Translation II – Ribosome Structure & Translation Initiation (BCH5413) from transcripts
Introduction & Lecture Context
Instructor: Prof. Bert Flanegan, Department of Biochemistry & Molecular Biology (Course BCH5413).
Lecture theme: Translation III – Ribosome Structure & Translation Initiation.
This lecture critically builds on previous foundational material, specifically covering messenger-RNA (mRNA) stability and the intricate amino-acyl (charging) reactions of tRNAs, which are essential for delivering the correct amino acids.
The third and final “player” in the complex process of protein synthesis, ribosomes, are introduced in detail today, highlighting their central role as the machinery for protein synthesis.
Unit learning objectives:
Recognise and distinguish ribosome architecture between bacteria (prokaryotes) and eukaryotes, noting their evolutionary conservation and increasing complexity.
Understand the fundamental ribosomal catalytic function, particularly the peptidyl-transferase activity, and recognize that this activity resides within ribosomal RNA (rRNA), making the ribosome a catalytic RNA or ribozyme.
Trace all mechanistic steps that meticulously bring a ribosome to the correct start codon on a mature eukaryotic mRNA, from initial cap recognition to the formation of the full initiation complex.
Appreciate the crucial sequence context rules, particularly the Kozak consensus sequence, and understand variations in initiation such as leaky scanning and their functional consequences.
Ribosome Structure & Localisation

General role:
Serves as the crucial scaffold for the sequential and precise binding of charged tRNAs to their complementary mRNA codons, ensuring the correct amino acid sequence.
Acts as the core enzyme for stepwise peptide-bond formation, with rRNA performing the key catalytic function (peptidyl-transferase activity) ➔ thus, the ribosome is a catalytic RNA/ribozyme.
Protein elongation is a rapid process, occurring at an approximate rate of , demonstrating the efficiency of this cellular machinery.
Cytoplasmic Distribution (Slide 3: EM image illustration)
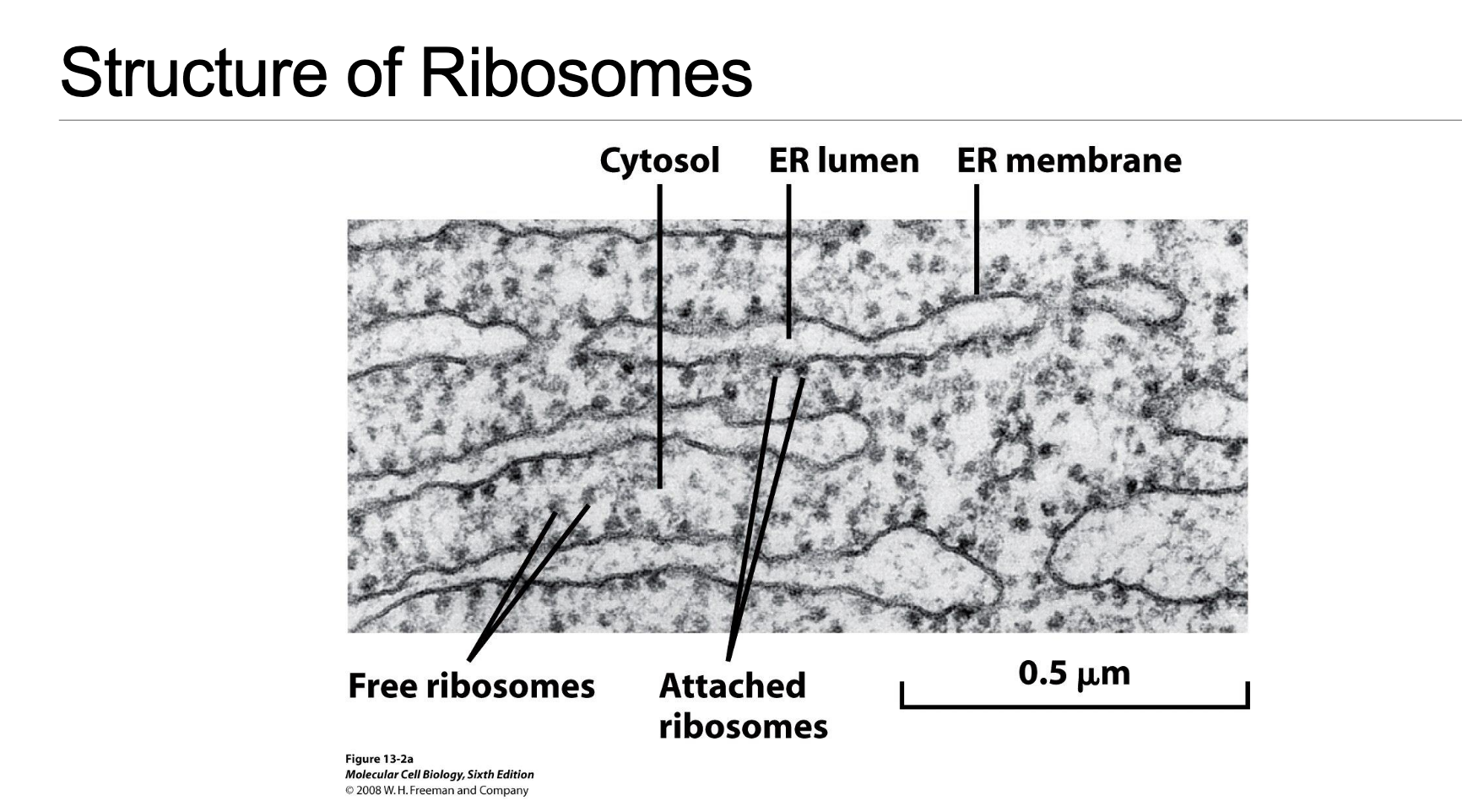
Rough Endoplasmic Reticulum (rER):
The endoplasmic reticulum appears “rough” due to the dense presence of ribosomes studding its outer membrane, giving it its characteristic appearance.
This ribosome–membrane interaction is crucial, forming a specialized translocon channel that allows nascent peptide chains to enter the ER lumen co-translationally. This entry is a precursor step for proteins destined for secretion outside the cell or for integration into cellular membranes.
Free cytosolic ribosomes:
These ribosomes float freely within the cytosol, unattached to membranes.
They primarily translate proteins destined for cytosolic functions or for import into various organelles (e.g., mitochondria, nucleus, peroxisomes) within the cell, as opposed to those secreted or incorporated into membranes.
Bacterial (Prokaryotic) Ribosome (Slide 4)
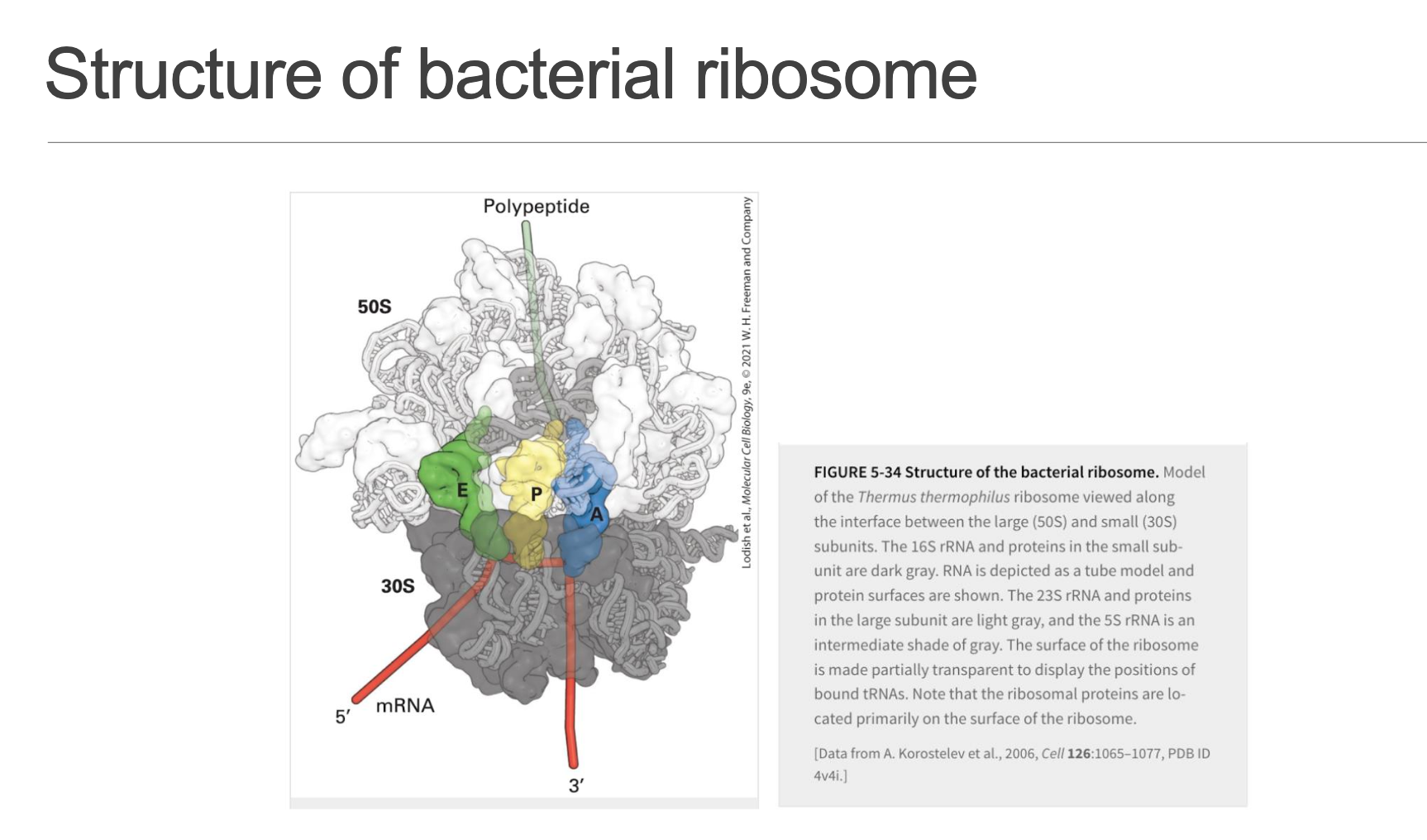
Composed of two unequal subunits, distinguished by their sedimentation coefficients (Svedberg units), which reflect their size and shape:
Large subunit:
Small subunit:
These combine to form a complete functional ribosome.
The mRNA path runs precisely through the inter-subunit cleft, providing a protected and efficient track for translation. It features three crucial tRNA binding sites:
A (Amino-acyl) site: The primary entry point for all incoming charged tRNAs, each carrying its specific amino acid, matching the mRNA codon.
P (Peptidyl) site: Occupied by the tRNA holding the nascent polypeptide chain, facilitating peptide bond formation.
E (Exit) site: The departure lounge for de-acylated tRNAs (tRNAs that have released their amino acid), which then leave the ribosome.
The peptidyl transferase centre, responsible for catalyzing peptide bond formation, resides exclusively in the rRNA of the large subunit (23S rRNA), a classic and compelling example of a ribozyme.
Eukaryotic Ribosome (Slide 5)
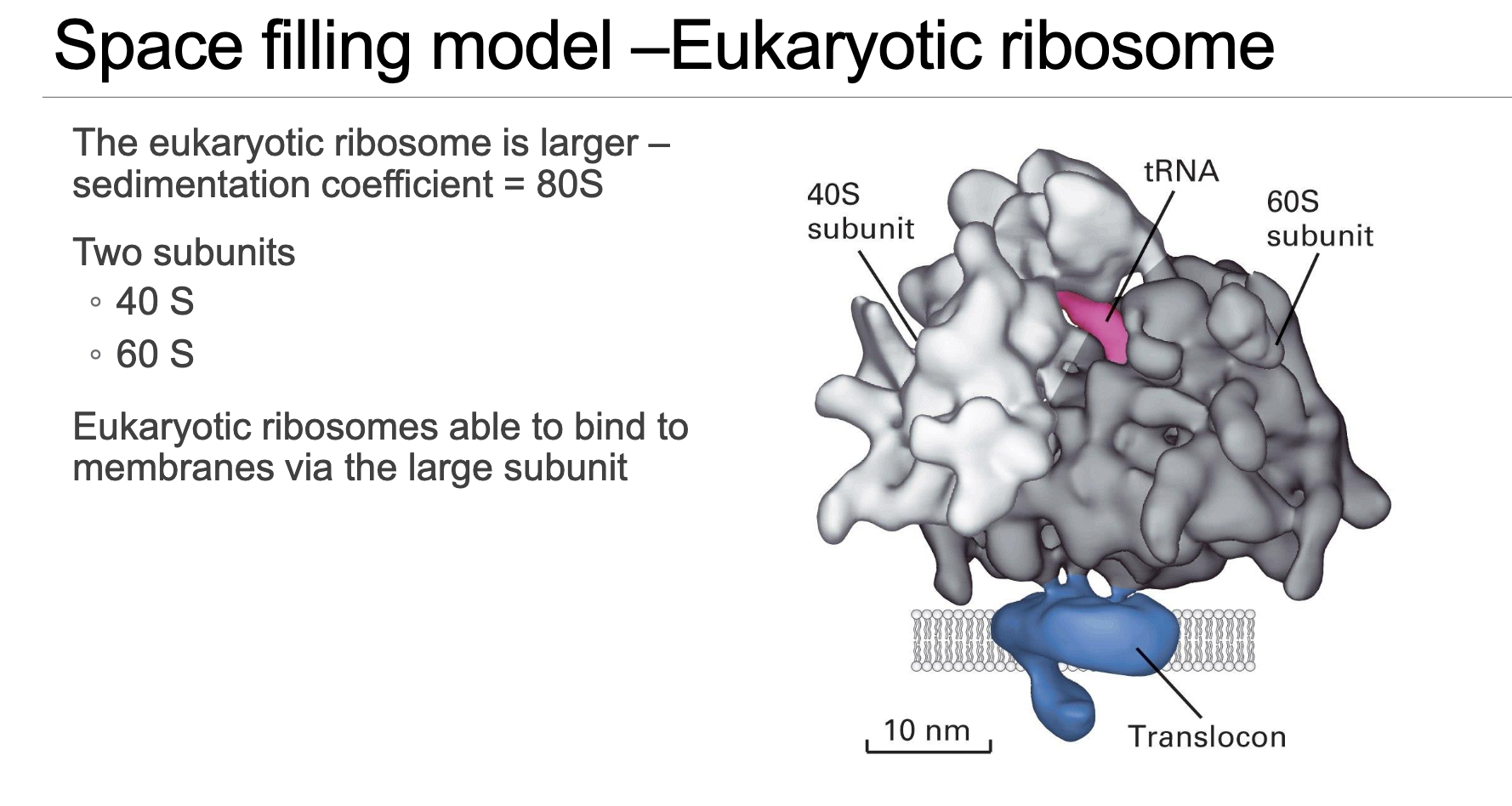
Larger in overall size compared to bacterial ribosomes but maintains a remarkably conserved overall architecture, reflecting their common evolutionary origin and fundamental function.
Sediments at as a complete functional unit, composed of:
40S small subunit.
60S large subunit.
The 60S large subunit specifically provides the critical interaction interface with the ER translocon, explaining the precise physical docking mechanism of rER-bound ribosomes that are synthesizing secreted or membrane proteins.
rRNA & Protein Composition (Human example) (Slide 6)
40S subunit: Comprises 18S rRNA + 33 distinct proteins.
60S subunit: Comprises 5S, 5.8S, 28S rRNAs + 50 distinct proteins.
Despite a numerically fewer species of RNA molecules compared to proteins, the mass of rRNA significantly exceeds the mass of ribosomal proteins within each subunit, underscoring the RNA’s structural and catalytic importance.
Evolutionary Conservation (Slide 7)
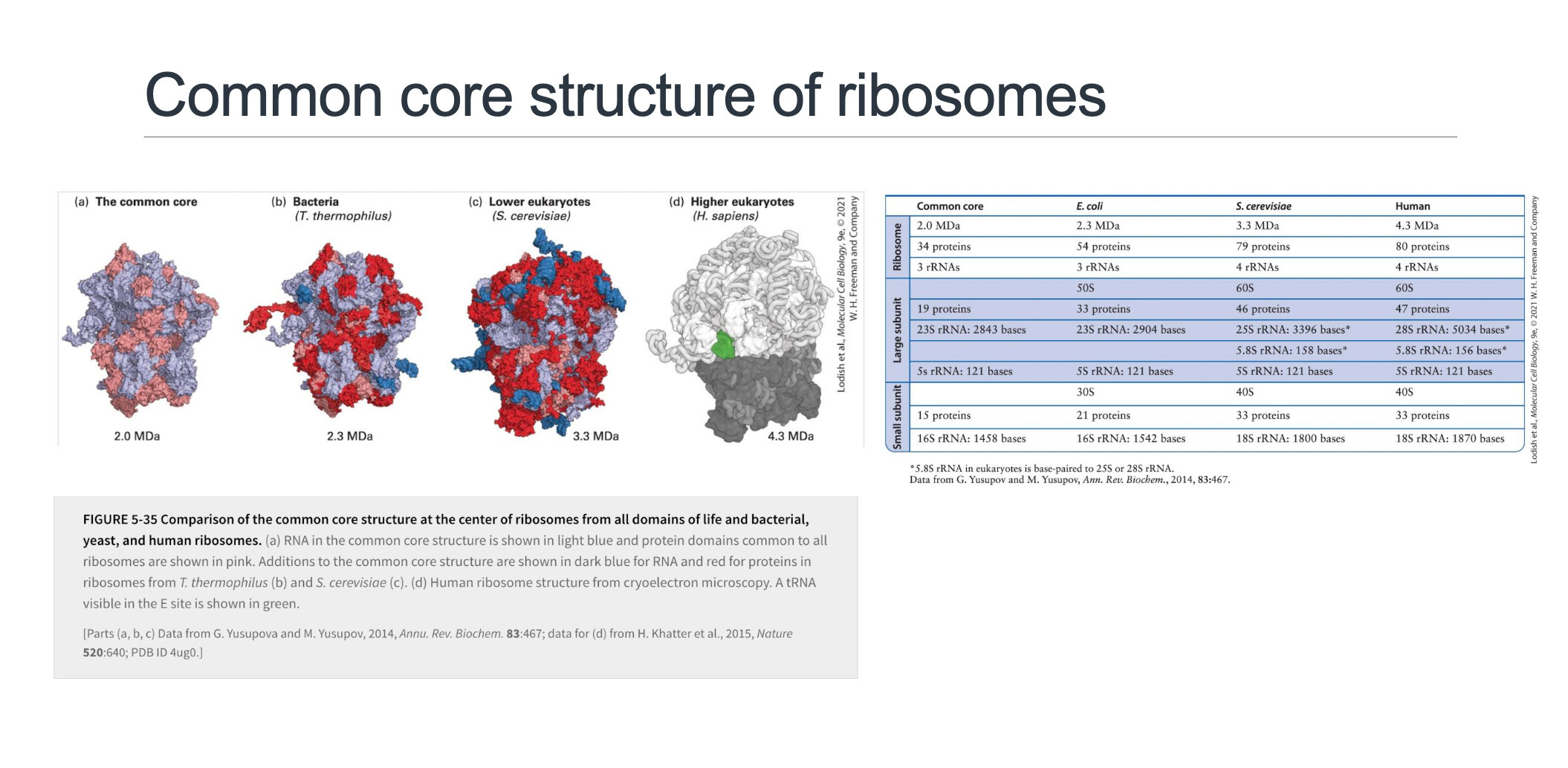
High-resolution crystal and cryo-EM reconstructions of ribosomes from diverse organisms consistently reveal a common 2 MDa RNA-protein core that is remarkably conserved across all three domains of life (Bacteria, Archaea, and Eukarya). This core represents the minimal functional unit required for translation.
Beyond this core, there is a progressive accretion of extra rRNA segments (often depicted in dark blue) and additional proteins (dark red) as organisms evolve, leading to increasingly larger and more complex ribosomes:
Core: .
Bacteria (e.g., E. coli): ; with 54 proteins and approximately rRNA nucleotides.
Yeast (a model lower eukaryote): ; incorporating 79 proteins and approximately rRNA nucleotides.
Human: The largest and most complex, at ; containing 80 proteins and approximately rRNA nucleotides.
Functional takeaway: The conserved core ensures the same fundamental mechanistic process of translation. The evolutionarily added extra elements primarily serve to fine-tune regulation, control localisation within the cell, and facilitate interaction with the increasing complexity of mRNA features in higher organisms.
Overview of Eukaryotic mRNA Features Relevant to Initiation (Slide 8)
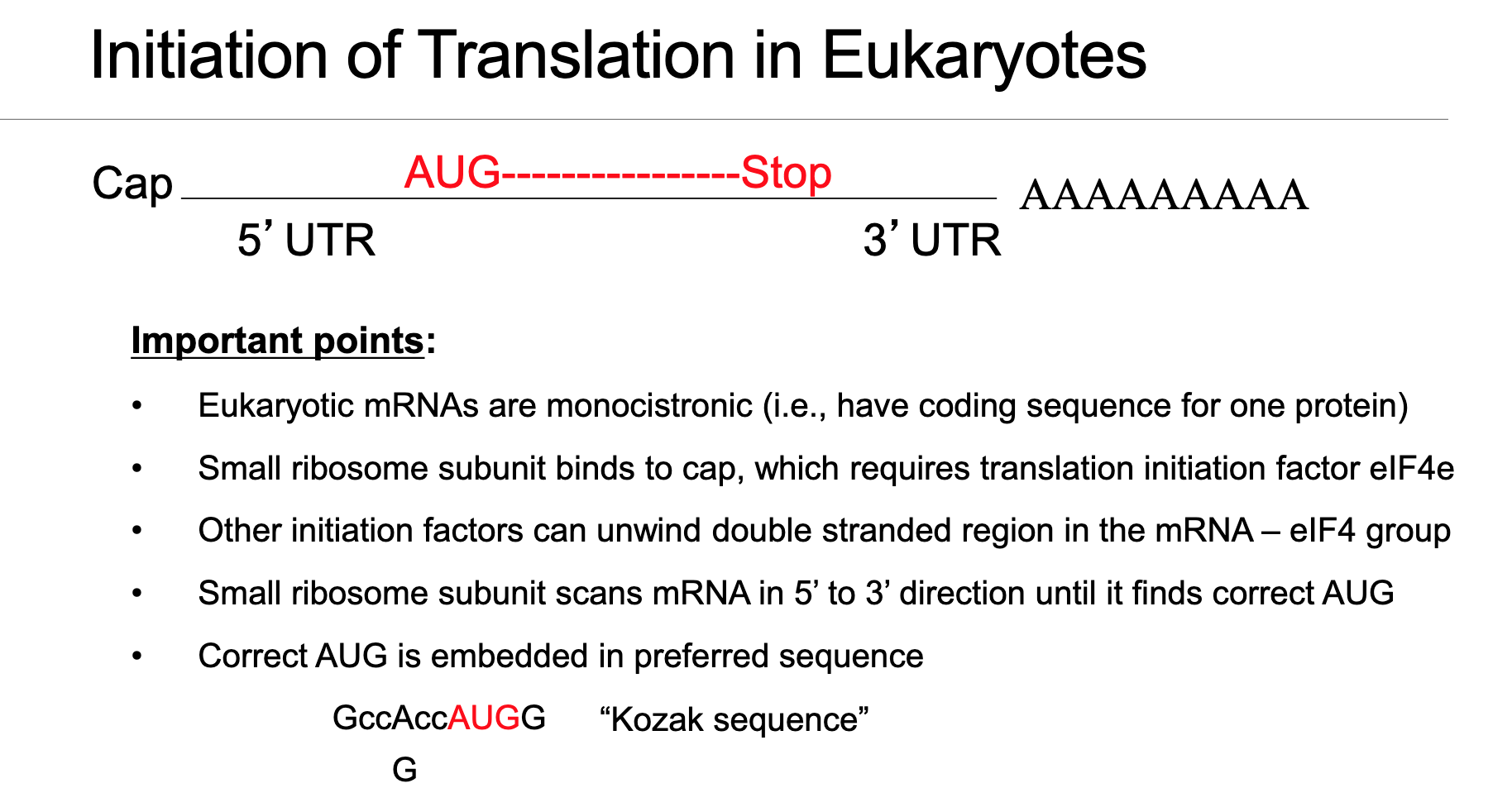

Eukaryotic mRNA is highly structured with several key elements arranged in a 5’→3’ direction, all of which play roles in efficient and regulated translation initiation:
7-methyl-guanosine cap (m⁷GpppN): Located at the 5’ end, it protects the mRNA from degradation and serves as a crucial recognition signal for the initiation machinery.
5’ UTR (untranslated region): A segment between the cap and the start codon, which can contain regulatory elements like secondary structures or upstream open reading frames.
Start codon AUG: The triplet nucleotide sequence (Adenine-Uracil-Guanine) that specifically defines the translation initiation site, coding for methionine.
Protein-coding sequence (CDS): The region that carries the genetic information to be translated into a polypeptide chain.
Stop codon (UAA/UAG/UGA): Signals the termination of translation.
3’ UTR: The untranslated region at the 3’ end, often containing regulatory elements for mRNA stability and translation control.
Poly(A) tail: A long chain of adenine nucleotides at the 3’ end, which enhances mRNA stability and recruitment of translation factors, primarily bound by PABPC (poly-A-binding protein, cytoplasmic).
Monocistronic design: Eukaryotic mRNAs typically follow a monocistronic design, meaning one CDS generally gives rise to one protein. This contrasts sharply with bacterial mRNAs which are often polycistronic, encoding multiple proteins from a single mRNA molecule.
Scanning Model of Translation Initiation (Slide 9-11)
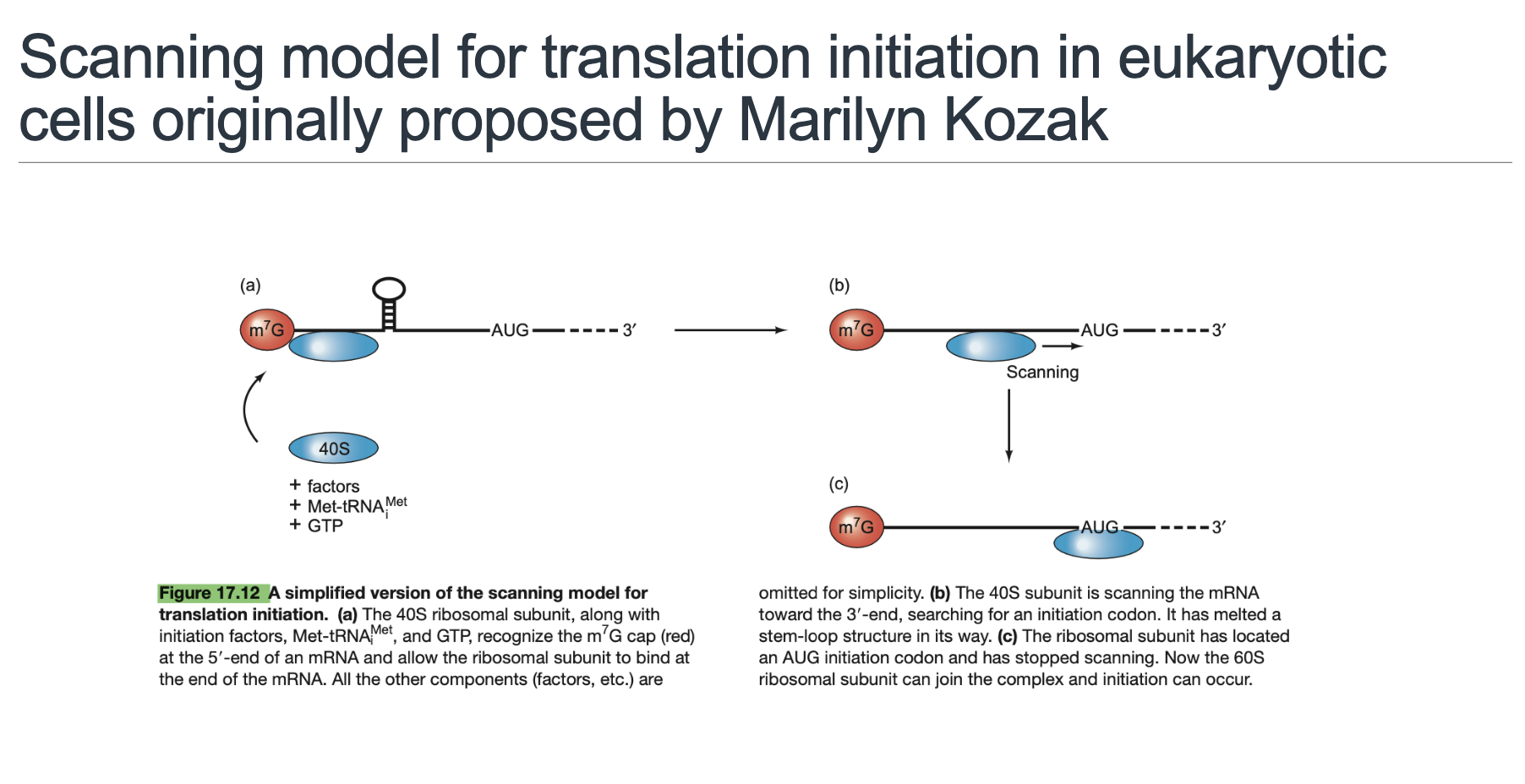
This crucial concept, fundamental to eukaryotic translation initiation, was established by Dr. Marilyn Kozak, after whom the Kozak sequence is named.
The stepwise narrative of how the ribosome finds the correct start codon involves a process of scanning. Here's a more detailed breakdown:
Cap Recognition (Slide 9): The process begins with the recognition of the 5’ m⁷G cap by the protein eIF4E. This protein forms a complex with the scaffold protein eIF4G, the RNA helicase eIF4A, and the RNA-binding enhancer eIF4B. This complex prepares the mRNA for ribosome binding.
40S Pre-Initiation Complex (43S) Assembly (Slide 10): Simultaneously, the 40S small ribosomal subunit pre-assembles with several initiation factors (eIF1, eIF1A, eIF3) and the initiator (bound to eIF2–GTP). This entire assembly forms the 43S pre-initiation complex.
mRNA Recruitment via Bridging (Slide 10): The large scaffold protein eIF4G acts as a crucial bridge, interacting with the cap-bound eIF4E complex at the 5’ end of the mRNA while also recruiting the 43S pre-initiation complex. This interaction brings the small ribosomal subunit to the 5’ end of the mRNA.
mRNA Circularisation (Slide 10): An important regulatory step, the PABPC (Poly-A-Binding Protein Cytoplasmic) bound to the poly(A) tail interacts with eIF4G (which is already bound to the cap region). This interaction brings the 5’ and 3’ ends of the mRNA into close proximity, effectively circularizing the mRNA. This circularization enhances the efficiency of re-initiation and ribosome recycling after one round of translation.
5’→3’ Scanning by 40S (Slide 11): The 40S pre-initiation complex, now loaded onto the 5’ end, begins to move linearly along the 5’ UTR in a 5’ to 3’ direction. This scanning process is powered by the ATP-dependent helicase activity of eIF4A/B, which unwinds any stable secondary structures (e.g., hairpins) in the 5’ UTR that could otherwise impede or halt scanning if their barriers are too stable.
Start-Codon Recognition and GTP Hydrolysis (Slide 11): Scanning continues until the 40S subunit encounters the start codon (AUG) in a favorable sequence context (Kozak consensus). This precise recognition triggers a conformational change that involves the GTP hydrolysis on eIF2, facilitated by eIF5. This hydrolysis releases eIF2-GDP and other associated eIFs from the 40S subunit, effectively halting the scanning process.
60S Joining and 80S Formation (Slide 11): Following eIFs dissociation, the 60S large ribosomal subunit is recruited and joins the 40S subunit. This crucial step is assisted by eIF5B–GTP. Upon 60S joining, the final GTP hydrolysis (on eIF5B) occurs, ejecting eIF5B and eIF1A. This completes the assembly of the functional 80S initiation complex, which is now perfectly poised for the elongation phase of protein synthesis.

Kozak Consensus (Slide 12)
The canonical strong context for eukaryotic start codon recognition is defined by the sequence: where R represents either Adenine (A) or Guanine (G). The parentheses around
gccindicate that these bases are commonly present but less critical for overall strength.Critical positions for strong initiation:
position: The nucleotide at three bases upstream of the A of AUG (i.e., immediately before the AUG triplet). It is critical for strong initiation if it is an A or G.
position: The nucleotide immediately following the G of AUG (i.e., the first base 3’ of the AUG). A Guanine (G) at this position is highly important for strong initiation.
Strength tiers and their implications:
Strong context: Both key nucleotides ( R and G) are present. This configuration ensures highly efficient and accurate initiation at that specific AUG.
Adequate/weak context: One or both of the critical nucleotides are absent (e.g., a T or C at or any base other than G at ). This leads to lower initiation efficiency at that AUG, which may permit the scanning ribosome to bypass it and search for a downstream AUG (a phenomenon known as leaky scanning).
Alternative Start-Site Use – Leaky Scanning (Slide 13)
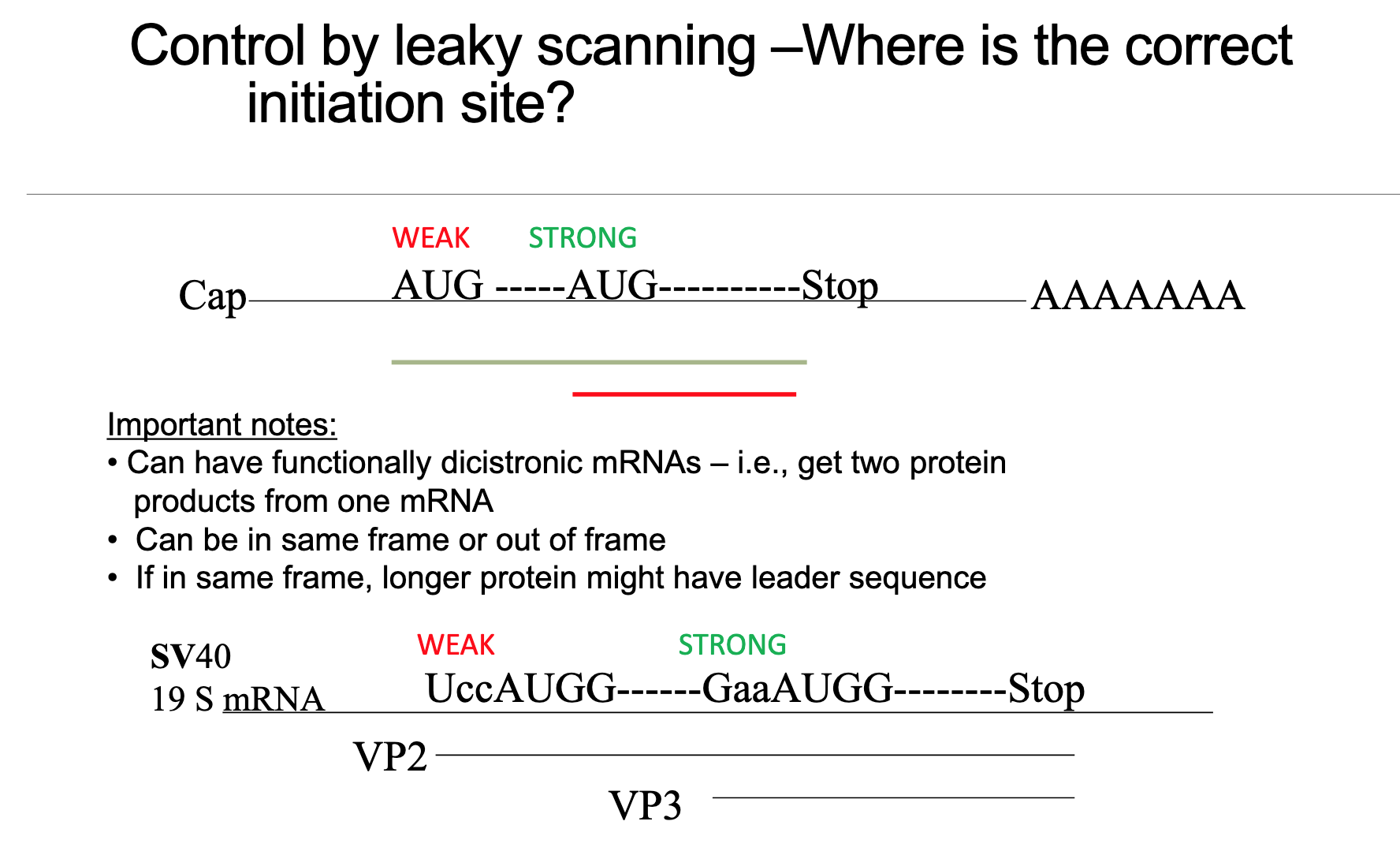
Leaky scanning occurs when an upstream AUG in the 5’ UTR is situated within a weak Kozak consensus context. Due to the suboptimal context, the 40S small ribosomal subunit occasionally (or frequently, depending on the weakness) ignores this upstream AUG and continues to scan downstream to a later, typically stronger, AUG.
Practical consequences:
This mechanism allows for the production of two or more distinct proteins from a single mRNA molecule, effectively making a eukaryotic mRNA functionally “bicistronic” (or even polycistronic, though less common than bacterial equivalents) despite its monocistronic design principle.
The proteins produced via leaky scanning often share the same C-terminal segment (as they are translated from the same coding sequence stretch after the second AUG) but differ in their N-terminal extension or may even originate from a different reading frame if the first AUG was not perfectly in-frame to establish the primary coding sequence.
SV40 19S mRNA example (Slide 13): A classic biological example of leaky scanning is observed in the Simian Virus 40 (SV40) 19S mRNA:
A weak upstream AUG (with a U at the position, making it a weaker Kozak context) leads to the production of the longer capsid protein VP2.
A stronger downstream AUG produces the major capsid protein VP3.
Approximately 90% of initiation events occur at the strong downstream site for VP3, while around 10% occur at the weak upstream site for VP2.
Experimental utility: Researchers, including Prof. Flanegan’s lab, routinely engineer Kozak strength in recombinant DNA constructs to precisely modulate which AUG is preferentially used, allowing them to control the expression of different protein isoforms or to optimize protein yield.
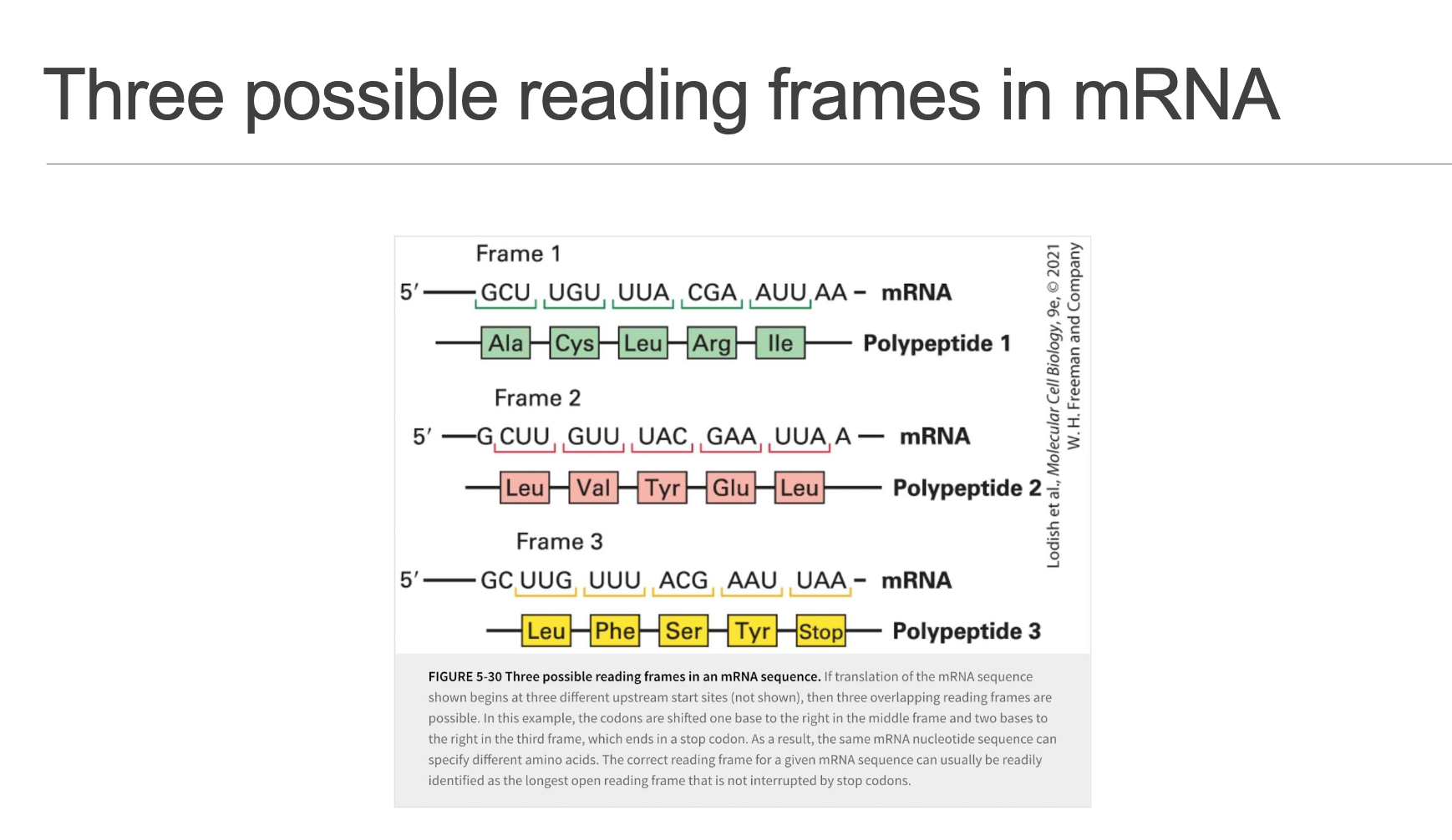
Reading Frames & Frame-Shifts (Slide 14)
Any mRNA potentially provides 3 different translational reading frames (offsets 0, +1, or +2 nucleotides from a given start point).
Initiation in the wrong reading frame or sporadic ribosomal frameshifting (where the ribosome shifts its reading frame during elongation) usually leads to the rapid encounter of a stop codon (UAA/UAG/UGA). Statistically, a random sequence is likely to contain a stop codon approximately every
h codons. This premature termination results in the production of significantly truncated and typically non-functional peptides.This highlights the absolute necessity for precise and accurate AUG selection during the initiation phase to ensure the synthesis of a full-length, functional protein in the correct reading frame.
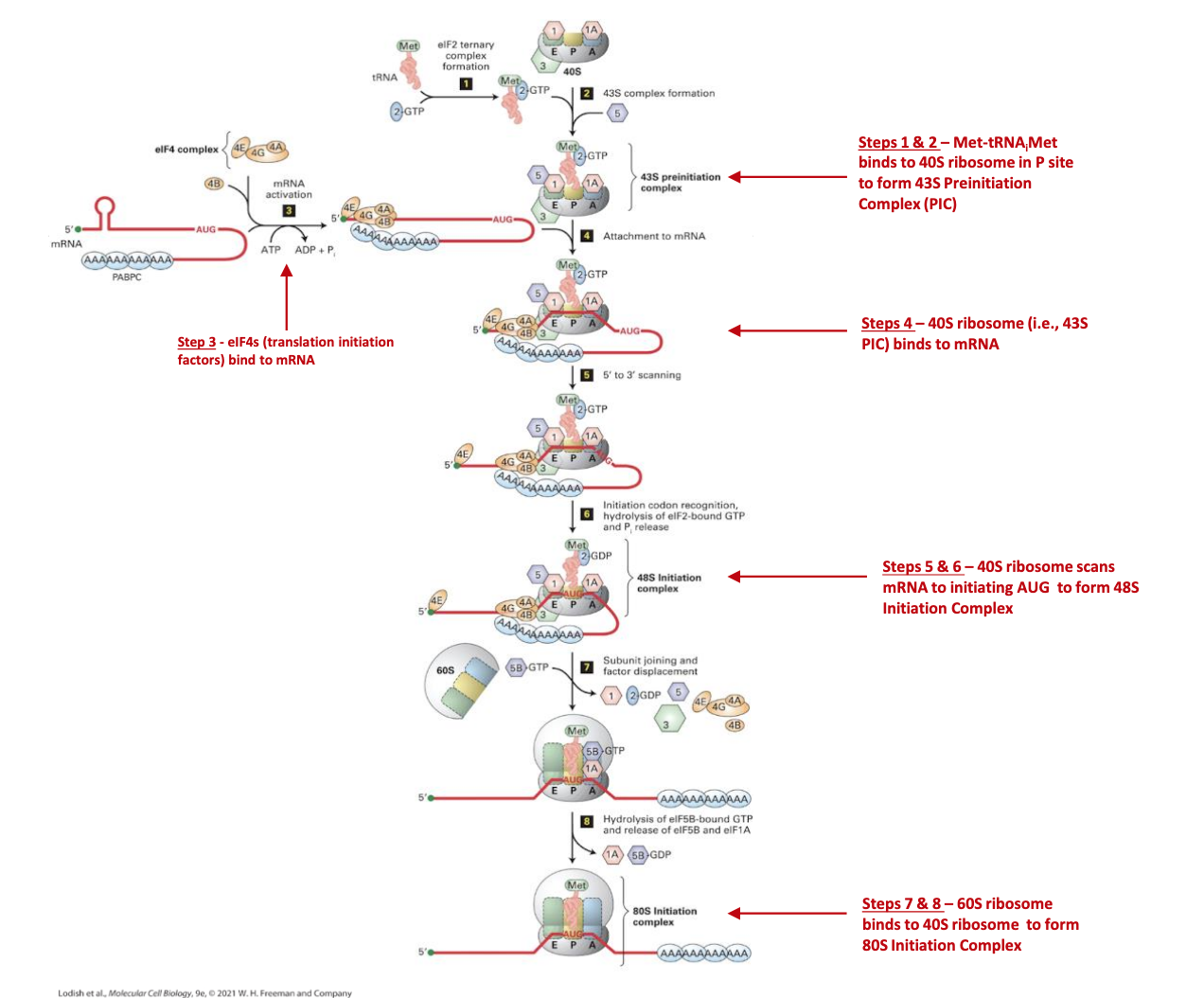
Detailed Step-by-Step Molecular Events (Slides 12–14)
These steps provide a comprehensive breakdown of eukaryotic translation initiation, integrating the roles of key initiation factors (eIFs) and the energy provided by GTP hydrolysis.
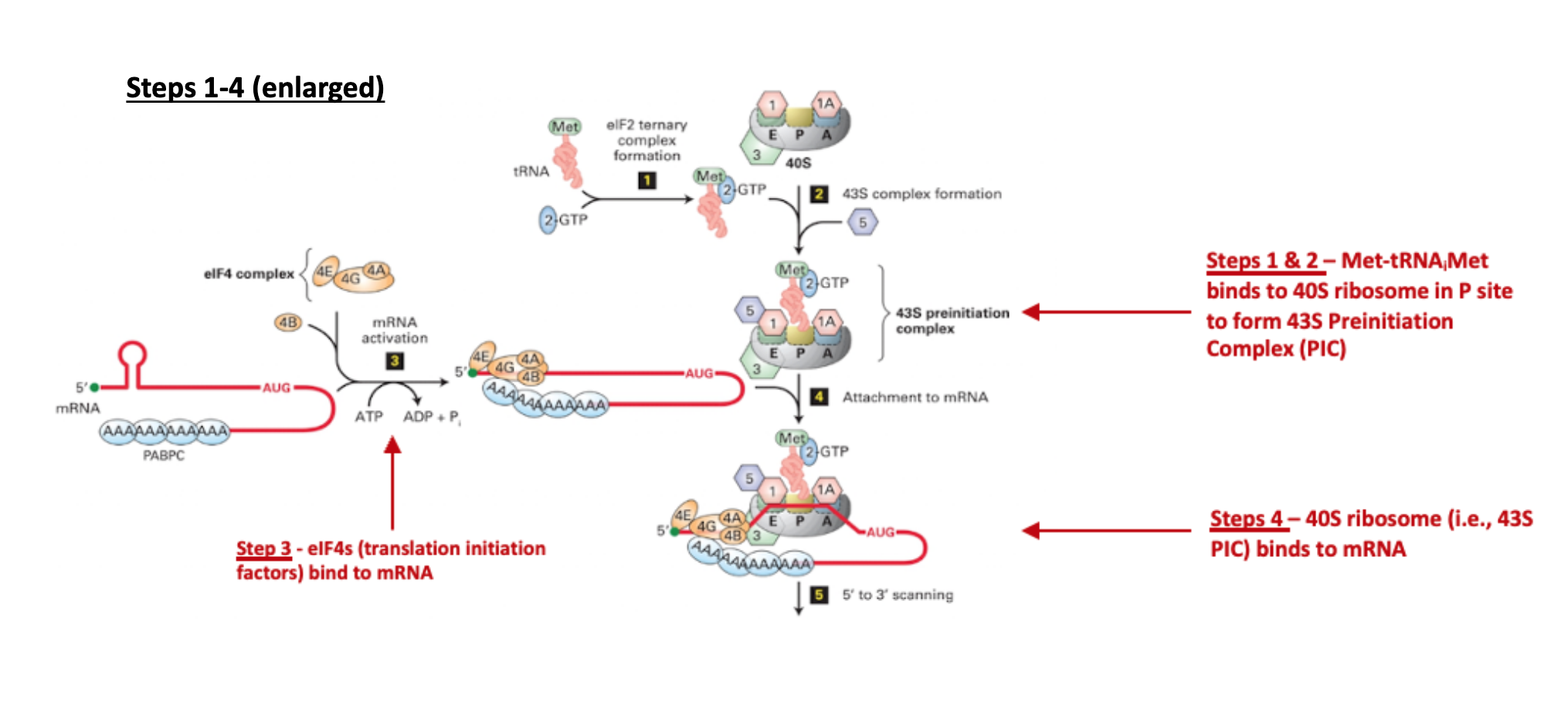
mRNA Activation / Closed-Loop Formation (Slide 12)
The process begins with the binding of eIF4E directly to the 5’ m⁷G cap of the mRNA, protecting it and marking it for translation.
Subsequently, eIF4G (a large scaffold protein) and the RNA helicase eIF4A (along with its enhancer eIF4B) join eIF4E at the cap. This complex, called eIF4F, facilitates recruitment of the ribosome.
Crucially, PABPC (Poly-A-Binding Protein Cytoplasmic), which is bound to the poly(A) tail at the 3’ end of the mRNA, interacts with eIF4G. This interaction effectively circularises the mRNA, bringing the 5’ and 3’ ends into proximity. This circularization is thought to promote efficient re-initiation after translation and protect the mRNA from degradation.
Formation of 43S Pre-Initiation Complex (Slide 12)
In parallel, the 40S small ribosomal subunit begins to assemble the 43S pre-initiation complex by binding several eukaryotic initiation factors: eIF1, eIF1A, and eIF3.
Simultaneously, eIF2–GTP (a GTP-binding protein) delivers the specialized initiator directly into the P-site of the 40S subunit. It is critical to note that the initiator tRNA is the only tRNA capable of entering the P-site directly without a preceding peptide chain.
mRNA Recruitment (Slide 13)
The eIF3 protein, which is part of the 43S pre-initiation complex, plays a pivotal role by interacting with the mRNA-bound eIF4G/A/B complex. This bridging interaction facilitates the loading of the entire 43S pre-initiation complex onto the 5’ end of the circularized mRNA.
Scanning (48S complex) (Slide 13)
Once loaded, the 43S complex (now often referred to as the 48S complex due to its association with mRNA) initiates ATP-powered scanning along the 5’ UTR in a 5’ to 3’ direction.
The eIF4A/B helicase activity is crucial during this phase, as it unwinds any inhibitory secondary structures (e.g., stable hairpins) encountered in the 5’ UTR.
This directed movement continues until the 40S subunit encounters the first Kozak-competent AUG start codon that is recognized as suitable for initiation.
Start-Codon Recognition (Slide 14)
Upon successful base pairing between the AUG start codon on the mRNA and the anticodon of the in the P-site, the scanning movement of the 40S subunit halts.
This correct recognition triggers a critical conformational change, leading to the GTP hydrolysis on eIF2. This hydrolysis is facilitated by eIF5, converting eIF2–GTP to eIF2–GDP + Pi. The dissociation of eIF2–GDP and other associated eIFs from the 40S subunit marks a point of no return for initiation.
Large-Subunit Joining (Slide 14)
After the dissociation of eIF2 and other factors, eIF5B–GTP steps in to escort the 60S large ribosomal subunit to the 40S subunit, facilitating their precise association.
A final GTP hydrolysis (on eIF5B-GTP) occurs, promoting the ejection of eIF5B (as eIF5B–GDP + Pi) and eIF1A from the now forming 80S complex.
80S Initiation Complex (Slide 14)
The successful completion of these steps results in the formation of the fully assembled 80S initiation complex.
At this stage, the A-site (Amino-acyl site) is vacant, ready to receive the first elongator tRNA.
The P-site (Peptidyl site) is precisely occupied by the initiator Met-tRNA, which is base-paired with the start codon.
The ribosome is now fully ready to recruit the first elongator tRNA (which will be delivered by eEF1A-GTP) and subsequently enter the elongation cycle, where sequential amino acids are added to the growing peptide chain.
Subsequent crucial steps, including peptide-bond formation (catalyzed by the peptidyl transferase activity of 28S rRNA) and translocation driven by eEF2, will be topics for forthcoming lectures.
Catalytic Role of Ribosomal RNA (Slide 15)
A profound discovery in molecular biology, the peptidyl-transferase centre within the ribosome is composed entirely of 23S rRNA in bacteria and 28S rRNA in eukaryotes, with no direct catalytic contribution from ribosomal proteins. This unequivocally establishes the ribosome as a ribozyme.
This observation highlights the notion that RNA molecules are not merely informational carriers but can also perform complex enzymatic catalysis, reinforcing earlier coursework on other catalytic RNAs such as self-splicing introns and RNase P.
Kinetic & Regulatory Considerations (Slide 16)
The rapid elongation rate of protein synthesis () significantly influences the overall protein synthesis throughput and the cell's ability to respond to changing demands.
Translation initiation is typically the rate-limiting step in protein synthesis and, consequently, represents the major control point for regulating gene expression globally. Key regulatory mechanisms include: (1) phosphorylation of eIF2α, which can lead to global translational repression, and (2) regulation of the availability or activity of eIF4E. These regulatory mechanisms will be covered in later lectures.
The presence of stable secondary structures in the 5’ UTR of mRNA (high negative hairpins) can significantly attenuate initiation efficiency by impeding the scanning ribosome. Viruses, in particular, often exploit or evolve mechanisms to counteract or even utilize such structures for their own translational control.
Connections to Previous & Future Lectures (Slide 17)
This lecture builds directly on foundational concepts previously covered, including mRNA capping and polyadenylation, mRNA stability mechanisms, and the critical accuracy of amino-acyl tRNA synthetase enzymes in charging tRNAs with the correct amino acids.
It meticulously prepares the ground for advanced topics like the detailed mechanics of translational elongation, the process of translational termination, various mechanisms of global translational control (e.g., in response to cellular stress), and the action of pharmacological inhibitors, such as antibiotics (which often target bacterial ribosomes) and anticancer agents (which can target eukaryotic translation).
Practical / Biomedical Relevance & Ethical Notes (Slide 18)
The ribosome is a highly effective and widely exploited target for many antibiotics. The significant structural differences between bacterial 50S/30S and eukaryotic 60S/40S ribosomal subunits are exploited to design drugs that selectively inhibit bacterial protein synthesis without harming host cells, emphasizing the evolutionary divergence despite a conserved core.
Mutations in specific ribosomal proteins or enzymes involved in rRNA modification are increasingly linked to a class of genetic disorders known as ribosomopathies (e.g., Diamond-Blackfan anemia), highlighting the critical role of ribosomal integrity in proper cellular function and development.
A deep understanding of the Kozak context is routinely employed in gene-therapy vector design, synthetic biology efforts (for designing synthetic gene circuits), and the optimization of recombinant protein yield in biotechnology, enabling precise control over gene expression.
Viral genomes (e.g., SV40) often ingeniously hijack host leaky scanning mechanisms to generate multiple structural proteins from compact, single mRNA genomes. This exemplifies the intricate co-evolutionary arms race between host translational machinery and pathogens, where viruses adapt to exploit cellular processes for their replication strategies.
Numerical & Statistical Highlights (Compiled) (Slide 19)
Ribosome elongation speed: . This represents the rate at which amino acids are added to a growing polypeptide chain.
Sedimentation coefficients of functionally assembled ribosomes:
Bacteria:
Eukaryotes:
Molecular weights of intact ribosomes, illustrating increasing complexity through evolution:
Core: (million Daltons)
Bacteria (E. coli):
Yeast (lower eukaryote):
Human:
Total ribosomal protein counts across different organisms (approximate):
Core (ancestral): 34 proteins
Bacteria: 54 proteins
Yeast: 79 proteins
Human: 80 proteins
rRNA nucleotide counts:
Bacteria: Approximately
nucleotidesHuman: Approximately
nucleotides
Frequency of first-AUG initiation in eukaryotes: Approximately 90\ \text{%} of translation initiation events occur at the first available AUG codon from the 5' end of the mRNA, provided it's in a strong context. The remaining approximate 10\ \text{%} instances are attributed to mechanisms like leaky scanning or other alternative initiation pathways.
Summary Checklist for Exam Review
[ ] Name and describe the roles of the three main translation components: mRNA, charged tRNAs, and ribosomes.
[ ] Identify the A (Amino-acyl), P (Peptidyl), and E (Exit) sites within the ribosome and explain their specific functions in the translation cycle.
[ ] Clearly contrast bacterial versus eukaryotic ribosome sizes (Svedberg units) and their respective subunit compositions (rRNA types and protein numbers).
[ ] Explain the concept of catalytic RNA (ribozyme), detailing its location within the ribosome (peptidyl transferase center in the large subunit rRNA) and its significance.
[ ] Be able to draw or annotate a diagram of a typical eukaryotic mRNA molecule, indicating the crucial elements: the 7-methyl-guanosine cap, the Kozak-embedded AUG start codon, and the poly(A) tail.
[ ] Trace the approximate sequence of molecular events from the initial recognition of the mRNA cap to the final assembly of the 80S initiation complex, naming the key eukaryotic initiation factors (eIFs) and identifying the critical GTP-driven steps (conceptual order of events is more important than memorizing every single factor ID).
[ ] Accurately reproduce the Kozak consensus sequence and predict the likely effect of point mutations at the crucial or positions on initiation efficiency.
[ ] Describe the mechanism and functional consequences of leaky scanning, and be able to provide and explain the SV40 VP2/VP3 example as a classical illustration.
[ ] Be able to reason about different translational reading frames, the consequences of frame-shifts, and the result of encountering premature stop codons.
[ ] Link the regulation and control of translation initiation (e.g., via eIF2α phosphorylation) to overall gene-expression regulation and specific disease contexts (e.g., ribosomopathies).
Instructor note: Conceptual understanding will be rewarded more heavily than rote memorization of every factor name or numerical detail. Focus on the 'why' and 'how'.