Chapter 4: Taxonomy and Phylogeny of Animals
[[Linnaeus and Taxonomy[[
Taxonomy: The study of the principles of systematic ordering and naming of organisms
- Part of a broader science of systematics
* Study of variation among animal populations to reveal their evolutionary relationships - Provides evolutionary biology
* Adjusting system to accommodate evolution has produced many problems - Carl von Linne
* (Latin pen name: Carolus Linnaeus) (1707-1778)
* Produced extensive system of taxonomy for both plants and animals
* Binomial Nomenclature
* 2 names: genus name + species epithet/name
* Species grouped in hierarchical categories
* Taxa (singular: taxon)
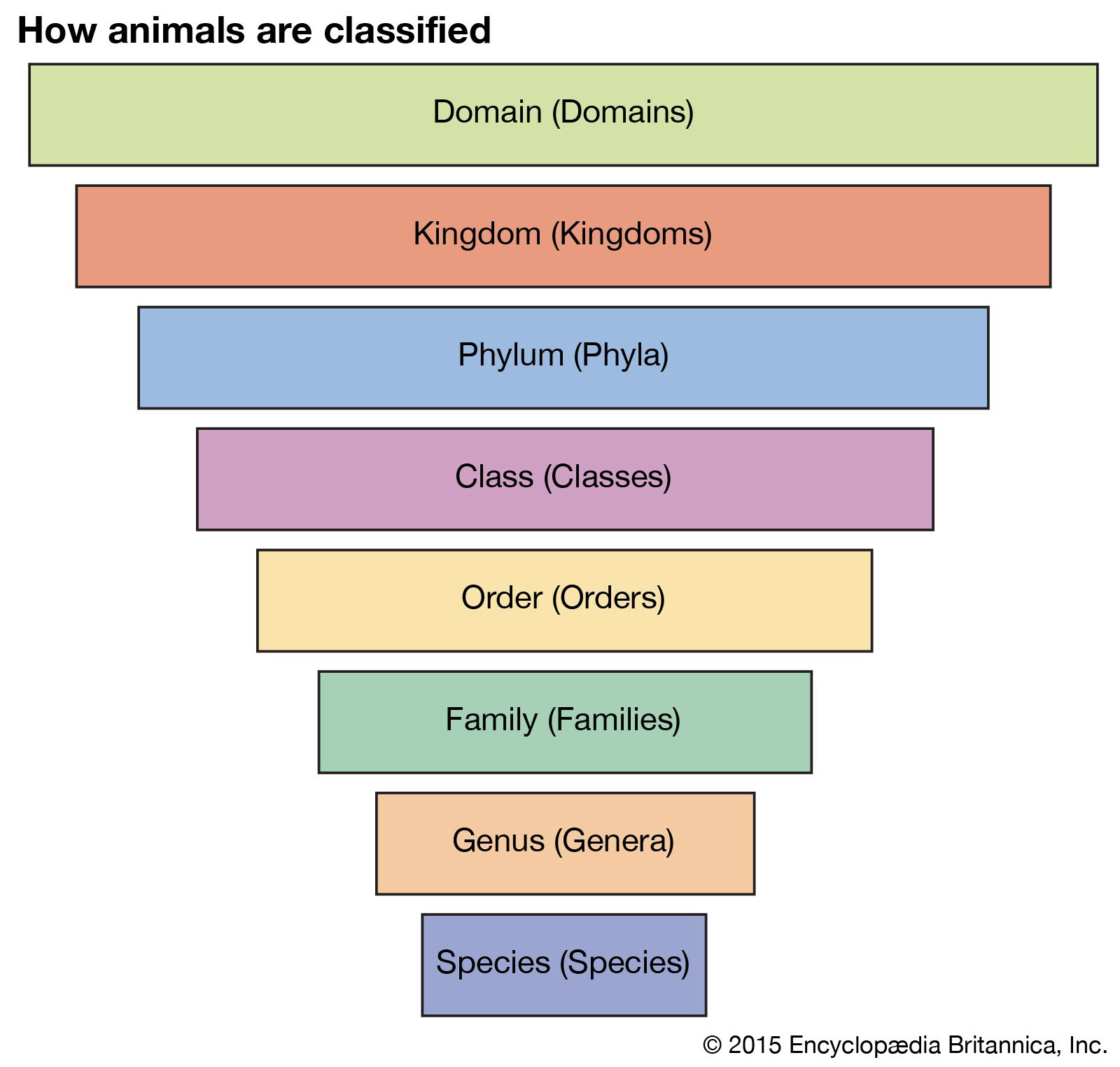
# [[Species[[
- Difficult to define
* Important criteria for recognition of species
- Common Descent
1. Members of a species must trace their ancestry to a common ancestral population
- Species must be the smallest distinct groupings of organisms sharing patterns of descent and ancestry
1. To ID such groupings:
1. Morphological characters
2. Chromosomal characters
3. Molecular characters
- Members of a species must form a reproductive community that excludes members of other species
==Concepts of Species:==
- Most influential: Biological Species Concept
* Inspired by Darwinian evolutionary theory
* “A species is a reproductive community of populations (reproductively isolated from others) that occupies a specific niche in nature.” - Ernst Mayr, 1983 - Species = interbreeding population of individuals having common descent
* Not based on organismal morphology
* But can still help to diagnose biological species
* Variation should be relatively smooth and continuous within species and discontinuous between them - Criticisms
* Refers to contemporary populations
* How can we trace the temporal duration of a species’ lineage through its past history?
* Example: Humans
* When are human fossils no longer Homo sapiens, but different species?
* Species do not exist in groups of organisms that reproduce only asexually
* How much reproductive divergence is necessary to consider two populations separate species?
Evolutionary Species Concept
- Proposed by George Gaylord Simpson in the 1940s
- Adds evolutionary time dimension to the biological species concept
- “A single lineage of ancestor-descendant populations that maintains its identity from other such lineages and that has its own evolutionary tendencies and historical fate”
- Applies both to sexually and asexually reproducing organisms
- Abrupt changes in diagnostic features mark a boundary between different species in evolutionary time
- Updated in 1989 by Alan Templeton
* Population geneticist
* Included the expectation that populations of a species evolve as a genetically cohesive unit by natural selection and genetic drift
* Cohesion Species Concept
* Any individual in a species is a possible common ancestor of the entire species at some future time
Phylogenetic Species Concept
- Irreducible (basal) grouping of organisms diagnosably distinct from other such groupings and within which there is a parental pattern of ancestry and descent
- Emphasizes common descent
- Both asexual and sexual groups covered
- Any spatially separated population that has undergone character evolution that distinguishes it is recognized as a species
- Would describe a larger # of species than would any other concept
\
Current disagreements about species definitions are exciting.
- Will lead to enormous advances in biology and fundamental reconsiderations of the meaning of species
==DNA Barcoding of Species==
Does not resolve the definition of species
- Useful for Identifying an unknown organism
DNA Barcoding
- Technique for diagnosing organisms to species using sequence information from a standard gene present in all animals
* COI (cytochrome c oxidase subunit 1)
* Mitochondrial gene
* Standard “barcode” region for animals
* Process
* DNA extracted
* Gene targeted
* Many copies made
* Sequenced
* Sequence checked against a public reference library of identified species (database)
* Can ID unknown species
[[Taxonomic Characters and Reconstruction of Phylogeny[[
Once species are named, how do we decide where each one fits on our tree (phylogeny)?
- Based on characters that vary among species
* Any feature that a taxonomist uses to study variation within or among species
* Can be morphological, chromosomal, and/or molecular
* Homologous characters are useful for constructing phylogenies
* Similar features that are the result of common ancestry
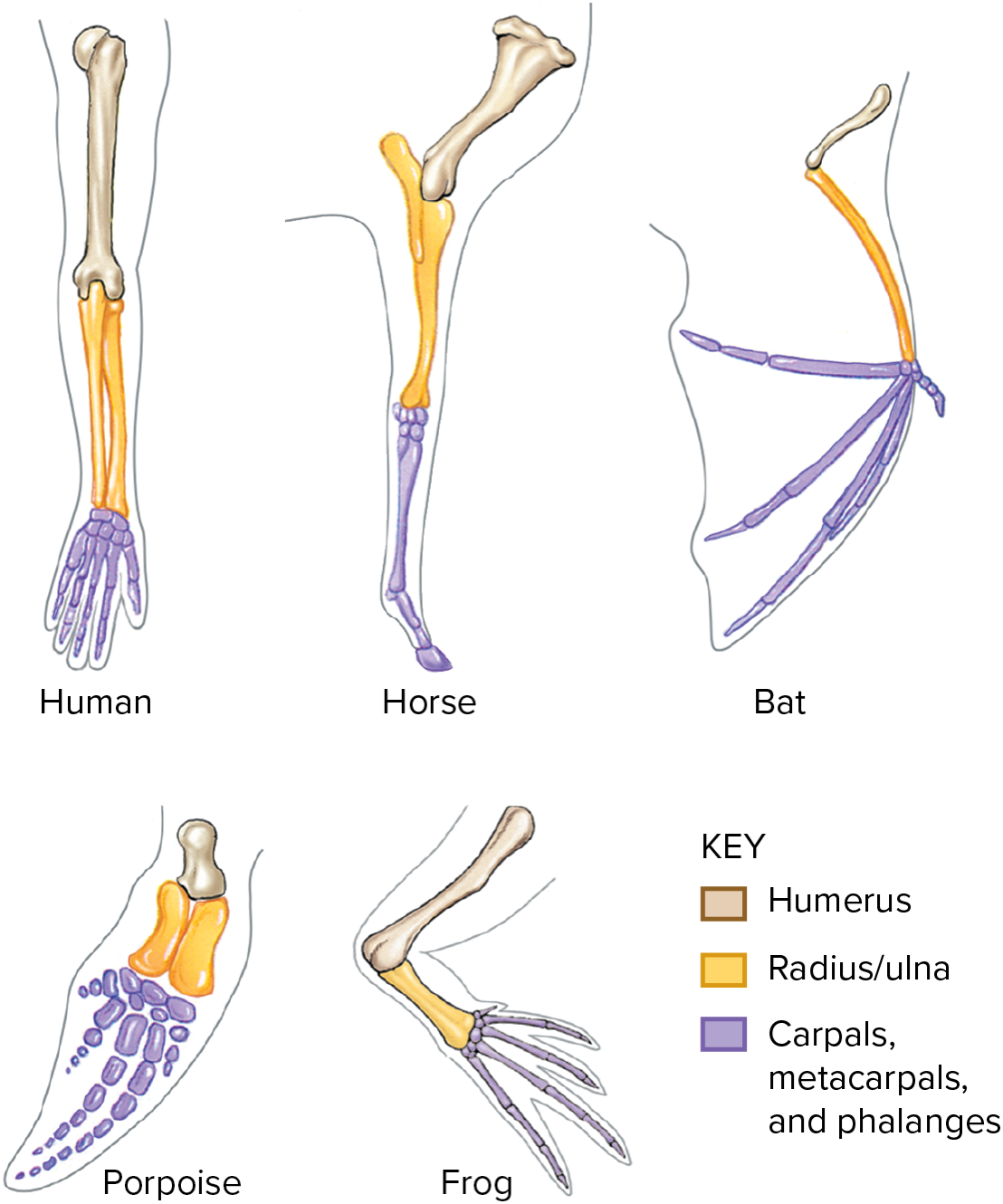
- Similarity does not always reflect common ancestry
* Homoplasy
* Result of independent or convergent evolution
* Analogous Characters
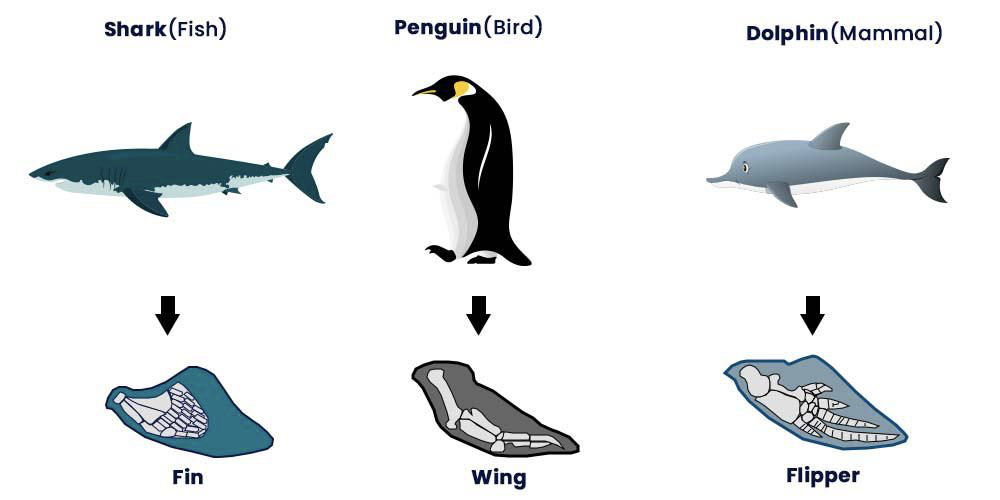
[[Using Character Variation to Reconstruct Phylogeny[[
==First Step:==
- For each character, which character state was present in the most recent common ancestor of the entire taxon?
* Ancestral Character State
* Contrasting states = Derived Character States
* Derived characters shared by all members of a clade = synapomorphies
* Clade = fundamental unit of the phylogenetic grouping of species
* Ancestral lineage + all descendants from lineage
* Basic Method
* Compare characters in group of interest (= ingroup) to those of an outgroup
* Reference group
* Known to be related to the study organisms
* Less closely related to any member of the ingroup than the outgroups are to each other
* Ancestral for ingroup
==Next Steps:==
- Use synapomorphies as evidence of homology to infer that a particular group of species forms a clade
- Pattern formed by all synapomorphies within ingroup reveals a nested hierarchy of clades within clades
* Goal: Identify all clades nested within ingroup
* Could reveal the structure of common descent among species
* Diagram = cladogram
* Not the same as a phylogenetic tree
* Needs more info
* Duration of evolutionary lineage, ancestor info, etc.
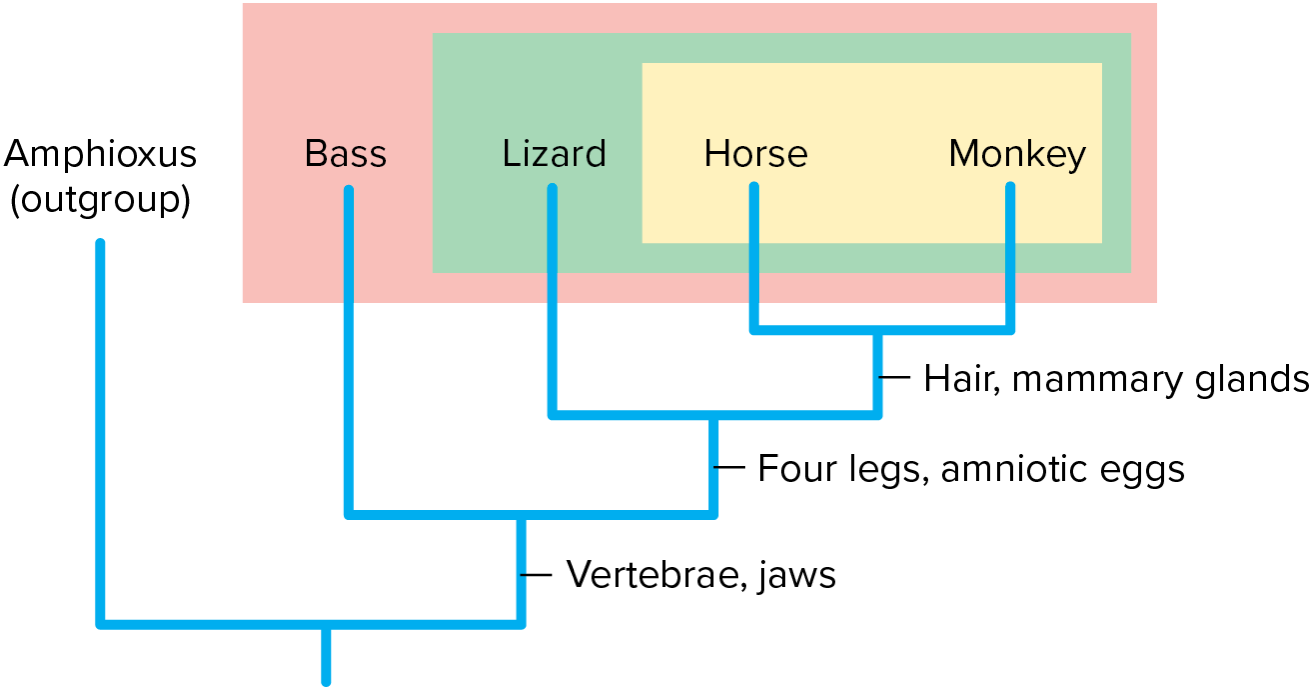
[[Theories of Taxonomy[[
2 currently popular theories of taxonomy
- Both based on evolutionary and phylogenetic principles
* Difference: how those evolutionary principles are used
- ==Evolutionary Taxonomy==
* Predates phylogenetic systematics
* Retains many aspects of Linnean taxonomy
* Species grouped in a nested hierarchy of increasingly more inclusive higher taxa
* All taxa must:
* Have a single evolutionary origin
* Include the most recent common ancestor of all members of the taxon - ==Phylogenetic Systematics (cladistics)==
* Developed in the mid-20th century by Willi Hennig
* Emphasizes criterion of common descent
* All taxa must be monophyletic
* Includes common ancestor and ALL descendants
* Members are linked by nested sets of characters that trace the evolutionary history of the group
* These principles underpin cladistics
* Method of investigating evolutionary relationships based on analyses of the distribution of characters
Our Approach:
- Emphasize monophyletic taxa
- Consistent with the criteria of both theories
[[Major Divisions of Life[[
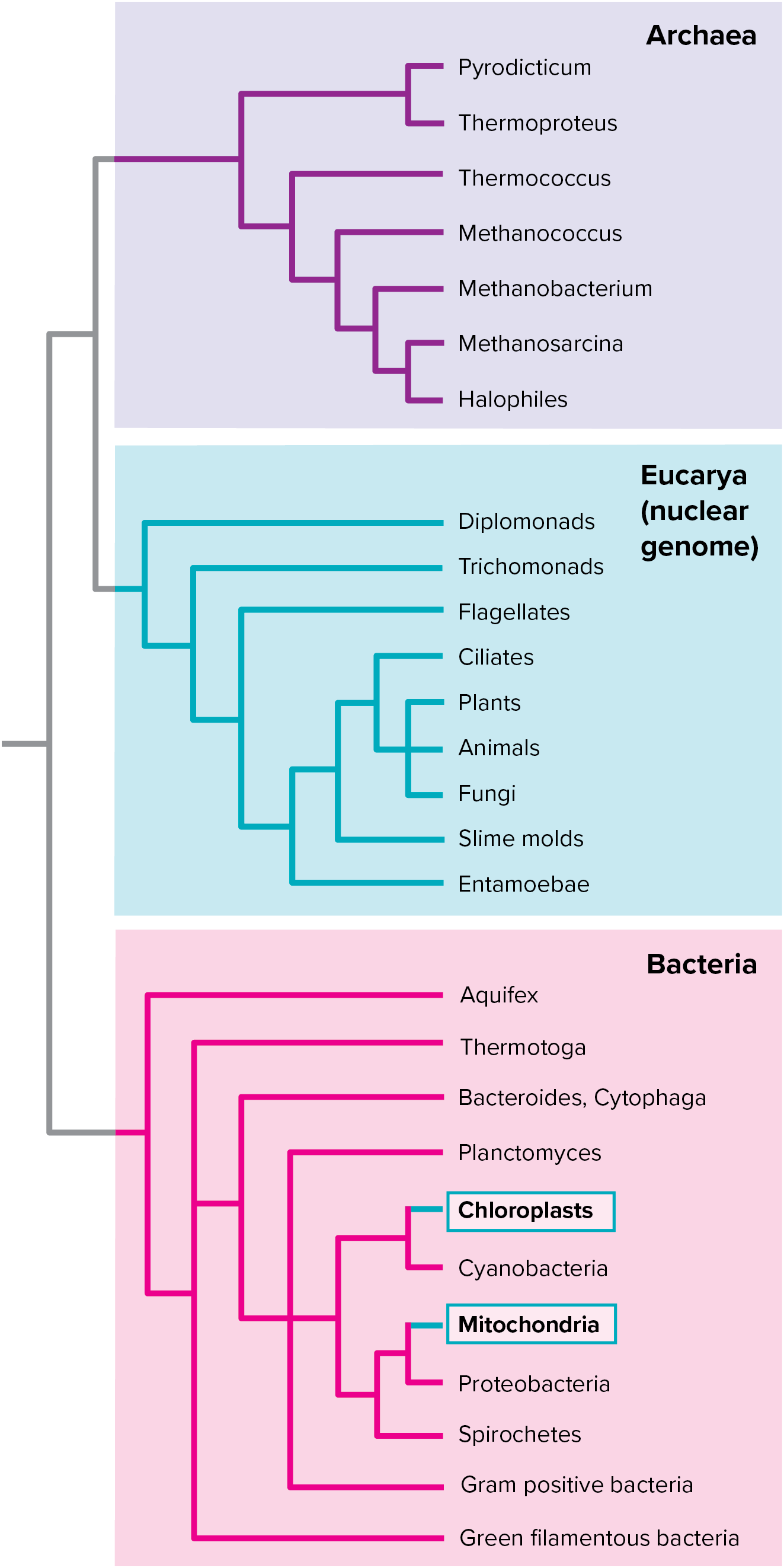
All life is categorized into 3 monophyletic domains:
- Bacteria
* Prokaryotes - Archaea
* Prokaryotes
* Differ from bacteria in membrane structure and ribosomal RNA sequences - Eukarya
* All eukaryotes
\