11. Antibiotic Production I-> Discovery and Mechanism of Action
What is an antibiotic?
An antibiotic is any compound that inhibits the growth, or actively kills, microorganisms
Imprecise definition – what about detergents, or generally toxic compounds? If it kills you as well as the bacteria, is it an antibiotic?
Therefore usually refers to compounds that selectively target bacteria
Two major modes of action:
Bacteriostatic: Inhibits bacterial growth without actively killing
Bactericidal: Actively kills bacteria
Most antibacterials are better described as potentially being both bactericidal and bacteriostatic, depending on concentration and the bacterial infection being treated
Broad-spectrum antibiotic: active against a wide range of pathogenic bacteria.
Narrow-spectrum antibiotic: active against a specific family (or limited number) of pathogenic bacteria.
Minimum inhibitory concentration (MIC): the lowest concentration of a drug that will inhibit the visible growth of an organism after overnight incubation.
An MIC <4 μg/mL is usually desirable, but an acceptable MIC range depends on various factors, including the antibiotic, the bacteria and the site of infection
Clinically used antibiotic classes
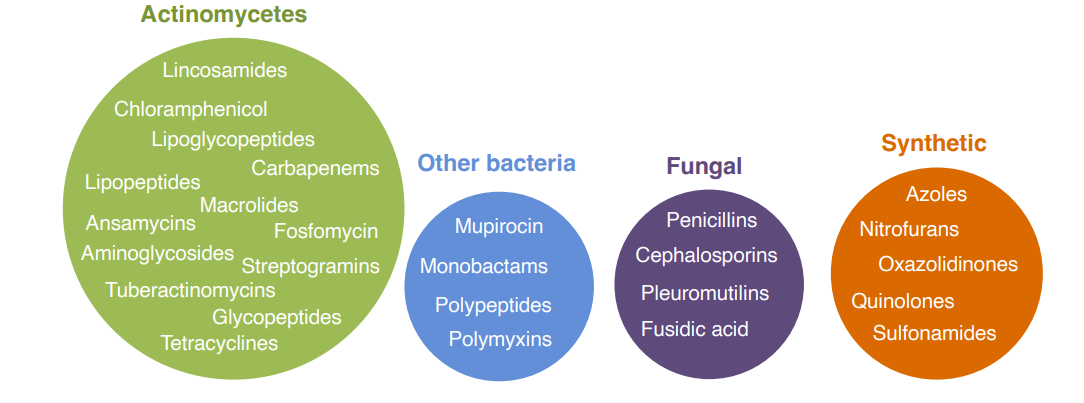
Actinomycetes are Gram-positive, non-motile bacteria often found in the soil
Also produce a variety of other clinically or agriculturally important compounds, such as anti-cancer agents, immunosuppressants, anti-fungals, herbicides and insecticides
Traditional approach to antibiotic discovery
Environmental sample (traditionally soil)
Plated to identify bacteria and fungi in sample
Individual organisms screened for their ability to produce antibiotics
Fermentation and subsequent purification to isolate pure antibiotic
New clinically useful antibiotics
Industrial production of antibiotics
Often need to improve productivity from a few mg per litre of culture to > 1 g
Wild type antibiotic-producing strain
Selection and maintenance of stable producer → Want reliable production and growth characteristics
Mutagenesis and selection of antibiotic overproducers → Random or selective mutations in producing strain can yield big increases in production
Fermentation medium optimisation → Antibiotic yield can be very medium specific. A cheap and easy to use medium is also an advantage.
Fermentation process optimisation and scale-up → Fermentation and purification processes needs to be suitable for industrial scale fermenter (earlier work often carried out on small scale)
Industrial Process
Important classes – natural products
Beta-lactams: penicillins
Amoxicillin → Semi-synthetic derivative of penicillin, produced by Penicillium chrysogenum
Beta-lactams: cephalosporins
Cefacetrile → Semi-synthetic derivative of cephalosporin, produced by C Acremonium chrysogenum
Aminoglycosides
Kanamycin A → produced by Streptomyces kanamyceticus
Macrolides
Erythromycin → produced by Saccharopolyspora erythraea
Glycopeptides
Vancomycin → produced by Amycolatopsis orientalis
Polymyxins
Colistin → produced by Paenibacillus polymyxa
Tetracyclines
Tetracycline → produced by Streptomyces aureofaciens
Ansamycins
Rifampicin → Semi-synthetic derivative of rifamycin, produced by Amycolatopsis rifamycinic
Important classes – synthetic compounds
In general, synthetic antibiotics are structurally more simple than natural product antibiotics
Very difficult (and/or expensive) to synthesise complex natural products, which are almost all produced industrially by fermentation
Oxazolidinones
Linezolid
(Fluoro)quinolones
Ciprofloxacin
Azoles
Metronidazole
Sulfonamides
Mafenide
Antibacterial targets
How do you kill or inhibit bacterial growth?
Target an process that is essential for bacterial survival (or is at least essential under certain conditions)
Sounds simple but there are surprisingly few successful targets for clinically-used antibiotics
Cell wall biosynthesis → reduce the structural integrity of the cell
Protein synthesis → prevent the cell from making essential proteins (and also corrupt this process so that the wrong proteins are made)
Nucleic acid synthesis → prevent the cell from making essential nucleic acids (DNA and RNA)
Ideally, targets should be unique to bacteria to avoid side-effects in humans
β-lactam antibiotics (e.g. Penicillin G)
Most widely used class of antibiotic
Broad-spectrum β-lactam antibiotics are active against both Gram-negative and Gram-positive bacteria
β-lactam target – the peptidoglycan
The peptidoglycan forms an integral part of the bacterial cell wall
A polymeric material formed from sugars (glycans) and amino acids
β-lactam mechanism of action (MOA)
Penicillin binding protein (PBP) is a transpeptidase that catalyses peptidoglycan cross-linking
Penicillins mimic the substrate of the transpeptidase, thus binding in the active site
The beta-lactam reacts with the active site serine to inactivate the enzyme
Antimicrobial resistance (AMR)
Carbapenase: an enzyme that degrades carbapenems, β-lactam antibiotics used for the treatment of serious infections caused by multidrug-resistant bacteria.
Enterobacteriaceae: a large family of Gram-negative bacteria that includes many widespread pathogens, such as Salmonella, Escherichia coli, Yersinia pestis, Klebsiella and Shigella.
Antibacterial resistance → How do bacteria evade antibiotics?
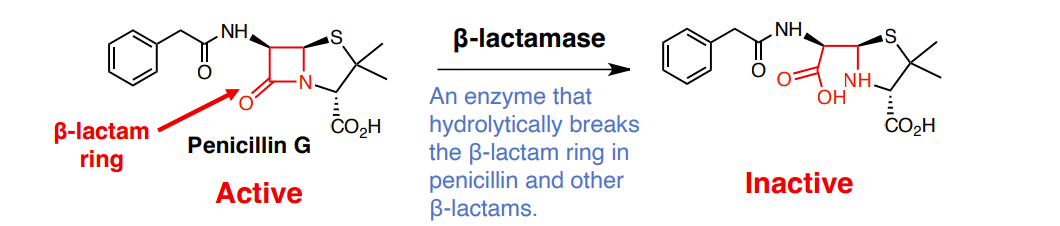
1. Destruction: enzymatically degrade or modify the antibiotic to render it inactive.
e.g. β-lactamases that degrade beta-lactams
 e.g. enzymatic modification of aminoglycosides
e.g. enzymatic modification of aminoglycosides
N-Acetyltransferases (AAC): Catalyse acetyl CoA-dependent acetylation of an amino group
O-Adenyltransferases (ANT): Catalyse ATP-dependent adenylation of a hydroxyl group
O-Phosphotransferases (APH): Catalyse ATP-dependent phosphorylation of a hydroxyl group
2. Exclusion: prevent the antibiotic from entering the cell, or remove it from the cell before it can do damage
e.g. the tetracycline TetA efflux pump
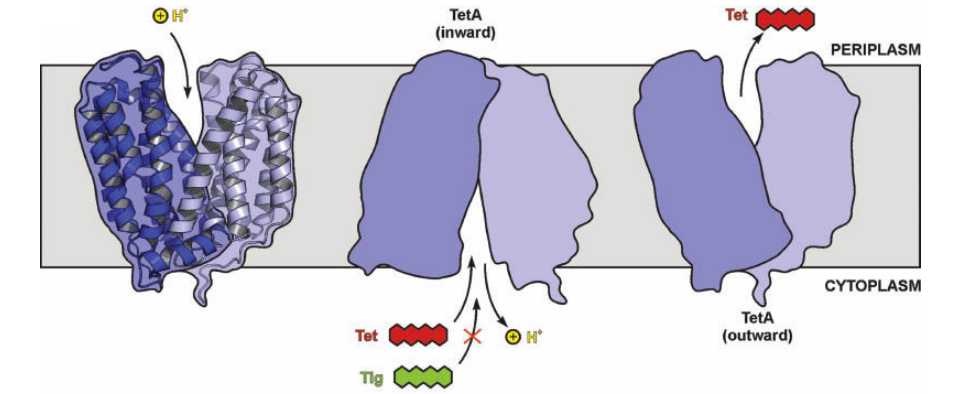
TetA uses a proton gradient to actively export tetracycline (Tet)
The antibiotic tigecycline (Tig) can evade this exporter
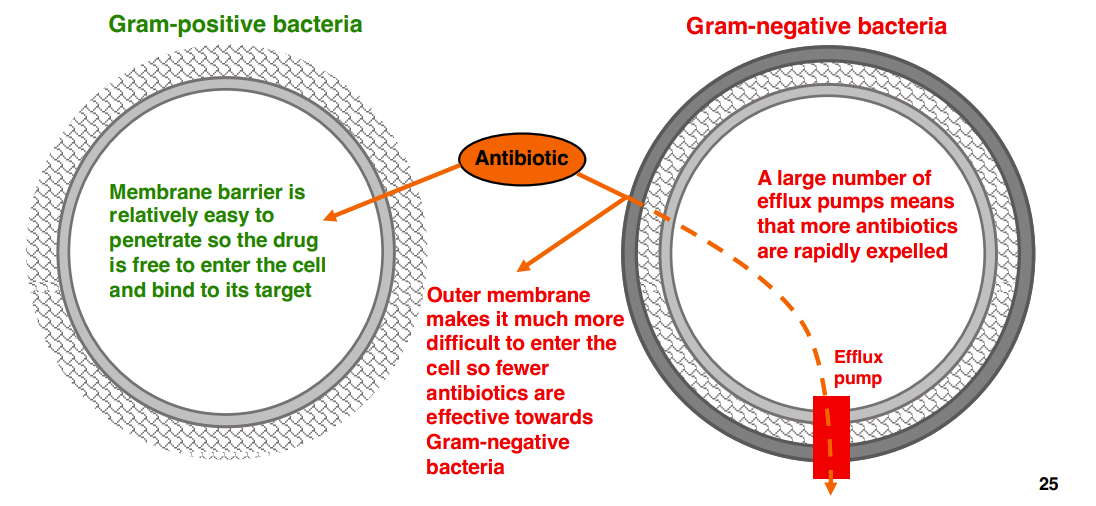
Also: Gram-negative bacteria outer membrane (intrinsic resistance)
 3. Target modification: modify the antibacterial target so that the antibiotic can no longer bind. Alternatively bypass the antibiotic target so that it is no longer essential.
3. Target modification: modify the antibacterial target so that the antibiotic can no longer bind. Alternatively bypass the antibiotic target so that it is no longer essential.
a. Point mutation to target → Changes to target structure that prevent strong binding of antibiotic (must enable the target to retain its required function)
b. Modification of target → Protection of the target by enzymatic modification e.g. The erythromycin ribosome methylase (Erm) methylates 23S rRNA, which prevents binding of erythromycin, as well as other classes of antibiotic (lincosamides and streptogramins)
How does resistance develop? Adaptive evolution on a rapid timescale
Random mutagenesis: mutants that confer increased survival towards an antibiotic are selected for during antibacterial treatment
Horizontal gene transfer: entire genes (or clusters of genes) are transferred from one bacterium to another as mobile genetic elements. More dangerous due to the capacity for spreading complex multi-gene resistance mechanisms.