Wk 5.2 Biosynthetic Secretory Pathway
Introduction and Objectives
Biosynthetic Secretory Pathway Overview
Focus on the transport of secretory and organelle proteins across the endoplasmic reticulum (ER) membrane.
Understand the biosynthetic secretory pathway processes, including sorting, compartments traversed, and post-Golgi sorting/maturation.
ER Membrane Binding
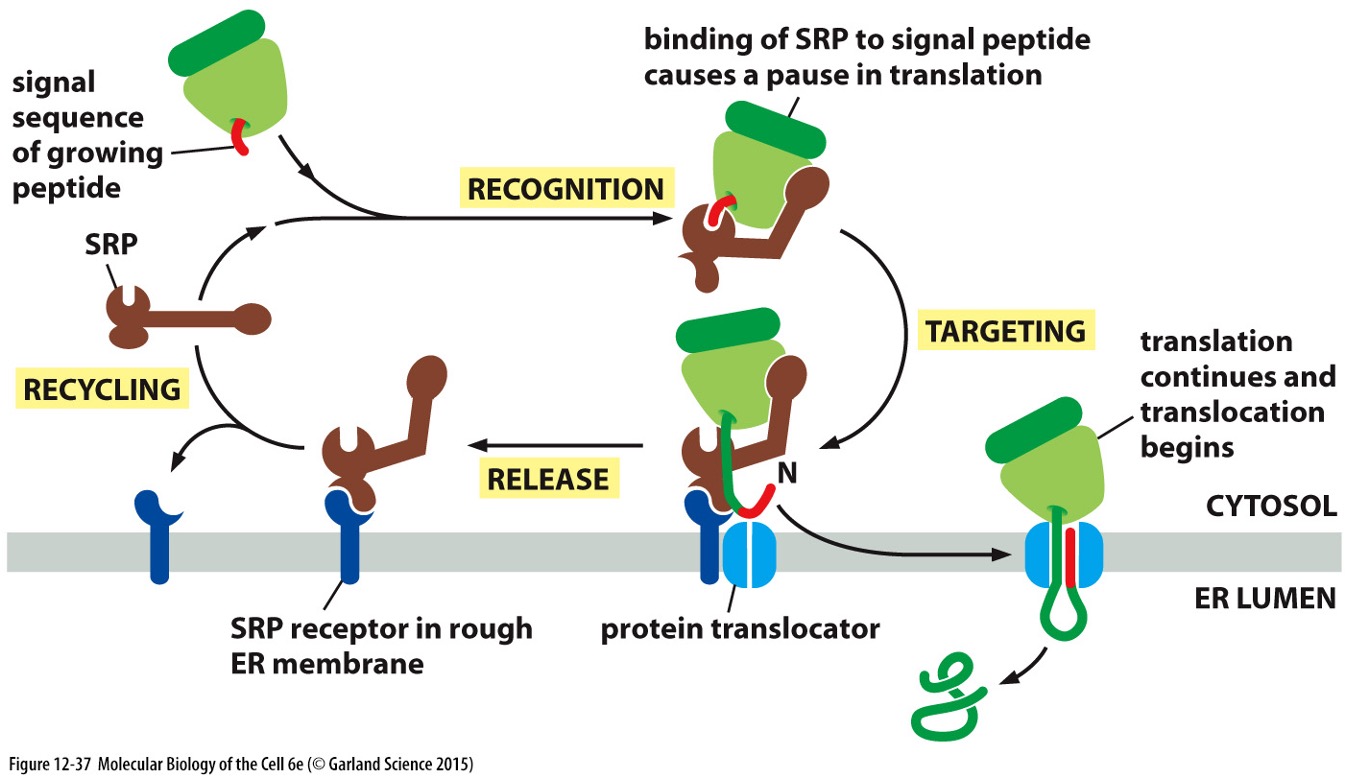
Ribosome Binding Process (SRP Cycle)
Signal Sequence: Part of the growing peptide recognised by the Signal Recognition Particle (SRP).
SRP Binding:
SRP binds to the signal peptide, causing translation to pause.
Targeting:
SRP connects to the SRP receptor in the rough ER membrane.
Translation Resumption:
Translation resumes, and the polypeptide is translocated into the ER lumen.
Protein Sorting Signals
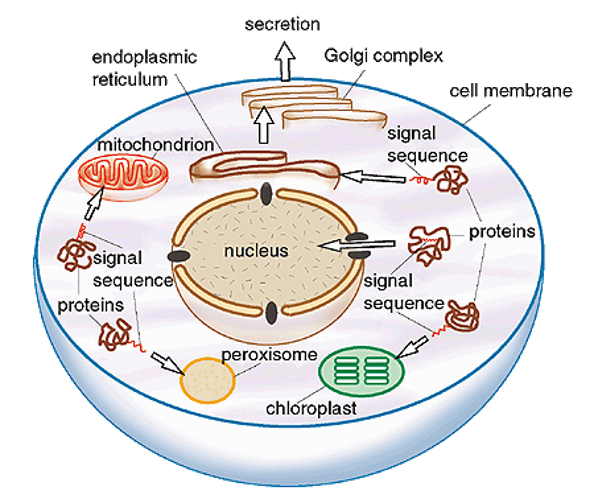
Signals and Receptors
Proteins contain sorting signals (15-60 amino acids, often at N-terminus) that direct them to the correct cellular location.
Cleaved by signal peptidase following delivery
Signals may include internal sequences that form specific configurations (e.g., nuclear localisation signals). aka. signal patch
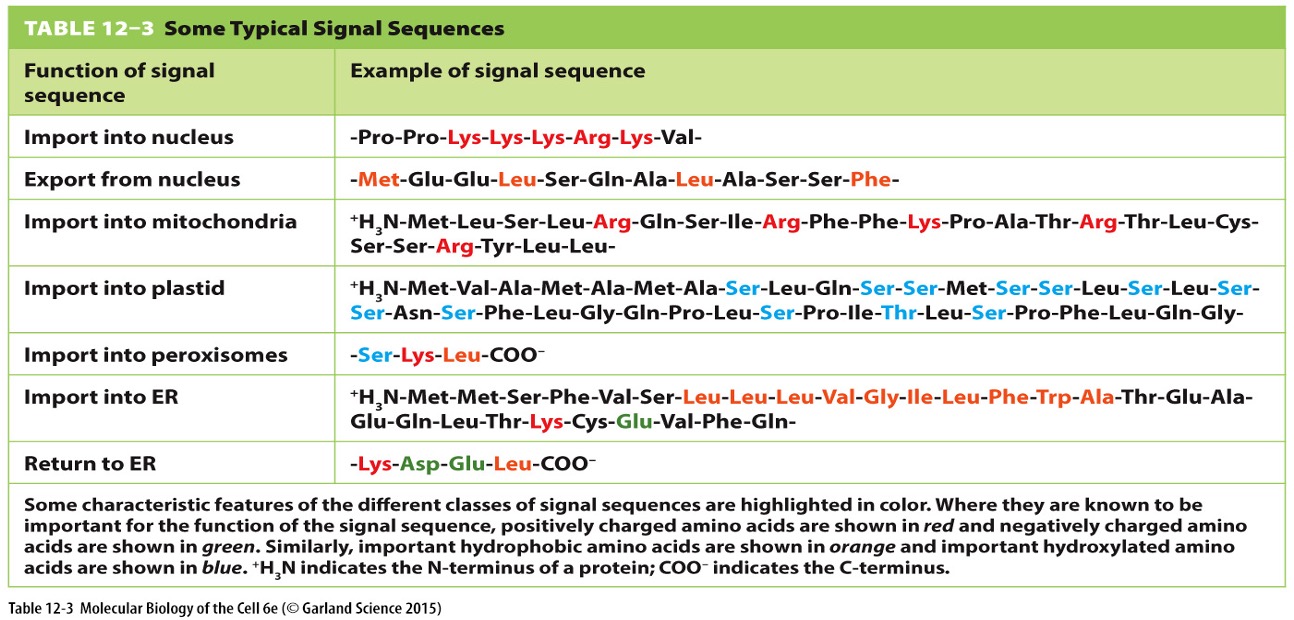
Protein Characteristics
Hydrophobicity/charge of the sequence often determines sorting, rather than exact amino acid composition.
Necessary and sufficient for targeting — switch signals, redirect protein

Secretory Pathway Characteristics
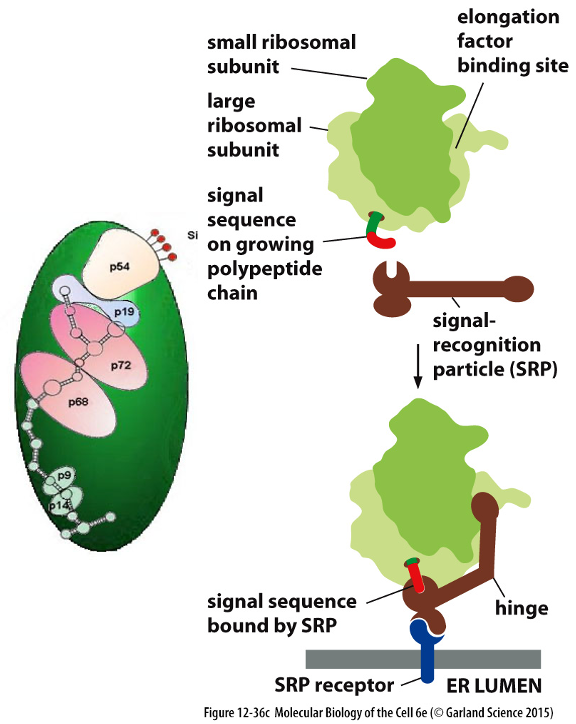
SRP Mechanism:
Signal Recognition Particle (SRP-protein complex) directs proteins to the ER membrane based on the presence of an ER signal sequence.
SRP binds to large ribosomal subunit (signal sequence binding pocket binds near nascent chain exit site on ribosome)
Protein Translocators (Sec61 Complex):
Forms a water-filled pore in the ER membrane allowing polypeptide chains to pass through.
Can open in two ways: by moving a “plug” out of way or opening sideways via a seam
Signal sequence is released into the membrane after opening of seam
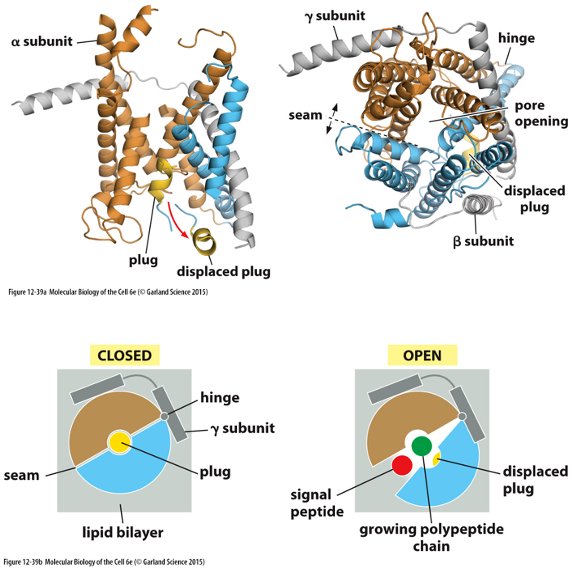
The Translocator Opens by Binding the Signal Sequence
Organelle and secreted proteins transported to final location
Secreted proteins enter biosynthetic-secretory pathway
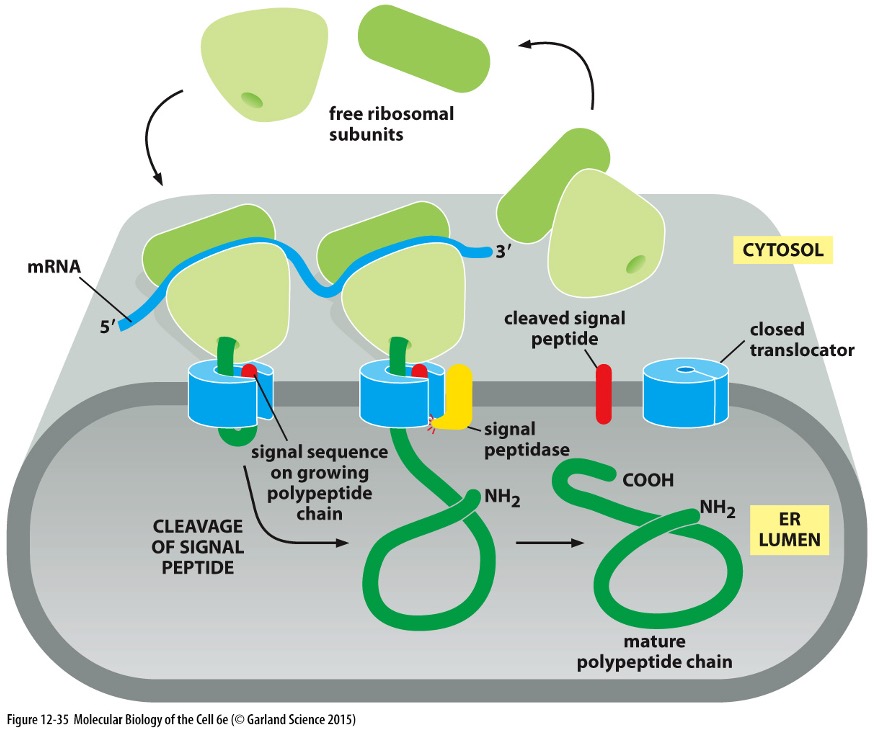
Post-Translational Modifications in ER
Folding and Assembly:
Correct protein folding occurs in the ER; proteins assemble into multimeric forms.
Disulphide bonds formed in the ER stabilise protein structure.
Oxidative linkage of sulphydryl (-SH) groups — a spontaneous reaction (sufficient oxidant)
Proper pairing Cys is essential for protein structure/ activity
Also catalysed by protein disulphide isomerase (PDI)
Form specific order » small domains first, then distant segments
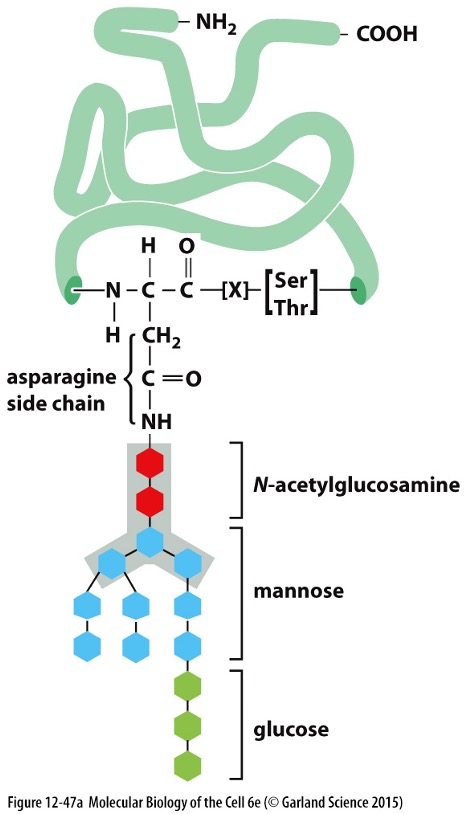
Glycosylation:
Most membrane and secretory proteins undergo glycosylation in the ER and Golgi apparatus, adding oligosaccharides.
Important for protein stability, targeting, and function.
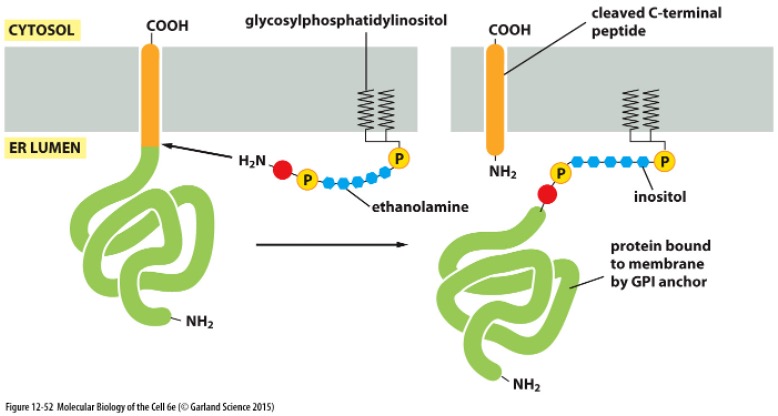
Special glycosylation » GPI anchor
Some proteins acquire a covalently attached glycosyphosphatidyl-inositol (GPI) anchor
Added to C-terminus
Marks proteins for membrane insertion
These proteins are only attached by GPI anchor
Allows for soluble extracellular release by cells
Biosynthetic Secretory Pathway
ER » Golgi apparatus » destination (e.g. lysosomes, cell surface)
Only proteins that are properly folded and assembled can leave the ER
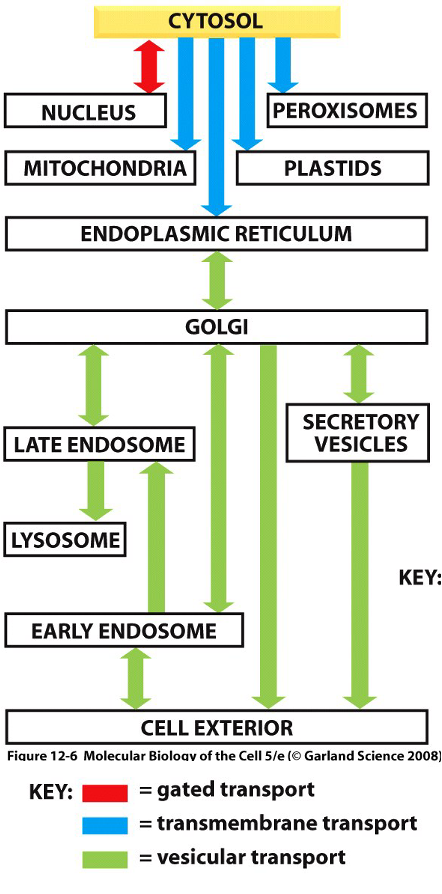
Proteins Can Moe Between Compartments in Different Ways
Gated transport
Between nucleus and cytoplasm
Nuclear pore complexed (nuclear pore envelope)
Selective ‘gate’, not free diffusion
Transmembrane transport
Protein translocators move proteins from cytoplasm to topologically distinct areas (e.g. mitochondria, ER)
Unfolded to across
Vesicular transport
Membrane enclosed transport intermediates (vesicles) move proteins between compartments
Loaded in lumen, bud from membrane, fuse to second compartment
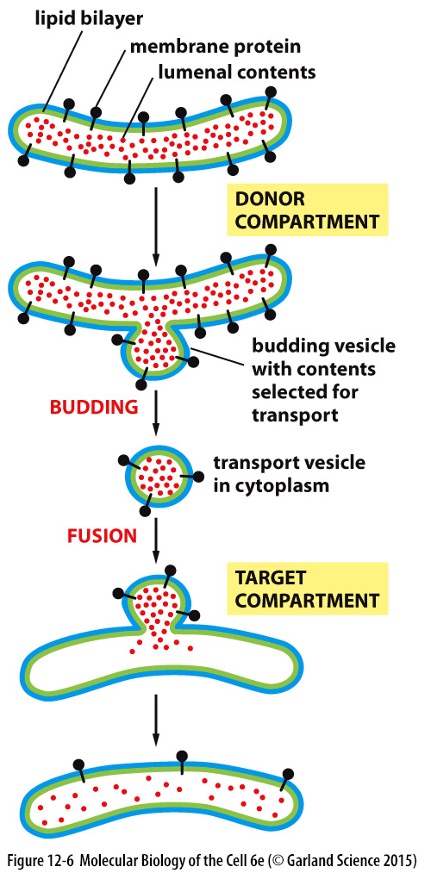
How Do Proteins Exit the ER?
Proteins in lumen of first compartment
Membrane bulges out » ’buds’ off
Proteins packaged into vesicles
Vesicle fuses with target compartment » releases content
Vesicular Tubular Clusters
After transport vesicles have budded they fuse – vesicular tubular clusters
Move along microtubules to carry proteins from ER to Golgi apparatus (bud off their own transport vesicles which carry escaped resident proteins and proteins involved in budding)
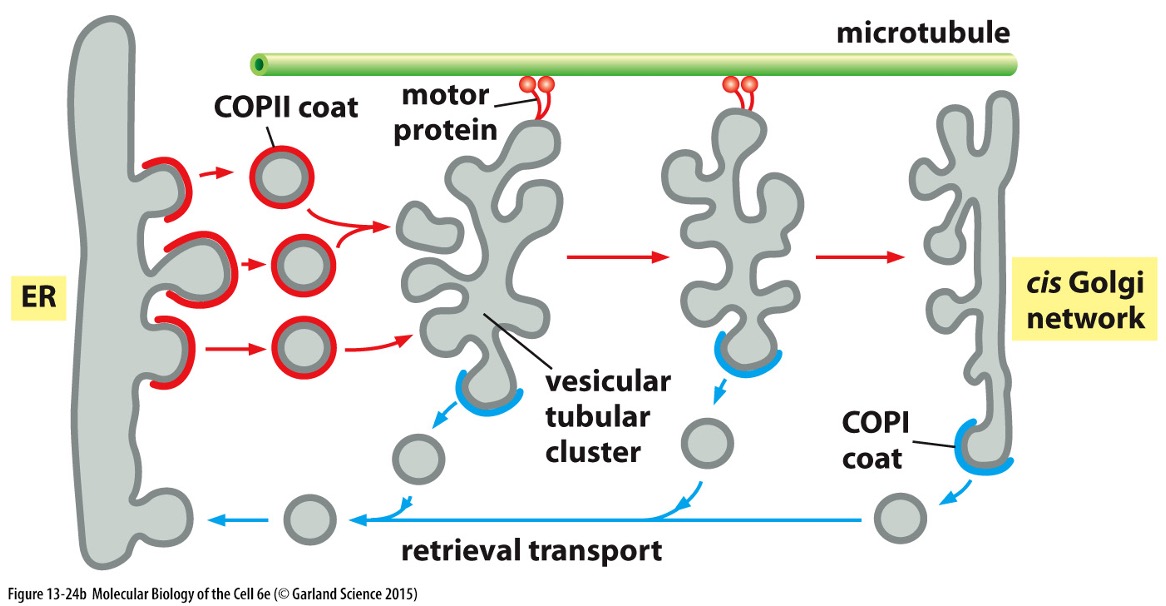
Direction depends on coating
COPII » ER to Golgi
COPI » retrieval transport
Golgi Apparatus Functions
Structure:
Comprises a series of flattened vesicles with cis, medial, and trans regions.
Proteins arrive from the ER and continue to undergo post-translational modifications.
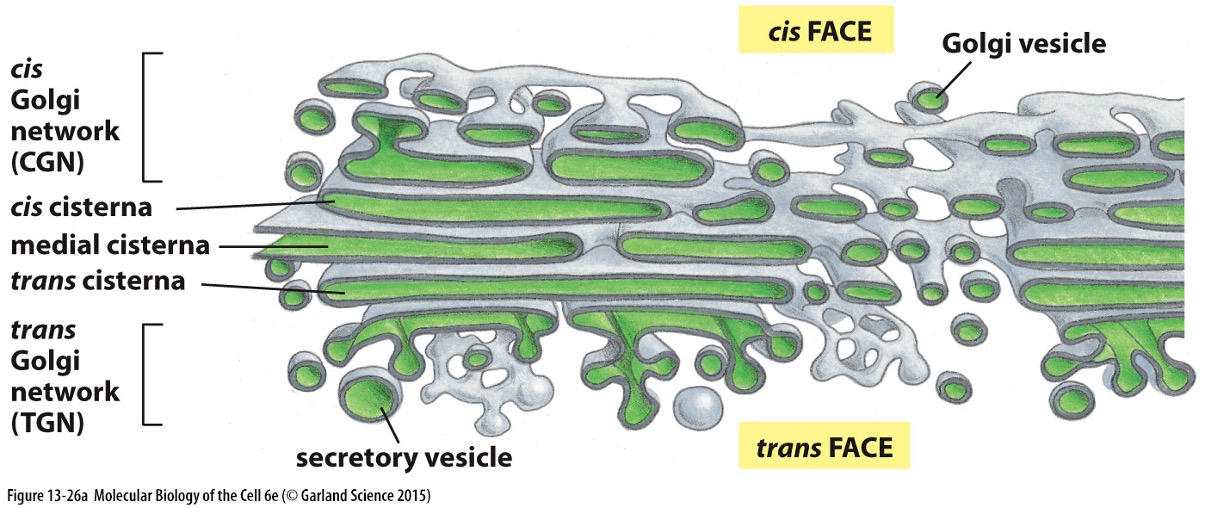
Further modification of oligosaccharide chains occurs during transport through Golgi
Each cisternae have different enzymes which modify proteins differently
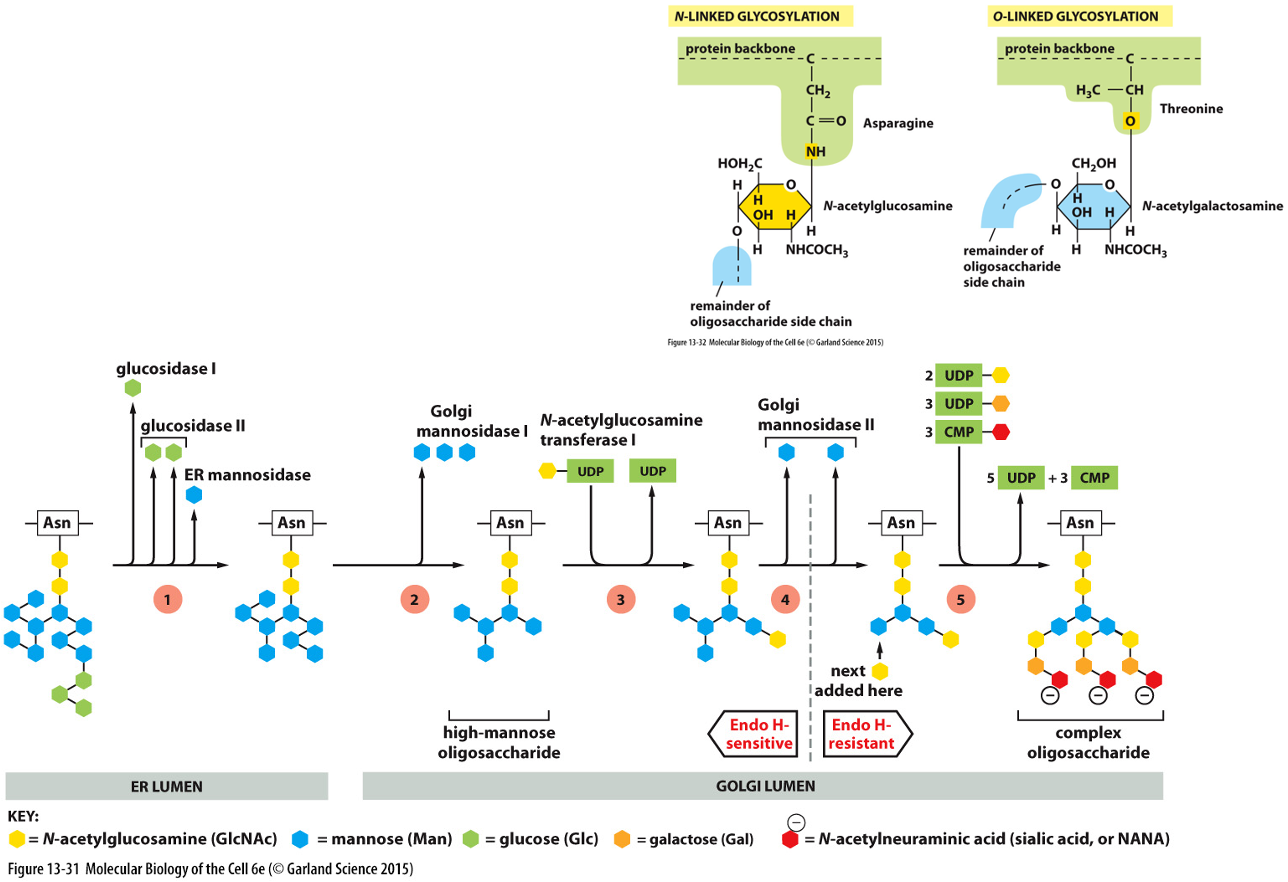
Oligosaccharide Processing in Golgi Apparatus:
Resident proteins are all membrane bound
Finishes processing of oligosaccharides that started in ER
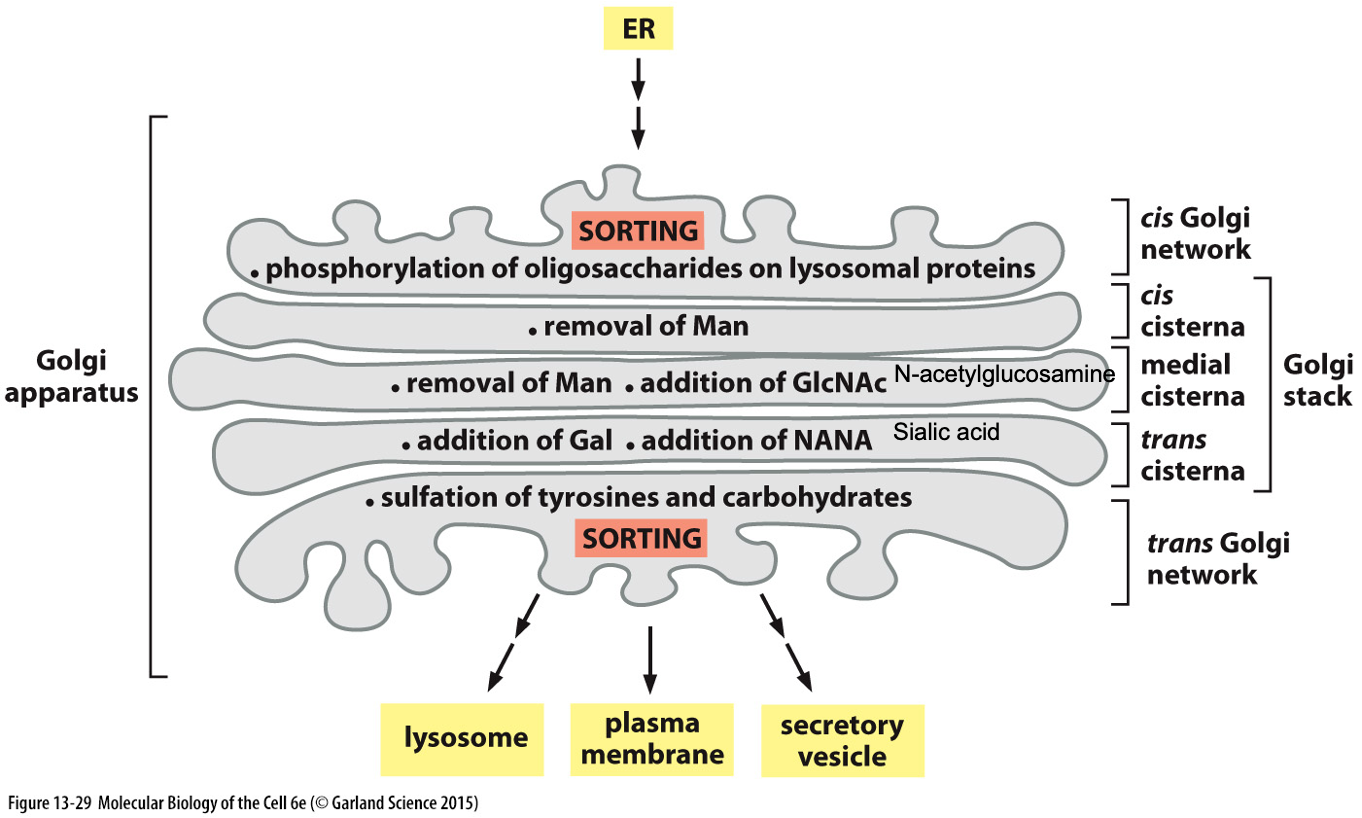
Models of Golgi Transport
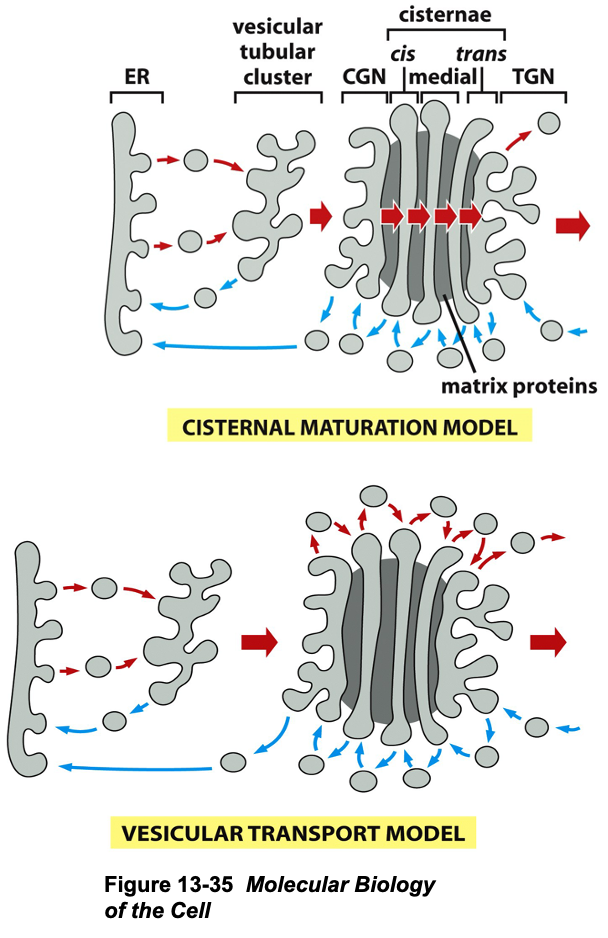
Transport Models:
Vesicular Transport Model:
Static structure where proteins move through cisternae carried by transport vesicle (cis » trans)
Cisternal Maturation Model:
Dynamic model describing how cisternae move,
Vesicular tubular cluster fuse to form cis (Golgi network) » medial (cisterna) » trans (cisterna)
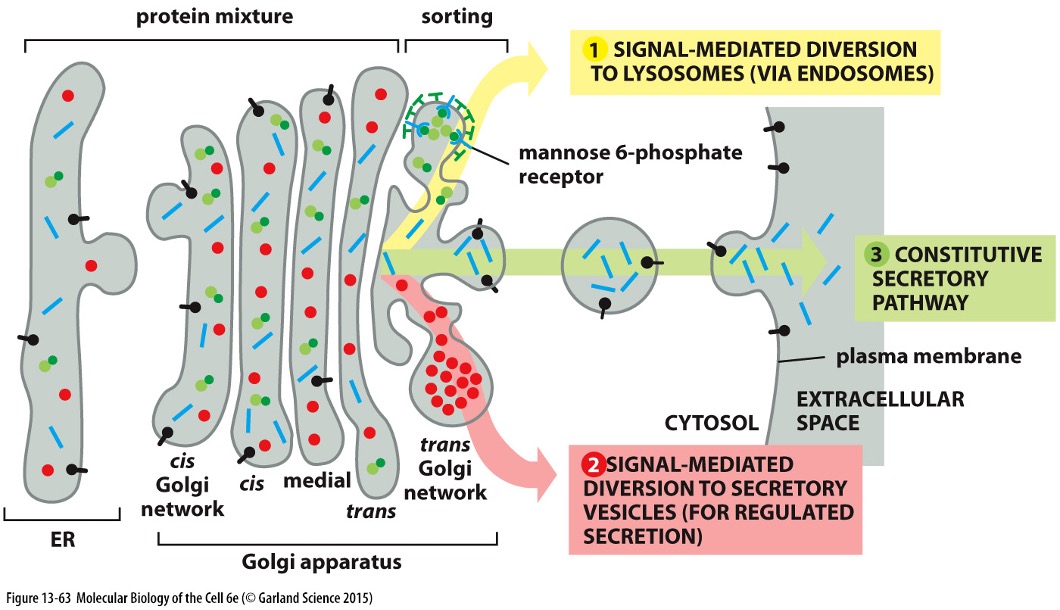
Protein Sorting and Secretory Pathway
Final Sorting in Trans-Golgi:
Sorting signals determine the protein's final destination (e.g., lysosome or secretion).
Enzymes that function in Golgi (e.g. glycosyltransferases) are retained by inserting in membrane (specific membrane spanning domain)
Secretory proteins are then packaged into transport vesicles and taken to the cell surface
Secretion Proteins » To Surface:
Proteins can be continuously secreted or regulated in response to stimuli (e.g., insulin and blood glucose).
Some proteins can undergo cleavage in the vesicles
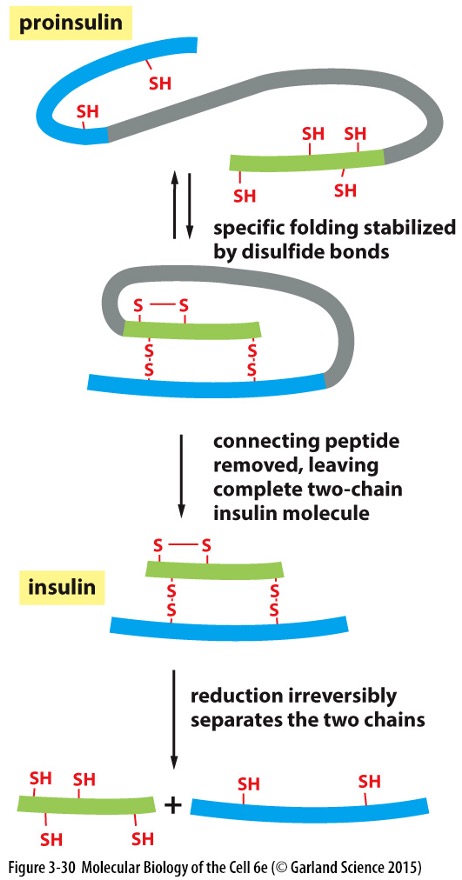
Specific Proteolytic Cleavages
Proproteins require additional processing to become active (‘mature’) which occurs in secretory vesicles
Secreted proteins are cleaved by proteases prohormone convertase 2 (PC2) and 3 (PC3) and furin
Example: Proinsulin » folds » cleaved » insulin
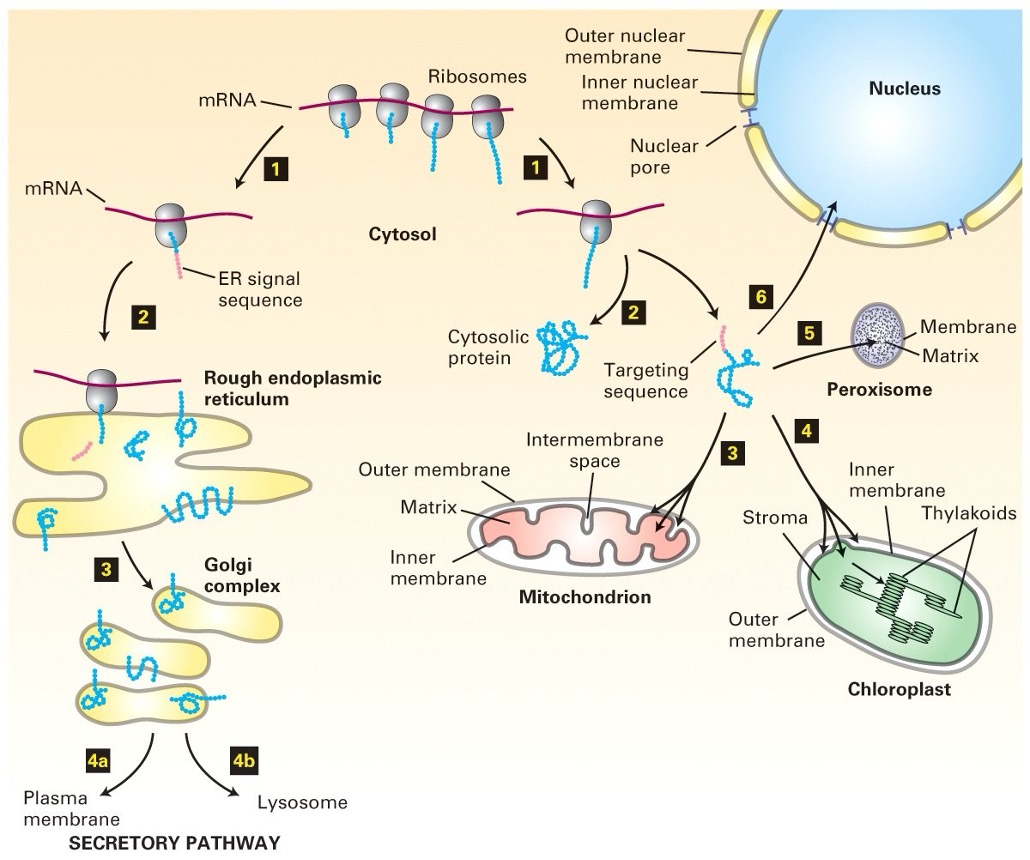
Summary of Protein Transport
Key Steps:
Protein synthesis occurs on ribosomes (both free and bound).
Proteins translocated across the ER and undergo modifications (disulphide bonds, glycosylation).
Move to Golgi for further processing.
Proteins reach the trans Golgi and are secreted or sent to specific organelles.
Further Reading
Recommended texts:
Molecular Biology of the Cell, 6th Edition, by Alberts et al.
Essential Cell Biology, 3rd Edition, by Alberts et al.