Replication and Recombination
Vocabulary:
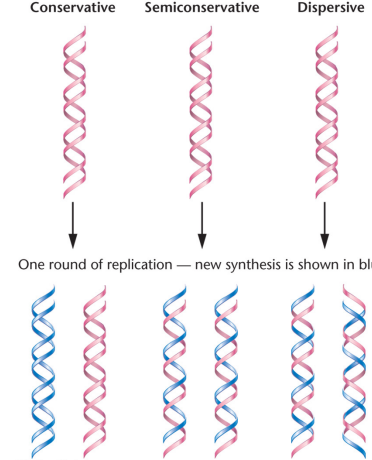
Conservative replication: False model of replication, one original helix was conserved and another completely new one was made from the strands, one old one new
Semiconservative replication: Correct model of replication, describes how one old strand and one new strand are a part of new DNA
Dispersive replication: False model of replication, describes how new DNA strands might have bits and pieces of the parent strands embedded in the new DNA in a patchwork fashion
Meselson and Stahl: In 1968, used two forms of Nitrogen to grow bacteria, who would incorporate them into their DNA. Tested for different nitrogens to see how DNA was split while replicated, found semi conservatively, used equilibrium density gradient centrifugation
Taylor-Woods-Hughes: In 1957 showed DNA replication was semiconservative in eukaryotes, used autoradiography to monitor 3H-Thymidine and the x-rays emitted, found traces in all sister chromatids
Origin of replication (OriC): Where replication begins, one for prokaryotes and many for eukaryotes
Replication fork: Formed by parts of the DNA helix being unwound. Since replication is bidirectional, there are two that migrate apart as replication continues
Ter: Where two replication forks merge and semiconservative replication ends
Replicon: The length of DNA that is replication following one initiation event at a single origin
RNA Polymerase: Uses ribonucleotides to synthesize RNA, adding nucleotides from 5’ → 3’, can initiate synthesis on an open template strand
DNA Polymerase: Uses deoxyribose to synthesize DNA strands, adds nucleotides to the new strand in the 5’ → 3’ direction, can only add bases onto the 3’-OH of an existing strand (cannot initiate). Typically uses Mg2+ cofactors to function
Deoxyribonucleoside triphosphate (dNTP): Used by DNA polymerase as a substrate, one nucleotide
Metal ions: Mg2+ and Mn2+, used to catalyze the addition of new nucleotides as co-factors
Pyrophosphate: Inorganic, cleaved off by DNA polymerase as a nucleotide is added to the 3’ end. 2 phosphates
Primer: An existing DNA strand that may be the beginning point for DNA synthesis elongation
DNA Polymerase III: Can proofread in the 3’ to 5’ direction, responsible for 5’ to 3’ polymerization essential in vivo. If synthesis stalls, will reverse and excise incorrect nucleotides
Fidelity: The accuracy with which polymerases replicate DNA, depends on proofreading abilities
DNA Polymerase I: Synthesizes DNA in the 5’ to 3’ direction, has exonuclease activity in both 5’ to 3’ and 3’ to 5’ directions. Mainly removes and replaces primers. Removes primer and replaces it with DNA. Primarily removes primers
DNA Polymerase II: Synthesizes DNA 5’ to 3’, only can correct 3’ to 5’, for DNA repair and restarting replication after damaged DNA halts synthesis. Primarily fills in gaps
DNA Polymerase III: Synthesizes DNA 5’ to 3’, can only check 3’ to 5’, mainly elongated existing DNA. Primarily replicates DNA
DNA Polymerase IV: Has no exonuclease activity and cannot proofread DNA, only synthesize it in the 5’ to 3’ direction, mainly for DNA repair
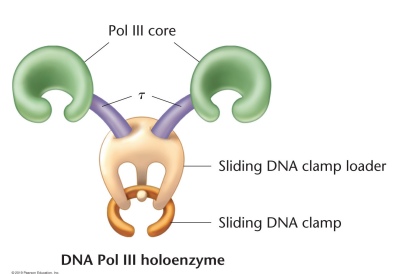
Holoenzyme: Active form of DNA Polymerase III, made up of complex subunits. Alpha for 5’ → 3’ polymerization, epsilon for 3’ to 5’ exonuclease, and theta for core assembly. The y complex loads the enzyme on template and acts as the clamp loader. Beta is the sliding clamp structure, and tau dimerizes core complexes
Sliding clamp loader: Subunit of DNA Pol III, responsible for loading the beta sliding clamps onto the DNA at the site of the primer-template junction, adds nucleotides to the template
Processivity: The strength of an enzyme and how long it can act without having to stall/restart
Beta sliding clamps: Subunit of DNA Pol III, holds polymerase onto the DNA and allows for high-processivity replication, wrap around and orient both leading and lagging strands
dnaA: Initiator protein, binds to OriC to cause a conformational change where the helix destabilizes and opens up, exposing ssDNA. Recruits DNA helicase enzyme. Recognizes a specific pattern on the major groove
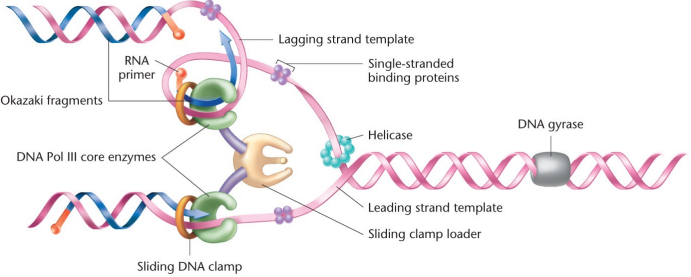
DNA Helicase: Made of dnaB polypeptides, binds to the lagging strand of a replication fork and moves in the 5’ to 3’ direction and breaks the hydrogen bonds between base pairs and moves the replication fork back. Uses energy from ATP hydrolysis. Recruits holoenzyme to bind to the replication fork and initiate replication
DNA gyrase (topoisomerase): Relieves strain ahead of the replication fork before helicase can split it into ssDNA, one of many DNA topoisomerases
Single-stranded binding proteins (SSBPs): Stabilize the open conformation of the helix and bind specifically to single strands of DNA, keep the bubble open and stops DNA from rebinding it itself
Primase (RNA polymerase): Recruited to the replication fork by helicase, synthesizes RNA primer. Doesn’t need a 3’-Oh end but provides it for DNA Pol III to work on elongation
DNA Primase: Synthesizes a short RNA primer to provide a 3’-OH group for the attachment of DNA nucleotides
Replication bubble: Created by two replication forks travelling in opposite directions, starts from one OriC
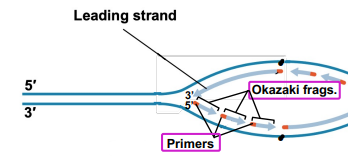
Leading strand: One portion of DNA as it is replicated by DNA Pol in the 5’ to 3’ direction, the same direction as the replication fork, only required one RNA primer
Lagging strand: One portion of DNA is it is being replicated by DNA Pol in the 3’ to 5’ direction, results in Okazaki fragments made discontinuously. Each fragment requires a separate RNA primer
Okazaki fragment: Result of lagging strand, portions of replicated DNA that must be rejoined together, individually made in the 5’ to 3’ direction and each have their own primer
DNA Ligase: Catalyzes formation of phosphodiester bonds, seals nicks and joins together Okazaki fragments with phosphodiester bonds to create the backbone
Concurrent synthesis: When both strands of DNA and synthesized at the same time, the lagging strand is looped. The physical but not biochemical direction is inverted and the DNA sliding clamp prevents core enzyme dissociation from the template
DNA Proofreading: Done by DNA Pol in the 3’ to 5’ direction after incorrect complimentary pairs are added, increased fidelity of synthesis by 100x
Termination: Can occur when two replication forks meet, or when particular DNA sequences (Ter sequences) are present
Initiator protein: Binds to origins and separates strands of DNA to initiate replication in bacterial cells
Autonomously replicating sequences (ARSs): Origins for yeast, 250-400, contain 11-bp consensus sequences flanked by other short sequences involved in efficient initiation, allow any DNA they are attached to to replicate
Prereplication complex (pre-Rc): Assembles at replication origins during the G1 phase
Origin Replication Complex (ORC): A six protein complex that recognizes replication origins in early G1 phases, then tags the origin as the site of initiation, causing the pre-Rc to leave. Distinguishes between segments of DNA that have been replicated and those that haven’t
DNA Polymerase Alpha: DNA polymerase in eukaryotes, A primer for RNA/DNA, initiates DNA synthesis, elongates DNA, low processivity
DNA Polymerase Delta: DNA polymerase in eukaryotes, involved with lagging strand synthesis, repair, recombination, and proofreading, elongates DNA, high processivity
DNA Polymerase epsilon: DNA Polymerase in eukaryotes, involved in leading strand synthesis, repair, recombination, and proofreading, elongates DNA, high processivity
DNA Polymerase Gamma: DNA Polymerase in eukaryotes, involved with mitochondrial DNA replication and repair
Polymerase switching: Happens once a DNA primer is in place, Pol Alpha is replaces by Pol delta or Pol epsilon depending on which strand for elongation
Chromatin: Eukaryotic DNA that is complexed with binding proteins
Histone: Proteins that may have various DNA wrapped inside it, with DNA replication 200 base pair nucleosomes wrap around eight of these
Chromatin assembly factors (CAFs): Carry out assembly, move along with the replication fork
Telomere: The end of a linear eukaryotic DNA sequence, where the nucleotide sequencing is repetitive. Acts protectively, preventing degradation by nucleases or improper repair by cell DNA repair mechanisms. Overall postpone erosion of the genes near the end, though if replicated enough will run out. Flags DNA as the end, and not a DSB
Double-stranded breaks (DSB): What occurs when a chromosome is broken internally due to DNA damage
Telomerase: Produced by germ cells, an enzyme that adds telomeres to DNA, specifically in the sperm/egg cells that germ cells produce. This enzyme is turned off in somatic cells, as turning it on may result from cancer
Recombination: The process whereby genetic information is exchanged between two DNA molecules at equivalent positions along two chromosomes. Important in DNA repair, generating diversity, and gene conversion
S-checkpoint: Part of mitosis, where the genome is checked for errors
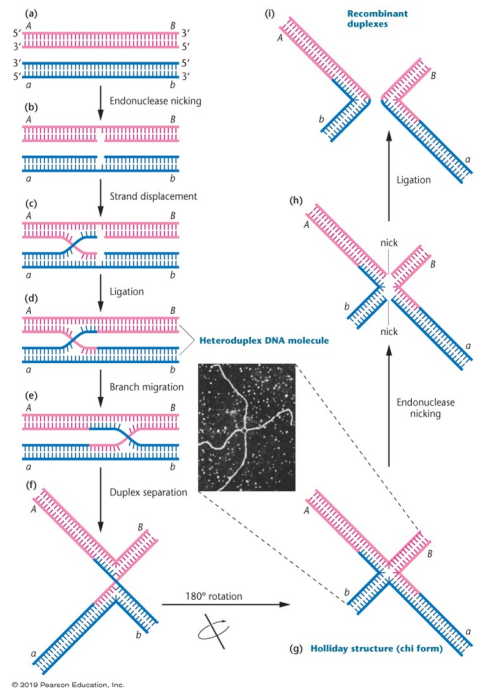
Holliday Model: Model of recombination, simple and doesn’t involve DNA synthesis. Has two homologs, internal strand endings are displaced and paired with complements on opposite duplexes. Ligase seals the loose ends, making a hybrid duplex
Heteroduplex DNA molecule: Part of the Holliday model of recombination, results from ligase sealing the ends of opposite duplexes, held together by a bridge structure
Double-strand break model: Complex model of recombination
Branch migration: Part of the Holliday model, when the cross bridge moves down the chromosome, as hydrogen bonds are broken and reformed
In eukaryotes, transcription occurs at multiple start points, allowing for the genome to be replicated in only minutes to a couple of hours. In Prokaryotes, there is only one starting location. Bacteria have a singular circular chromosome
The human genome has ~6.4 billion base pairs, duplication of the DNA is important for mitosis used in growth and repair, around 1000 base pairs replicated a second
DNA Polymerases I, II, and III all contain 3’ to 5’ exonuclease activity allowing them to proofread newly synthesized DNA, remove, and replace incorrect nucleotides. DNA Polymerase I additionally possesses this in 5’ to 3’ direction
Eukaryotic polymerases required four deoxyribonucleoside triphosphates, a template, and a primer. There is also more DNA, with linear chromosomes and heavier use of proteins
Every time a chromosome is copied/replicated, a little bit of the DNA at the ends is lost at the end of the lagging strand where the RNA primer was. The 5’ ends of daughter DNA strands is incomplete as there is no 3’ ends for DNA polymerase to add on to. Only a eukaryotic problem
Phosphodiester bonds are created by nucleophilic attacks by the activated 3’-hydroxyl on the phosphate of the incoming dNTP resulting in the formation of a new phosphodiester bond and release of inorganic phosphate (PPi). Metal ions are important in this process