07-Biochemistry-Lecture7 | Week 4 - Lecture 2
LECTURE 7: DNA REPLICATION
DNA METABOLISM
Definition: DNA metabolism refers to a set of tightly regulated processes that maintain the integrity, stability, and expression of genetic information.
Overview of Processes:
DNA Replication: A new copy of DNA is synthesized before each cell division with high fidelity.
DNA Repair: Errors arising during or after DNA synthesis are constantly checked for and repaired.
DNA Recombination: Segments of DNA are rearranged either within one chromosome or between two DNA molecules, providing genetic diversity to offspring.
Summary: DNA serves as the substrate that encodes its own metabolism.
OUTLINE OF REPLICATION
Main Topics:
General principles of replication
Structure and reaction of polymerases
Key concepts: processivity, error rates, proofreading
Stages of Replication:
Initiation, Elongation, Termination
Comparison: Prokaryotic vs Eukaryotic processes
DNA REPLICATION FEATURES
Three Fundamental Rules of Replication:
Semi-Conservative: Each new DNA molecule consists of one old strand and one new strand.
Bi-Directional: Replication begins at an origin and proceeds in two directions.
Directionality: Synthesis of new DNA occurs in the 5’ → 3’ direction and is semi-discontinuous.
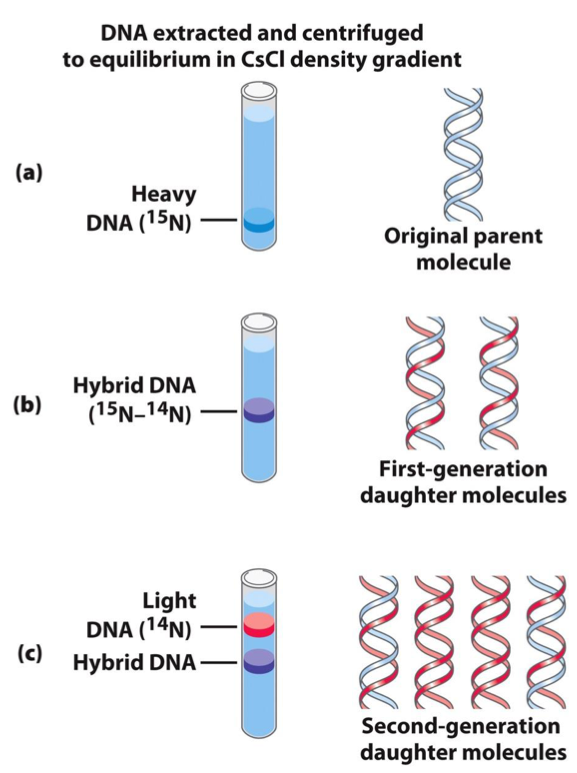
THE MESELSON-STAHL EXPERIMENT
Purpose: Demonstrated that DNA replication is semiconservative.
Methodology:
Cells were grown in a medium containing heavy nitrogen (15N) so that their DNA contained only 15N.
After transferring cells to light nitrogen (14N) for one generation, DNA was isolated and centrifuged in a CsCl gradient, yielding a hybrid DNA band indicative of semiconservative replication.
Results: The first generation showed a single hybrid band; the second generation showed one hybrid and one band of lighter 14N DNA, confirming semiconservative replication.
REPLICATION OF CIRCULAR DNA
Bidirectional Nature: DNA replication begins at an origin and usually progresses bidirectionally.
Visualization of Bidirectional Replication:
Stages are visualized by electron microscopy, producing a theta (θ) structure as both strands are replicated simultaneously.
Electron micrographs illustrate replication of plasmid DNA from a single origin.
SEMI-DISCONTINUOUS SYNTHESIS
Replicative Strands at the Fork:
A new strand is always synthesized in the 5’ → 3’ direction, reading the template in the opposite direction (3’ → 5’).
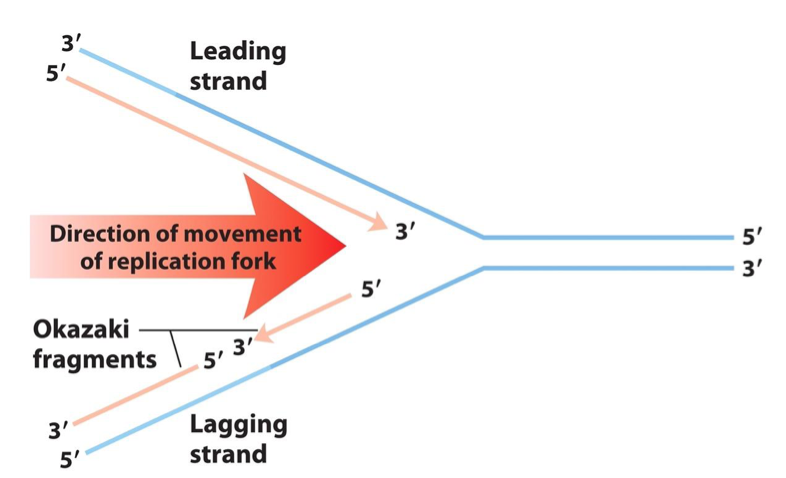
Leading Strand: Continuously synthesized in the direction of the replication fork.
Lagging Strand: Synthesized discontinuously in short pieces (Okazaki fragments) in the opposite direction to the fork's movement.
Okazaki Fragment Lengths: In bacteria, 1000 to 2000 nucleotides; in eukaryotes, 150 to 200 nucleotides.
STRUCTURAL FEATURES OF DNA POLYMERASES
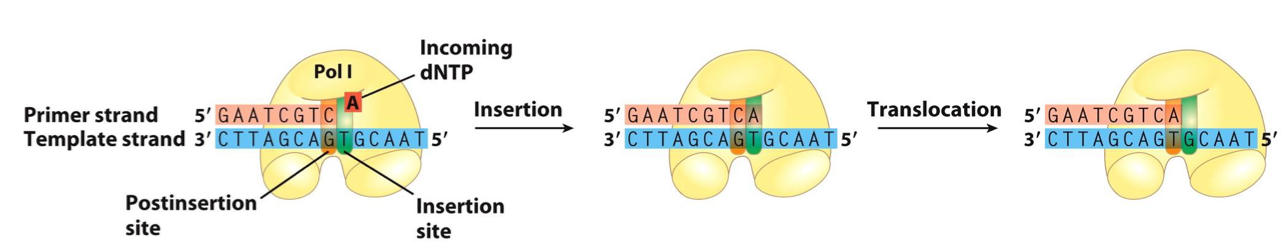
Core Structure: A DNA polymerase has a pocket with two regions:
Insertion Site: Where the incoming nucleotide binds.
Post-Insertion Site: Where the newly made base pair resides after the polymerase moves forward.
Shape: Most DNA polymerases resemble a human hand that wraps around the active site, as seen in DNA Polymerase I from Thermus aquaticus.
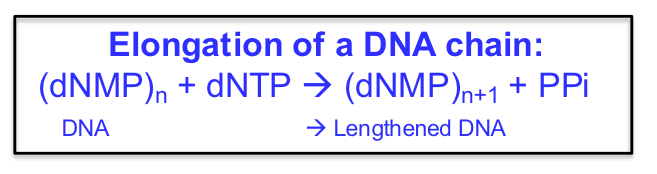
DNA ELONGATION CHEMISTRY
Template: Parental DNA strand serves as a template for elongation.
Substrates: Nucleoside triphosphates (dNTPs) are used as substrates in strand synthesis.
Mechanism of Nucleotide Addition: The nucleophilic OH group at the 3’ (the primer) end of the growing chain attacks the α-phosphate of the incoming trinucleotide.
This 3’-OH is REQUIRED
The 3’-OH is made a more powerful nucleophile by nearby Mg2+ ions
Pyrophosphate is a good leaving group
Reaction Representation:
MECHANISM OF ACTION OF DNA POLYMERASES
Catalytic Mechanism: Involves two Mg²⁺ ions and three highly conserved Asp residues.
Function of Mg²⁺ Ions: Facilitate attack by the 3’-hydroxyl group on the α phosphate and displace pyrophosphate (PPi).
Processivity of Polymerase: Refers to the number of dNTPs added before the polymerase dissociates from the template.
Requirements for Activity: Template, primer, and dNTPs are required.
FEATURES AND IMPORTANCE OF A PRIMER
Primer: A short strand complementary to the template that serves as a starting point for DNA synthesis.
Contains a free 3’-OH to begin the first DNA polymerase-catalyzed reaction
Can be made of DNA or RNA
3’-OH Group: Essential for DNA polymerase-catalyzed reactions, allowing the addition of nucleotides to the growing strand.
Base-Pairing: Each incoming nucleotide is selected based on base-pairing with the template strand. This insertion occurs in the insertion site.
The newly formed base pair migrates to the post-insertion site to make the active site available to the next pair to be formed.
CONTRIBUTION OF BASE PAIR GEOMETRY TO REPLICATION FIDELITY
Base Pair Geometry: Proper A=T and G≡C pair geometries allow them to fit into the polymerase active site. Incorrect pair geometries can be excluded from the active site, reducing errors.
HIGH FIDELITY OF DNA SYNTHESIS
Error Rates of DNA Synthesis:
Geometry of base pairing & proofreading accounts for high fidelity
DNA polymerase active site excludes base pairs with incorrect geometry, & it also has a proofreading activity.
DNA polymerase adds nucleotides with an error rate of 1 in 10⁵ (because of tautomerism → H-bonding to incorrect partner).
Proofreading still does not correct errors every 1/10² bases
Proofreading (3’ → 5’ exonucleolytic): 1 in 10².
Strand-directed mismatch repair: 1 in 10³.
Combined Rate: Approximately 1 error in 10¹⁰ bases replicated.
ERRORS DURING SYNTHESIS AND CORRECTION
Proofreading Mechanism: Most DNA polymerases have a 3’ → 5’ exonuclease activity that corrects mismatched base pairs.
Translocation Process: The enzyme is inhibited from moving to the next nucleotide until the incorrect nucleotide is removed.
Error Correction Process: The enzyme slides the DNA backward into the exonuclease site to correct mistakes.
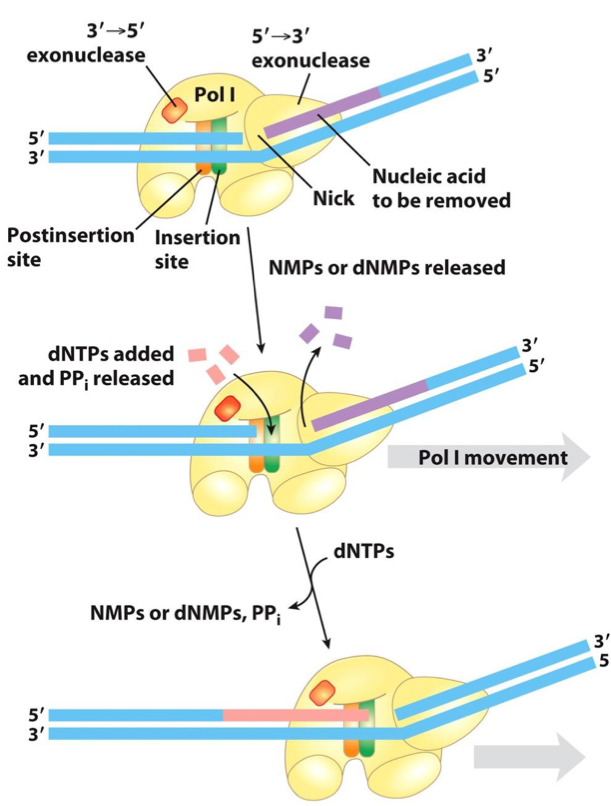
ERROR CORRECTION BY 3’→ 5’ EXONUCLEASE ACTIVITY OF DNA POLYMERASE I
The exonuclease activity is located behind the polymerase activity as the enzyme is oriented in its movement along the DNA.
A mismatched base impedes translocation of DNA polymerase I to the next site.
The DNA bound to the enzyme slides backward into the exonuclease site, and the enzyme corrects the mistake with its 3’→5’ exonuclease activity.
The enzyme then resumes its polymerase activity in the 5’→3’ direction.
Proofreading activity of Pol I improves the accuracy of polymerization by 102 to 103 fold.
KEY TERMINOLOGY FOR POLYMERASES
Fidelity: The error rate in the 5’ to 3’ synthesis.
Proofreading: The capability to correct synthesis errors.
Processivity: The number of nucleotides added before dissociation from the template.
Rate: The speed of nucleotide addition.
DNA POLYMERASE DIVERSITY IN E. COLI
DNA Polymerase I: Abundant but slow (10-20 nucleotides/sec) with low processivity (3-200 bases) → Mainly involved in clean-up.
DNA Polymerase III: Principal replication polymerase with high rate and processivity.
DNA Polymerases II, IV, and V: Involved mainly in DNA repair.
COMPARISON OF E. COLI DNA POLYMERASES
Role of Pol II, IV, and V: Primarily focused on DNA repair activities.
NICK TRANSLATION AND MULTI-FUNCTIONAL DNA POLYMERASE I
5’ → 3’ Exonuclease Activity: DNA Polymerase I also has this activity, which removes nucleotides in the direction of movement, facilitating nick translation.
DNA polymerase I degrades the DNA ahead of the enzyme (in 5’ → 3’ direction), thus removing nucleotides on its path.
Klenow Fragment: A distinct domain that separates from DNA Polymerase I, but lacks the 5’ → 3’ exonuclease activity.
comprises the 5’→3’ polymerase activity & the 3’→5’ proofreading exonuclease activity.
NICK TRANSLATION MECHANISM
Mechanism Description: Movement of the nick along with the enzyme due to its polymerase and exonuclease activities allows for removal of RNA primers during replication.
Bacterial DNA polymerase I has three domains, which catalyze: DNA polymerase activity, 5’→3’ exonuclease activity, and 3’→5’ exonuclease activity
The 5’→3’ exonuclease domain is positioned in front of the enzyme as it moves along DNA.
DNA polymerase I promotes a reaction called nick translation by:
Degrading the DNA strand ahead of the enzyme
Synthesizing a new DNA strand behind the enzyme
During nick translation, a break (nick) in the DNA is effectively moved along with the enzyme.
Nick translation plays a role in: DNA repair & Removal of RNA primers during DNA replication.
A nick remains after DNA polymerase I dissociates. This remaining nick is later sealed by another enzyme.
EUKARYOTIC DNA POLYMERASES
Diversity in Eukaryotes: 15 DNA polymerases present in eukaryotes, such as:
DNA Pol α: Extends primers initiated by primase.
DNA Pol δ: Used in the lagging strand.
DNA Pol ε: Functions in the leading strand.
DNA Pol β and others: Used for repair purposes.
DNA Pol γ: Responsible for mitochondrial DNA replication.
INITIATION OF REPLICATION
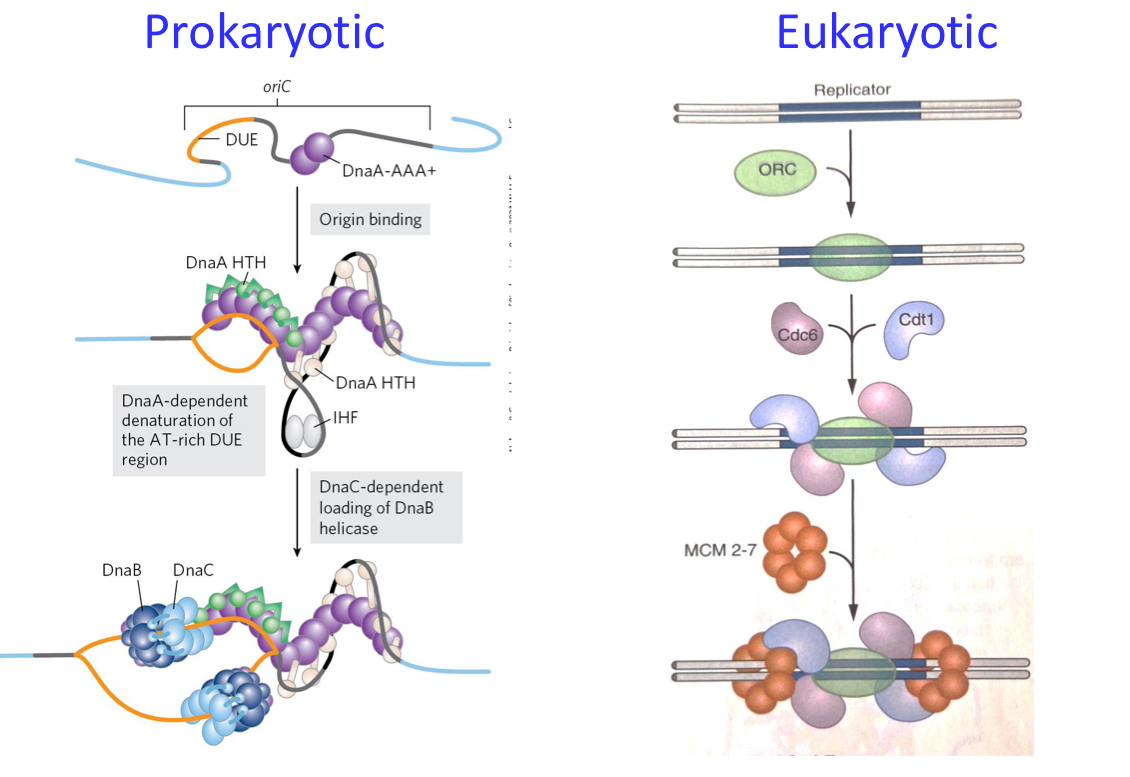
PROKARYOTIC INITIATION
Single Origin (oriC): Recognized by DnaA.
DnaA Role: Induces melting at DUE (DNA Unwinding Element) due to high AT content.
Other Proteins Required: DnaB (helicase), DnaC (helicase loader), clamp loader for polymerase, and SSB (single-strand binding protein).
Primase makes first primers.
Pol III starts synthesis of DNA
Only 1 replicstion/cylce:
DnaA ATP/ADP hydrolysis
SeqA sequestration
EUKARYOTIC INITIATION
Multiple Origins (1000): Recognized by the origin of replication complex (ORC).
Helicase Loading: MCM2-7 opens the site, cyclin signaling induces changes, RFC acts as a clamp loader and RPA stabilizes ssDNA.
Primase & DNA pol α make first primers.
DNA Pol ε & δ start synthesis of DNA.
Only 1 replication/cycle: sites licensed before S phase.
DNA POLYMERASE III (POL III)
Architecture: Composed of three core domains and a five-subunit clamp-loading complex. Maintains high processivity and interaction with helicase.
Functionality: Provides effective coordination during lagging and leading strand synthesis, crucial for overall replication speed and fidelity.
ELONGATION STAGE OF REPLICATION
PROKARYOTIC ELONGATION
Process Complexity: Involves multiple proteins including DNA polymerases, primase, and helicase.
Speed: Fast → Approximately 500 bp/s per polymerase, with a total replication time of about 40 minutes for a typical bacterial genome.
Primer Processing: RNA primers are removed by DNA Polymerase I.
EUKARYOTIC ELONGATION
Replication Duration: Slower than prokaryotic elongation, taking 6-8 hours to replicate the human genome.
Speed: Approximately 50 bp/s per polymerase.
Primer Removal: Carried out predominantly by Dna2 and FEN1.
COMPLEXITY OF DNA REPLICATION ENZYMES
E. coli requires over 20 enzymes and proteins for DNA replication
A set of proteins involved in replication, called the replisome, includes:
DNA-binding proteins to stabilize separated strands
Helicases (use ATP to unwind DNA strands)
Primases (make RNA primer)
DNA polymerases I and III (primer removal and DNA elongation)
DNA ligases to seal nicks
DNA gyrases (also called DNA topoisomerases II; relieve the stress caused by unwinding)
Eukaryotes require far more proteins, but similar activities
TOPOISOMERASE FUNCTION
Winding Problem: During replication movement, the ahead parent DNA helix must rotate to prevent overwinding, which could halt replication.
Role of Topoisomerases: Generating temporary single-strand breaks that relieve tension by allowing strand rotation, thus enabling continuous replication without halting.
LAGGING STRAND SYNTHESIS STEPS
Primer Removal and Replacement: Conducted by DNA polymerase I using its 5’ → 3’ exonuclease activity; remaining nicks are sealed by DNA ligase.
Nick Sealing Mechanism: 5’-PO4 must be activated before nucleophilic attack by 3’-OH, resulting in complete strand integrity.
GENERAL FEATURES OF THE REPLICATION FORK
The DNA duplex is unwound by: DNA gyrase & Helicase
Single-stranded regions are coated by SSB (single-strand binding proteins)
Primase periodically adds RNA primers.
The two sliding clamps of DNA polymerase III replicate the strands.
DNA polymerase I and DNA ligase act downstream on the lagging strand to:
Remove RNA primers and replace them with DNA (Pol I)
Ligate (join) the Okazaki fragments (Ligase)
TERMNATION OF REPLICATION
PROKARYOTIC TERMINATION
Usage of Ter Sequences: Bind Tus protein to signal replication ends; resolution of catenated circles occurs via topoisomerases.
EUKARYOTIC TERMINATION
Mechanism: Replicative forks meet and fuse, with unreplicated ends filled by telomerase, maintaining genomic stability through generations.
Termination of replication in E. coli
The Ter sequences (TerA through TerF) function as a trap for the replication fork.
Ter sequences bind Tus proteins (Terminus Utilization Sequence).
Topoisomerases are required in replication termination.
When opposing replication forks meet:
Replication leaves the completed chromosomes joined as catenanes (topologically interlinked circles).
Because they are covalently closed, they cannot be separated without topoisomerases.
In E. coli, a type II topoisomerase (DNA topoisomerase IV):
Plays the primary role in separating catenated chromosomes
Transiently breaks both DNA strands of one chromosome
Allows the other chromosome to pass through the break