Wk 1.3 Bacterial Genetics and Antibiotic Resistance
Learning Outcomes
Lecture 3
Understand the make up of bacterial genomes
Understand what changes can occur to bacterial genomes (mechanisms of genetic diversity) & how these can effect bacterial pathogenicity
Hint: What is meant by & what are homologous recombination, transposition, mutation, transformation,
conjugation, transduction?
Understand the role of plasmids in bacterial function
Bacterial DNA
Organized as a single double stranded circular chromosome in most prokaryotes.
Exceptions: Linear (e.g., Borrelia) or multiple chromosomes (e.g., Vibrio).
DNA is supercoiled and associated with basic proteins (histone-like).
Contains plasmids: small extrachromosomal double stranded DNA.
DNA Replication in Bacteria
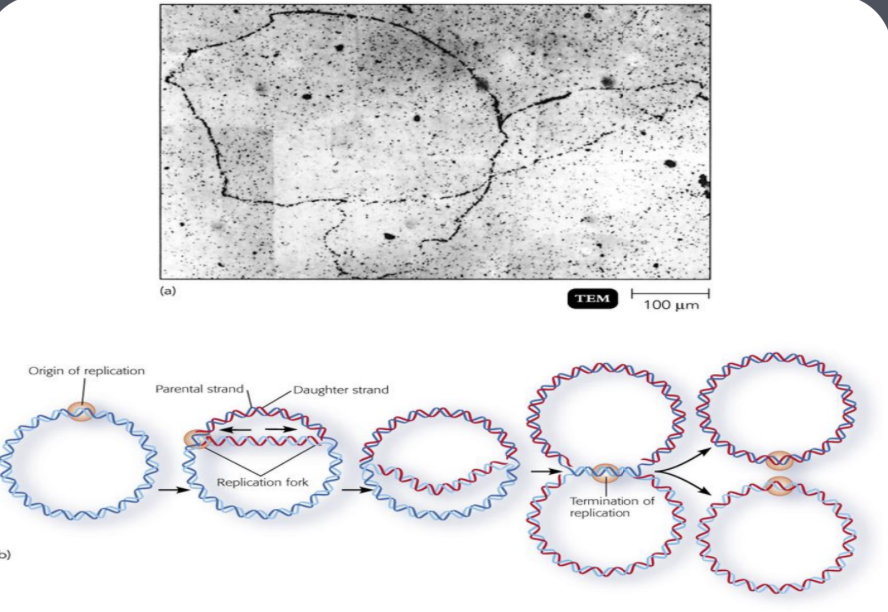
Initiation at a single point (origin) with synthesis at the replication fork. (place where at which the helix is unwound)
Two replication forks move outward from the origin until the a whole replication is copied
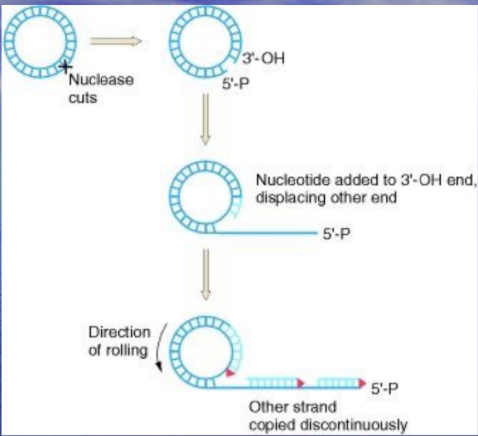
Chromosome is a single replicon, using the rolling circle mechanism during conjugation in E. coli.
At least 30 proteins are involved in E. coli replication.
Gene Structure
One gene-one enzyme hypothesis and one gene-one polypeptide hypothesis (cistron).
Some genes encode rRNA and tRNA.
Sequence of nucleic acid transcribed to generate an RNA product
Genes typically consist of discrete sequences that produce one way and one product
Single starting point and one reading frame (no overlapping)
Some do overlap
Introns present in some bacterial genes.
Gene Expression
Bacteria regulate genome expression to conserve energy and raw materials, maintaining protein balance, and allowing adaptation.
E. coli can synthesise 2000-4000 peptides, expressing a fraction at any one time.
Induction and Repression
E.coli growing in the presence of lactose induces synthesis of b-galactosidase (3000 molecules) while other bacteria in the absence leads to very low levels (<3 molecules).
beta-galactosidase is an inducible enzyme
Inducible enzymes triggered by inducers like allolactose. (ตัวเหนี่ยวนำ)
Repressible enzymes produced unless their pathway's end product is present (e.g., amino acids for biosynthesis).
will repress the expression of the enzyme required for its biosynthesis
Plasmids
Small, double stranded circular DNA in many bacteria; can exist independently of the host chromosome.
Separate replicon
Usually contains <30 genes; numerous plasmids (>40) may be present; just one
Not essential for survival, but play significant roles.
Plasmid Classification
Episome: can exist integrated or independently with chromosomes.
Conjugative: contribute to pili formation and conjugation.
F factors(fertility): fertility factors involved in conjugation.
R factors (resistant): confer antibiotic resistance.
Col plasmids: encode bacteriocins.
Virulence plasmids: produce toxins, siderophores, and aid adherence.
Metabolic plasmids: encode degradative enzymes.
Transposons
DNA segments that move within chromosomes or plasmids.
Cannot replicate independently
May contain genes required for transposition
Can carry genes like toxins or antibiotic resistance.
Effects of Transposons
DNA rearrangements (deletions)
Mutations
Gene activation by containing promoters (activate genes) or stop codons (block translation or transcription).
Transfer of Genetic Material (recombination DNA)
Occurs via transformation, transduction, and conjugation.
Transformation
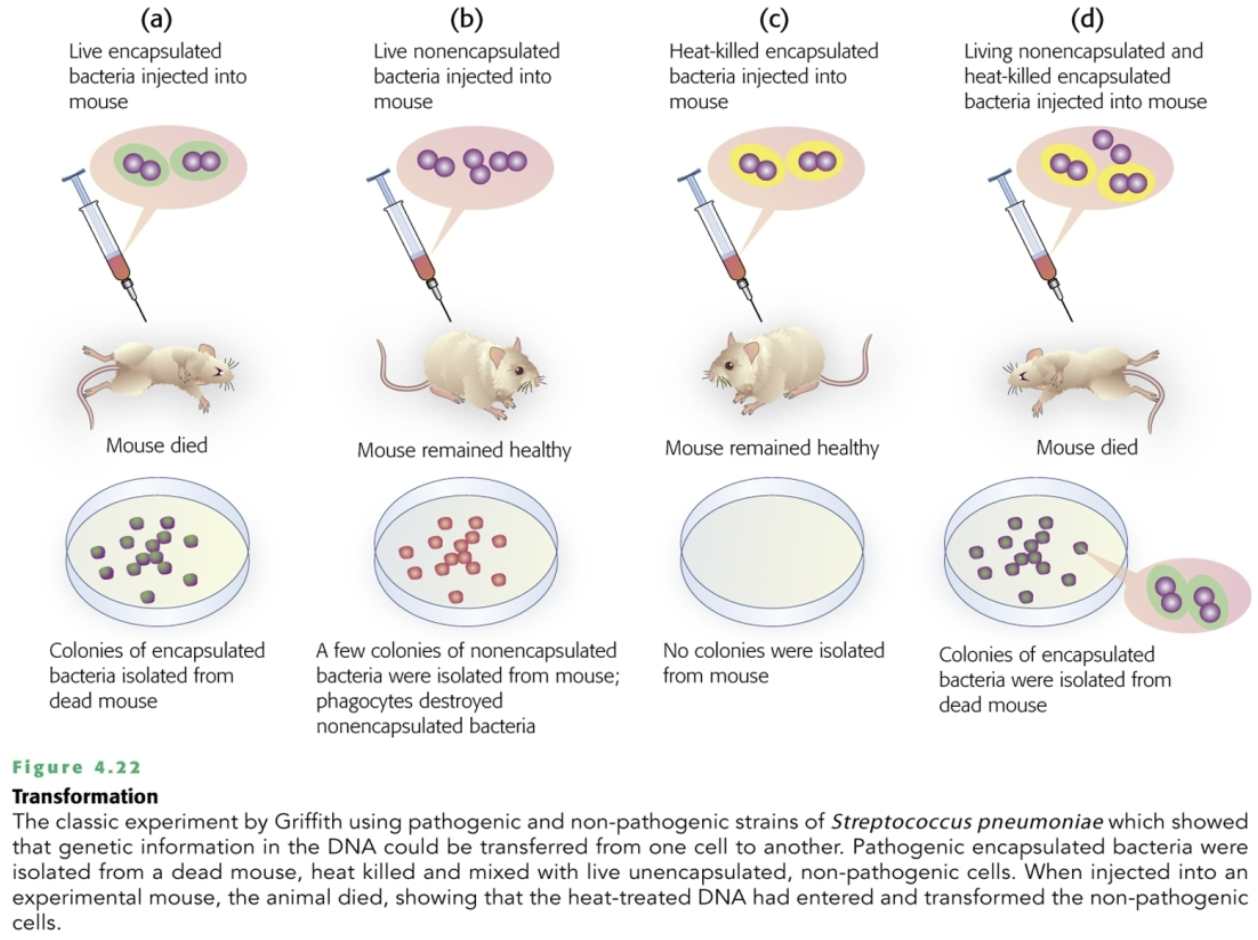
Bacteria can uptake DNA fragments from the environment into their genome (often from dead cells lysis).
Not all bacteria can do this
Usually between cells of the sam genera (Streptococcus, Staphylococcus, Bacillus)
Recognised in Griffith's 1940’s experiments (Streptococcus pneumoniae).
Transduction
Bacteriophages (phages) are viruses that infect bacteria
may induce a lytic cycle
producing phage particles within bacterium and its subsequent lysis releasing the newly formed progeny
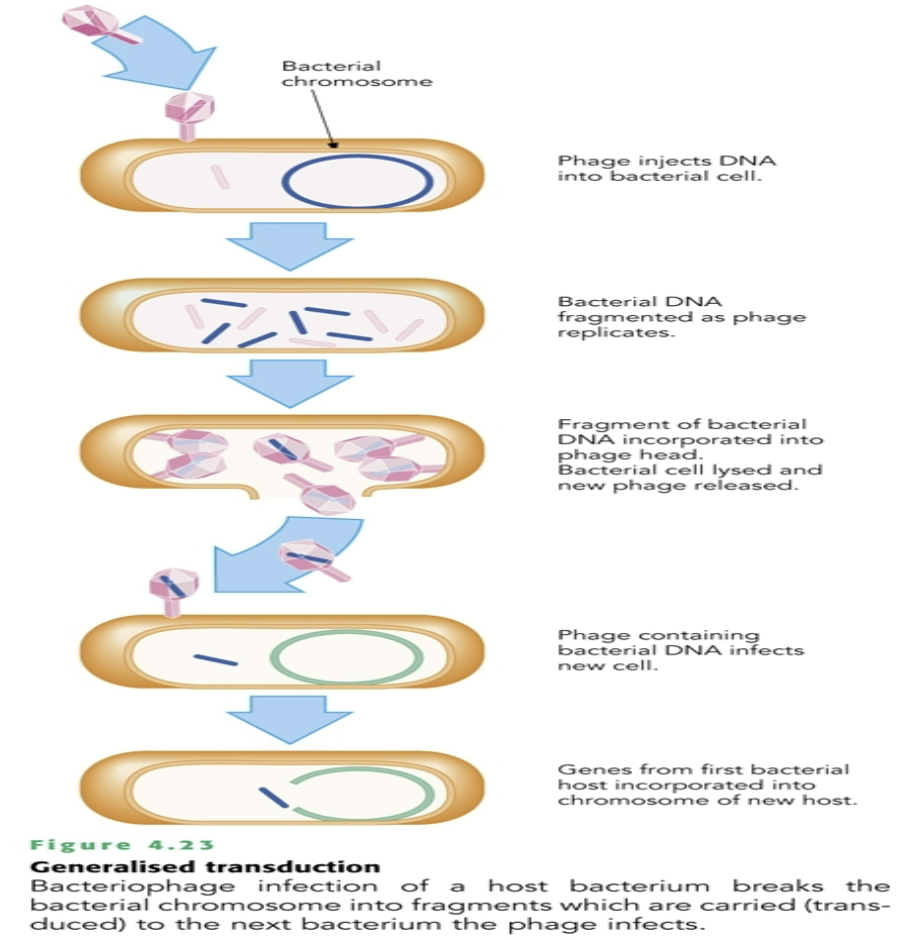
Fragmenting bacterial DNA.
When phage particles are being assembled some fragments of bacterial (donor) DNA may become incorporated into the phage DNA
Subsequent infections of other bacterial (recipient) cells transfers to the DNA
Refer as generalised transduction
Temperate phages DO NOT cause cell lysis BUT incorporate their DNA into bacterial chromosome (prophage)
Replicates with bacterial chromosome (lysogeny)
In some cases the prophage may carry genes for toxin production (Corynebacterium diphtheriae)
Conditions may arise that cause reversion to the lytic cycle
Phage particles assembled and may contain fragments of the bacterial DNA
Fragments will be those adjacent (ชิด) to the prophage in the lysogenic stat
New phage particles infect more bacteria
Referred to as specialised transduction
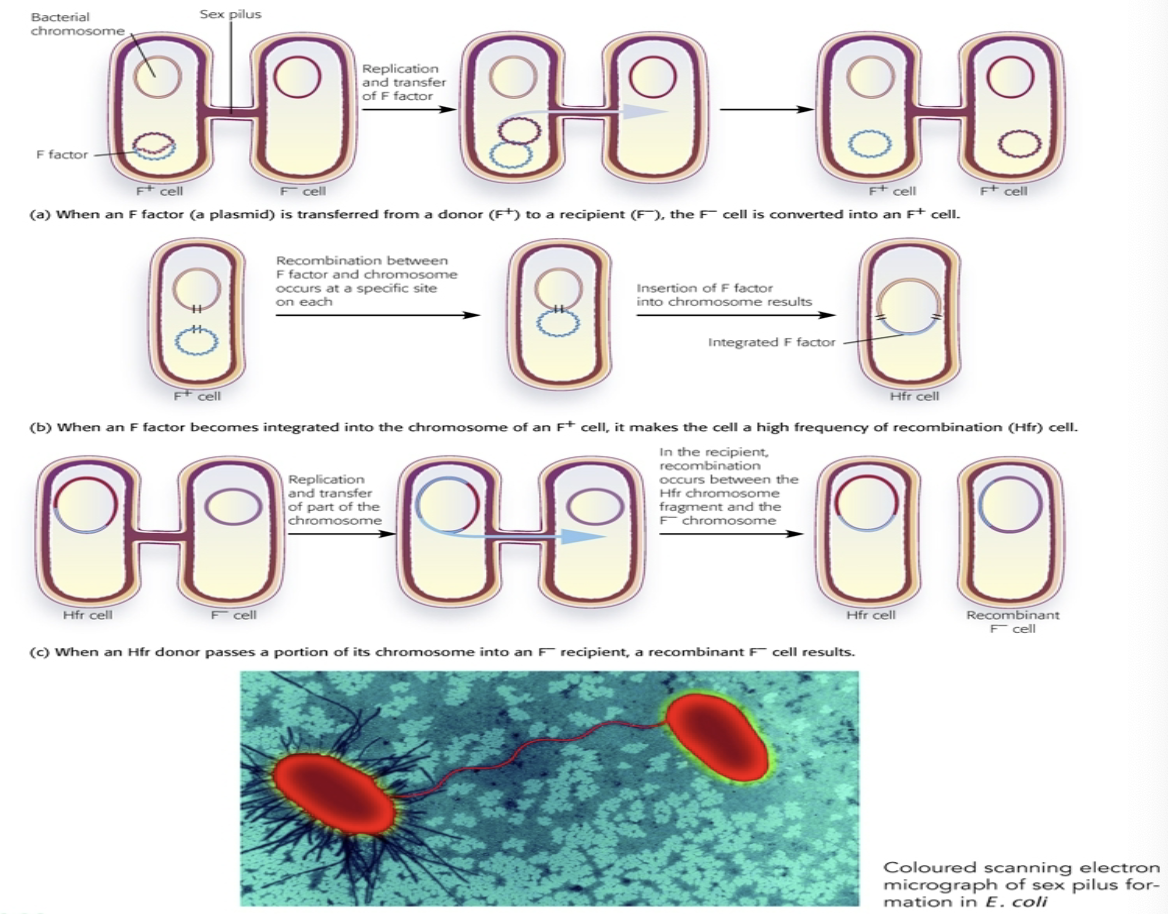
Conjugation
Requires physical contact between donor and recipient calls (unlike transformation and transduction)
Formation of sex pilus between 2 cells, genetic material is transferred via plasmids (may also be a transposon)
F factor plasmids direct sex pilus formation (F+)
F+ may enter recipient cell and continue to replicate independently conferring F+ status on the recipient (F-)
F+ may integrate with recipient chromosomal DNA to generate an Hfr cell
Act as donor cells in conjugation and generate a recombinant F-cell
Mutation
Defined as a permanent, heritable change in the sequence of nucleotide bases in the DNA sequence,
May be large deletions or insertion mutations but most are point mutation
Point mutations affecting a single base pair.
Point Mutations Types
Silent-no visible: no effect due to code degeneracy.
Missense-single base substitution:changing an amino acid.
Nonsense-conversion: sense codon converts to a termination codon.
Frameshift-insertion: deletion of base pairs altering the reading frame.
Origin of Mutations
Spontaneous mutations-mistakes: errors in DNA replication (1:109 cell divisions in bacteria)
Mutagens-substances: that induce mutations through chemical damaging DNA
Radiation-ionising: produces DNA damage through errors in replication produces DNA damage through errors in replication, non-ionising radiation generates thymine dimers
Mutations and Antimicrobial Drug Resistance
Mutations can confer drug resistance to a bacterial cell
Subsequent exposure of cell population to drug creates selective pressure
Susceptible cells are killed and resistant cells survive and multiply
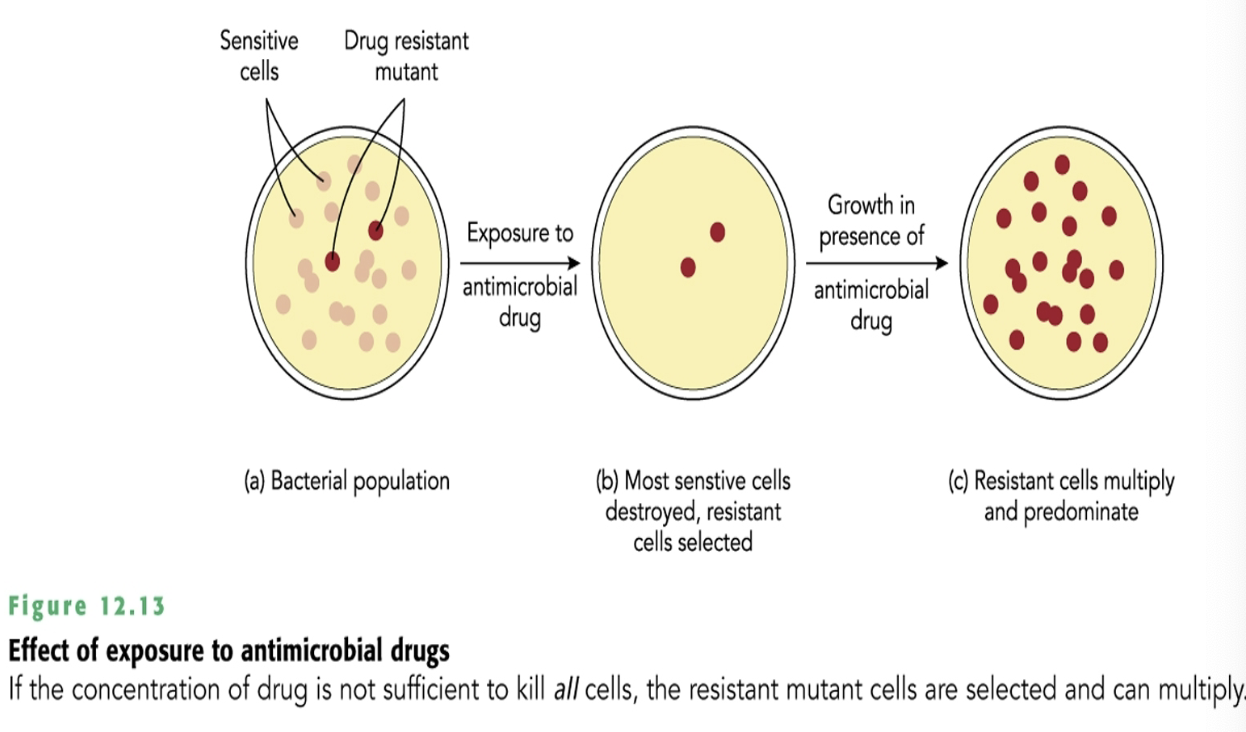
Reasons for Selection of Resistant Cells
Prolonged antibiotic use
Failure to complete treatment
Overuse/misuse of drugs.
Changes Leading to Antimicrobial Resistance (short answers)
1. Production of enzymes that inactivate or destroy the drug
Some bacteria produce enzymes that break down antibiotics, rendering them ineffective.
Example: β-lactamase enzymes degrade the β-lactam ring found in penicillins and cephalosporins, preventing these antibiotics from inhibiting bacterial cell wall synthesis.
2. Alterations in membrane structure and permeability
Bacteria can modify their outer membrane (especially Gram-negative bacteria) to reduce the drug’s ability to enter the cell.
This can involve:
Mutations in porin proteins (small channels in the membrane that allow antibiotics to enter).
Thicker cell walls (as seen in vancomycin-resistant bacteria).
Example: Pseudomonas aeruginosa reduces permeability to carbapenems.
3. Efflux pumps that expel the drug
Some bacteria have efflux pumps that actively transport the drug out of the cell, preventing it from reaching its target concentration.
This mechanism is common for:
Tetracyclines
Fluoroquinolones
Macrolides
Example: The Tet(A) and Tet(B) efflux pumps in tetracycline-resistant bacteria.
4. Alteration of drug binding sites
Bacteria can mutate or modify the target sites where antibiotics usually bind, reducing drug effectiveness.
Examples:
Penicillin-binding proteins (PBPs) altered → resistance to penicillins & cephalosporins (e.g., methicillin-resistant Staphylococcus aureus – MRSA).
Changes in ribosomal binding sites → resistance to aminoglycosides & macrolides (erythromycin).
5. Modification of metabolic pathways
Some bacteria bypass the normal metabolic pathway targeted by the antibiotic by using alternative enzymes or pathways.
Example: Sulphonamide resistance
Sulphonamides inhibit dihydropteroate synthase (an enzyme in folic acid synthesis).
Resistant bacteria produce an alternative enzyme that does not bind to sulphonamides, allowing folic acid synthesis to continue.
Transfer of Drug Resistance
Occurs by plasmids during conjugation, possible across species and closely related genera (e.g., Shigella and E. coli).
Transposons can also carry multiple resistance genes.
Animal feeds contain antibiotics and may contribute to the transfer of drug resistant organisms
Drug Resistance in Hospitals
The extensive use of antimicrobial drugs in hospitals provides a flourishing
Infection control procedures mandatory to prevent spread of resistant organisms to other patients
Drug Resistance in the Community
Caused by Overuse for minor infections
Misuse for viral infections with antibiotics
Alteration of pateint’s normal flora.
Increased resistant strains (e.g., Staphylococcus aureus).