M5L5 Targeting epigenetic mechanisms in cancer therapy
DNMT inhibitors - nucleoside analogues (5’azacytidine and 5-2’ decitabine) and non nucleoside analogues (procainamide)
RNA can be methylated by different marks - N6-methyladenosine most common, requires METTL3/14
METTL3 —> oncogenic targets eg MYC, EGFR upregulation
Histone variant H3.3
Mutations identified in H3F3A and H3F3B encoding histone variant H3.3
First oncohistone
Recurrent mutations affecting K21 and G34
Chromatin remodelers use ATP to slide or eject nucleosodes
Eg SWI/SNF family (eg BAF, pBAF), ISWI, CHD…
SWI/SNF mutated in ~20% of most cancer types
Histone modifications
Writers - HMT, HAT, Ub ligases
Readers - PWWP domains, bromodomains, chromodomains
Erasers - KDMs, HDACs
Acetylation
HATs eg GCN5, P300, CBP, Tip60
Adds acetyl group which neutralises pos charge of lysine —> relaxed chromatin structure
Irregularities —> cancer
4 classes of HDACs, including HDAC1-10 and SIRT1-7
HDAC inhibition —> open chromatin, gene expression, may reactivate silenced tumour suppressors
Methylation
KMTs are highly mutated across cancers
Gain of function mutation
Key determinant of altered methylation - hypoxia
DNA and histone demethylases are dioxygenases - requiring oxygen to work
Hypoxia blocks TET demethylase and KDMs blockingf CpG and histone demethylation
Blocks cell differentiation —> promotes tumorigenesis
Oncometabolites affect DNA and histone methylation
Isocitrate dehydrogenase (IDH) converts isocitrate to alpha ketoglutarate in Krebs cycle
IDH gain of function converts alpha-KG to 2-hydroxyglutarate (G2HG) which competitively inhibits alpha-KG substrates, driving cancer as a-KG is needed by TET and KDM for demethylation
IDH1 and IDH2 inhibitors developed
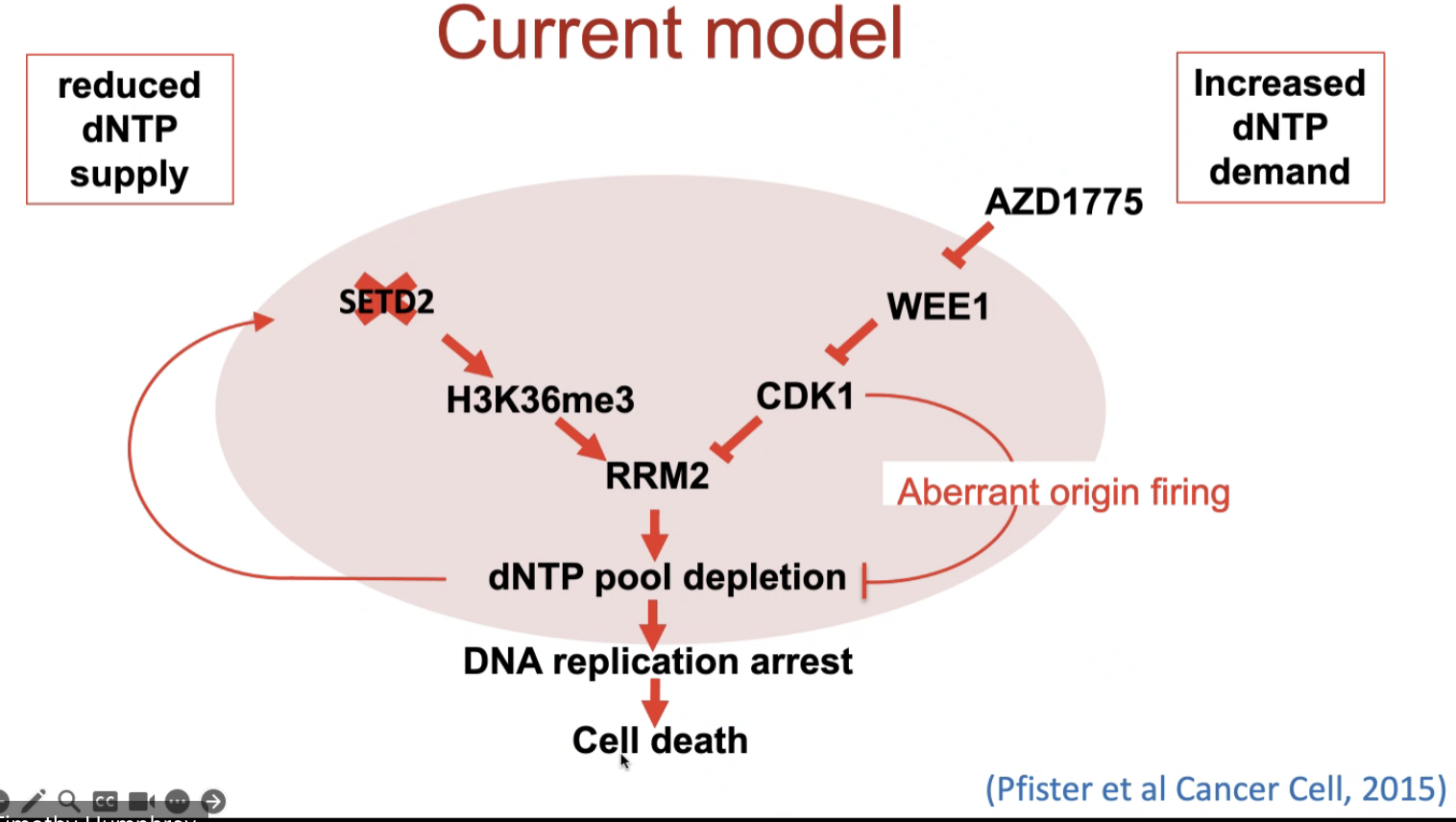
Targeting H3K36me3 deficient cancers
SETD2 is an HMT specific for H3K36 trimethylation
KDM4A can reverse trimethylation (could be upregulated in cancers)
SETD2 mutated in many cancers (tumour suppressor role)
H3.3 can inhibit H3K36me3
SETD2 is synthatic lethal with Wee1 in fission yeast
Ribonucleotide reductase (RRM2) levels are reduced when Wee1 is inhibited
RRM2 is degraded by Cdk1 in G2 phase due to its phosphorylation at Thr33 and subsequent ubiquitination
Wee1 loss leads to increased dNTP demand but SETD2 loss reduces dNTP supply