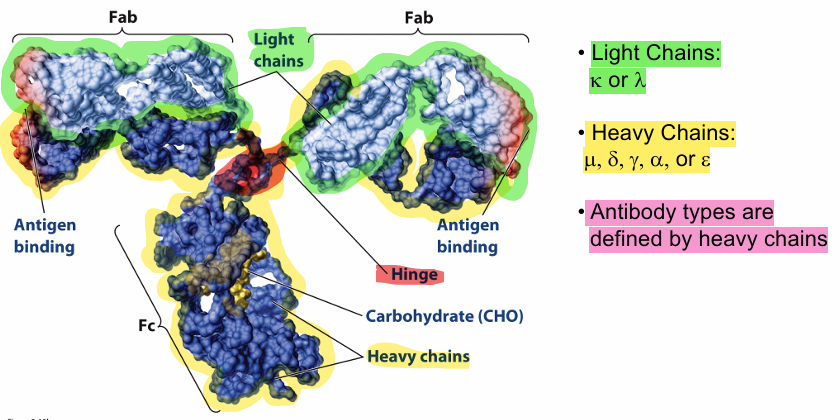
Antibody molecules can be cleaved into functionally distinct fragments
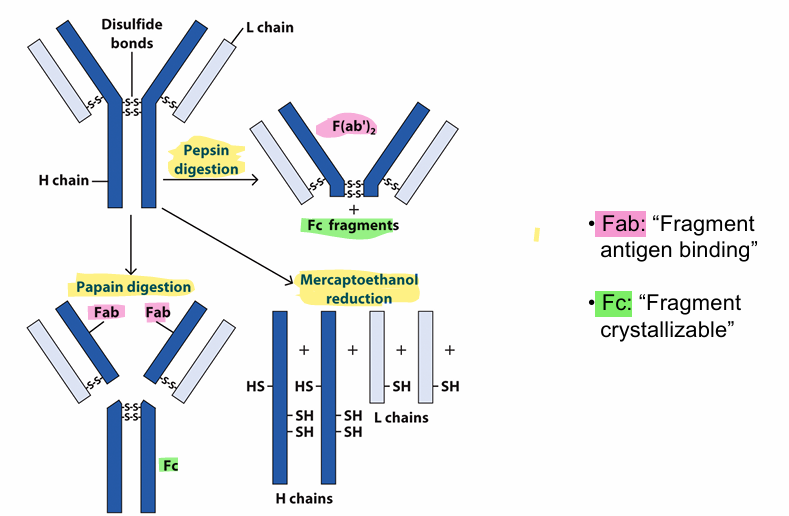
Pepsin Digestion: The enzyme pepsin cleaves below the hinge region of the antibody. It generates a F(ab’)2 fragment (Fab arms still connected via disulfide bonds) and destroys Fc
Papain Digestion: cleaves above the hinge region. It generates two separate Fab fragments and one intact Fc fragment.
Mercaptoethanol Reduction: This reducing agent breaks disulfide bonds. It separates all the polypeptide chains
Six Effector Mechanisms of Antibodies
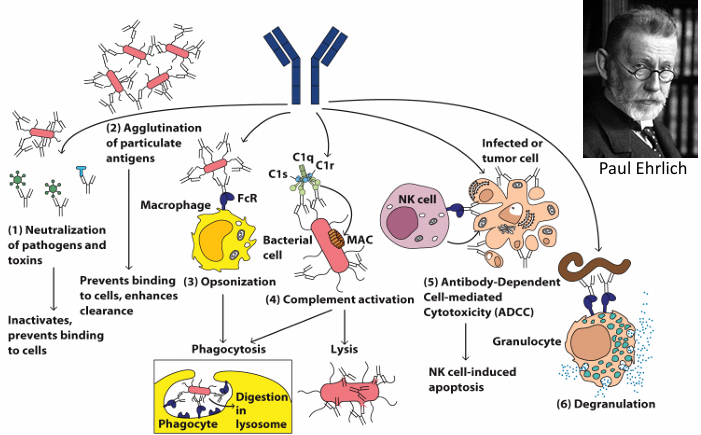
Neutralization of pathogens and toxins: Antibodies bind to pathogens inactivating them, preventing them from binding to cells.
Agglutination of particulate antigens: Prevents binding to cells, enhances clearance. Antibodies clump pathogens together making them easier to clear
Opsonization: Antibodies coat the surface of pathogens marking them for phagocytosis.
Complement activation: Antibodies trigger the classical complement pathway, leading to pathogen lysis, inflammation. etc.
Antibody-Dependent-Cell mediated Cytotoxicity (ADCC): Antibodies bound to infected cells interact with Fc receptors on NK cells which induce apoptosis
Degranulation: Antibodies bind to pathogens and trigger the release pre-stored granules which contain histamine, cytokines and enzymes
The complement system
The complement system: Components of plasma that complement antibody action to kill target cells by lysis. They are around 50 proteins that interact in a network as well as with other systems.
Complement functions include:
Alternative pathway: Directly kill cells
Classical pathway: Interact with antibodies to kill cells
Lectin pathway: Interact with lectins to kill cells
Opsonize pathogens to flag them for phagocytosis
Function as a chemoattractant to recruit leukocytes and induce inflammation
Immunoglobins and whether they activate the complement pathway:
Immunoglobin | Activates classical component pathway? |
IgG1 | YES |
IgG2 | YES AND NO |
IgG3 | YES |
IgG4 | NO |
IgA1 | NO |
IgA2 | NO |
IgM | YES |
IgE | NO |
IgD | NO |
Nomenclature:
C: proteins that belong in the Classical pathway
#: name of the component in order of discovery
b: designates the larger fragment with enzyme activity
a: designates the smaller fragment
Classical Pathway
Trigger: Initiated by the pathogen sensor C1 complex binding to pathogen bound antibodies like IgG or IgM
Goal: To form the membrane attack complex (MAC) and kill pathogens through lysis or opsonization
Steps of the Classical Pathway:
Initiation by C1 complex:
C1 Complex components:
C1q: 18 polypeptide chains in 6 collage like triple helices
C1r and C1s: 2 molecules each
Tails: Interact with 2 C1r and 2 C1s
Heads: Interact with IgM or IgG
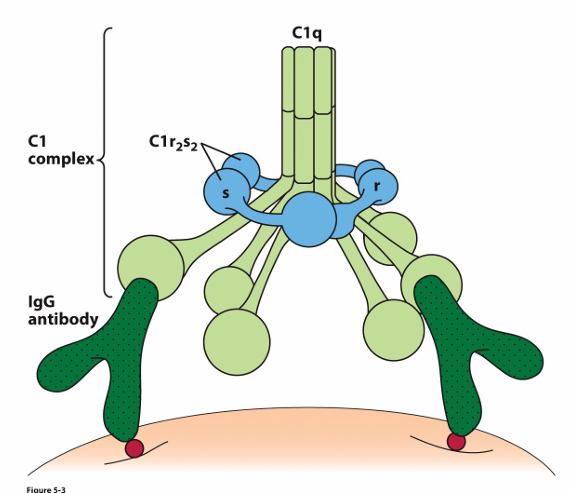
C1q Activation:
C1q binds to antigen-bound antibody, ( IgM (staple form) or IgG on pathogen surfaces).
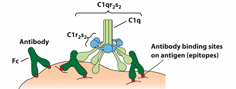
This induces a conformational change activating C1r, which then activates the second C1r and the two C1s molecules ….(C1r → 2nd C1r and 2 C1s)
Activation of C4 and C2:
C1s Activity:
Cleaves C4 into C4a (small) and C4b (larger)
C4b binds to pathogen surface
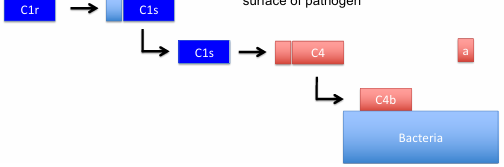
C2 binding and Cleavage:
C2 binds to C4b and is broken by C1s into C2a (larger) and C2b (smaller and not on the bacteria anymore)
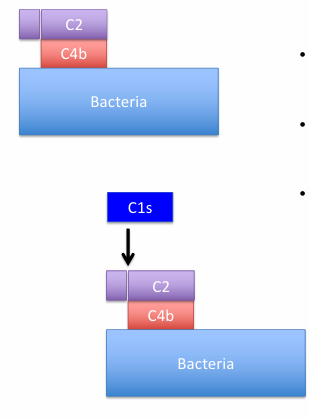
C4b2a complex forms and acts as the C3 convertase
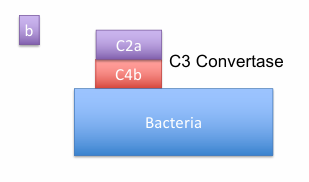
Formation of C3 Convertase:
C4b2a Complex
breaks C3 into C3a (small fragment, chemoattractant, not on the complex) and C3b (larger fragment, in the complex)
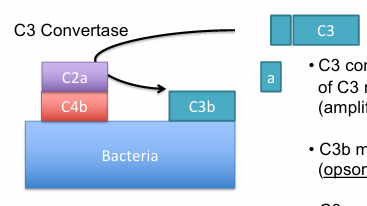
C3b coats the pathogen surface for opsonization and amplifies the complement cascade
Formation of C5 Convertase:
C4b2a3b Complex:
Formed when one C3b molecule binds to C4b2a
Functions as the C5 Convertase
C5 cleavage:
Breaks C5 into C5a and C5b ?????
Diagram/Flow (for Quick Review)
C1 Activation: C1q binds IgM/IgG → Activates C1r → Activates C1s.
C4 Cleavage: C1s breaks C4 → C4b binds pathogen.
C2 Cleavage: C4b binds C2 → C2 cleaved to form C4b2a (C3 convertase).
C3 Cleavage: C3 convertase cleaves C3 → C3b coats pathogen; amplifies cascade.
C5 Convertase: C4b2a3b cleaves C5 → Leads to MAC formation.
Lectin Pathway
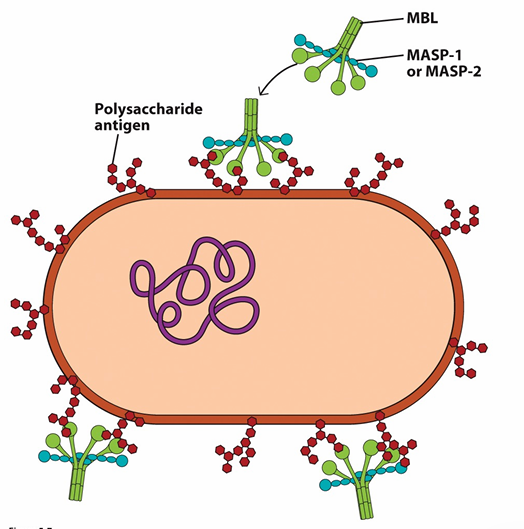
Trigger: MBL or ficolins bind to polysaccharide antigen on the pathogens surface
Goal:
Steps:??
MBL/ficolin binds pathogen surface.
Activated MASPs cleave C4 and C2.
Formation of C3 convertase (C4b2a) → Cleaves C3 into C3a and C3b.
C3b opsonizes pathogens and continues the complement cascade.
Alternative pathway
Trigger: Spontaneous cleavage of C3 into C3a and C3b. Requires no antibodies, lectins or pathogen-bound triggers.
Goal: form MAC complex
Steps of Alternative Pathway:
Spontaneous Cleavage
C3 undergoes spontaneous hydrolysis to from C3b.?
Formation of C3 Convertase
C3b binds Factor B, which is cleaved by Factor D to form C3bBb (a fluid phase C3 convertase)
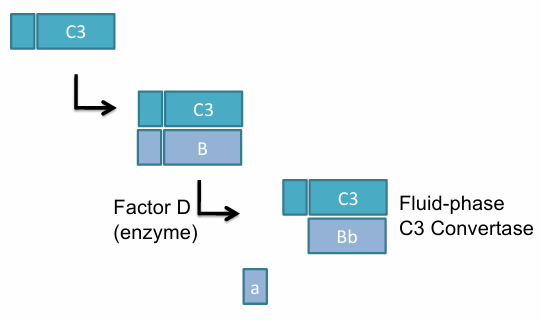
C3 is broken down into C3b and C3a(leaves) and C3b attaches the pathogen
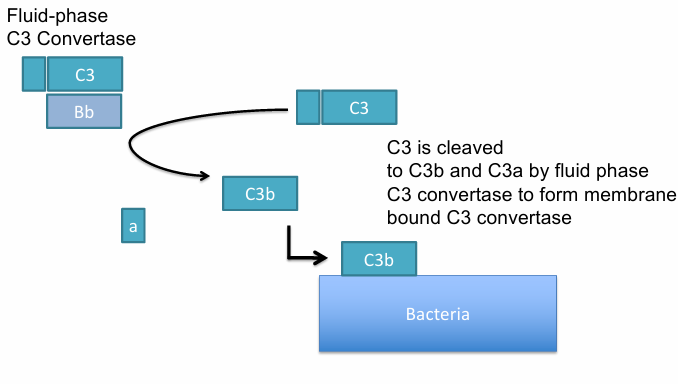
Stabilization:
Properdin stabilizes the C3bBb complex on microbial surfaces
C5 Convertase formation:
Addition to another B forms C3b, making C3bBbC3b, which acts as a C5 convertase. Required Factor D y Properdin
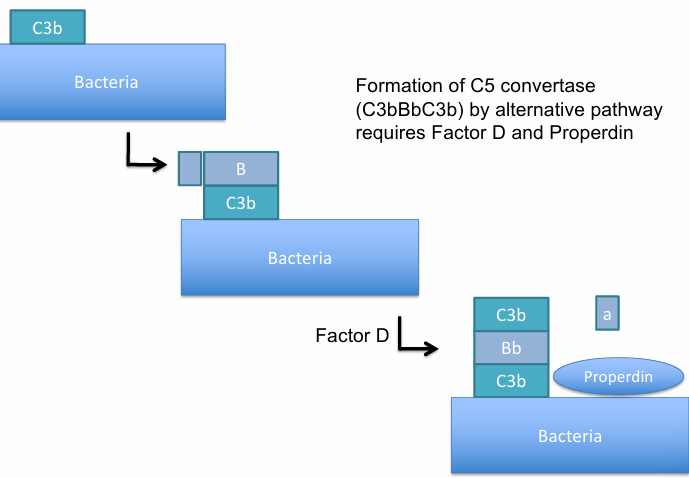
C5 cleavage:
C5 convertase cleaves C5 breaks into C5a and C5b → Leads to MAC formation.
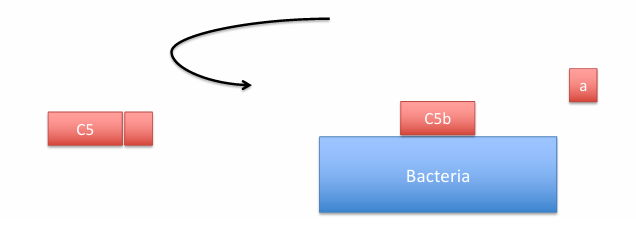
*C3a and C5a are potent inflammatory signals
Formation of the Membrane attack complex (MAC)
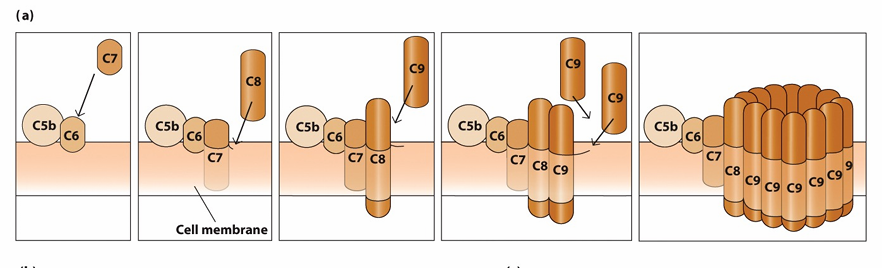
Regulation of Complement
Mechanism to prevent complement attack on self-cells:
Unstable complement enzymes
C1 inhibitor (C1NH): Dissociates C1q from C1s/C1r
Decay-accelerating factor (DAF) and complement receptor 1 (CR1): Dissociate C3 convertases
Genetic deficiencies in complement
C3 Deficiency: Susceptibility to bacterial infections
C5-C9 Deficiency: Recurrent infections with Neisseria species
C1q or C4 Deficiency: Linked to systemic lupus erythematosus (SLE)