Translation and Proteins
Vocabulary:
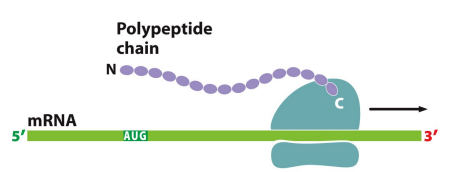
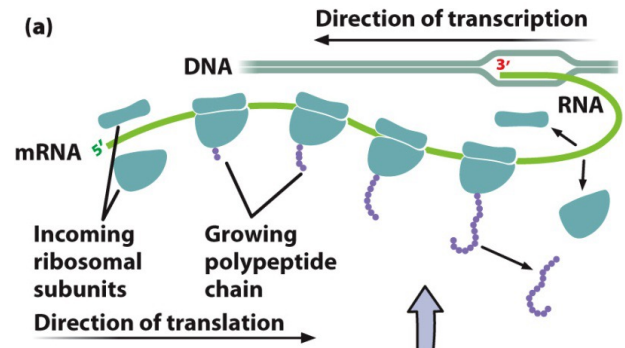
Translation: The biological polymerization of amino acids into peptide chains, requires amino acids, mRNA, ribosomes, and tRNA
Ribosome: An organelle used in translation, plays an essential role in expression of genetic information, made from rRNA
rRNA: A form of RNA that builds ribosomes, performs the central catalytic functions associated with translation, has large and small subunits
Svedberg Unit (S): A subunit of the ribosome
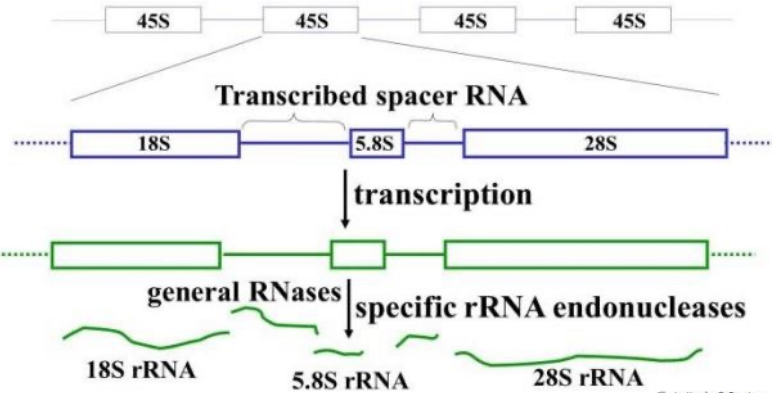
Ribosomal DNA (rDNA): The genes that build rRNA, moderately repetitive DNA fractions present in clusters (45S) at various chromosomal sites, each cluster contains tandem repeats separated by noncoding spacer DNA. Clusters repeated 200-400 times
Spacer DNA: A genetic material that separates different clusters of rDNA
45S transcription units: Transcribed as a precursor to rRNA, can be processed into 3 mature rRNAs, 18S, 5.8S, and 28S
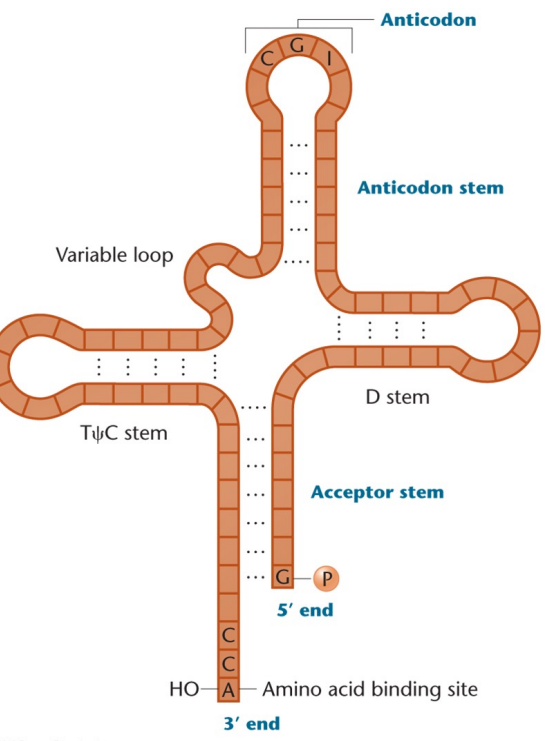
tRNA: A kind of RNA that adapts genetic information presented as specific triplet codons in mRNA to corresponding amino acids, contain anticodons and carry amino acids. Only 75-90 nucleotides long
Posttranscriptionally modified bases: Something contained in tRNAs structure, present after transcription of it and occurs as a result of cleaving a larger precursor
Cloverleaf structure: What tRNA looks like
Anticodon: Part of tRNA, complementarily base-pairs with the codon in mRNA
CCA sequence: Where the amino acid is covalently linked to in tRNA, at the 3’ end
Charging: The process of linking the appropriate amino acid to the tRNA
Aminoacylation: tRNA charging, before translation this must occur. Highly specific, the amino acid is converted into an active form through the use of ATP, resulting in the AMP and amino acid being covalently linked
Aminoacyl tRNA synthase: An enzyme that catalyzes aminoacylation, 20 classes, one for each amino acid. Once bound to an amino acid will find the correct tRNA and charge it
-Isoaccepting tRNA: The tRNA that will correctly match to the amino acid and aminoacyl tRNA synthase
-Aminoacyl tRNA: A charged tRNA, which may be used directly in protein synthesis
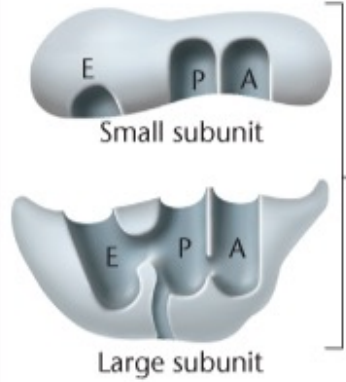
-Aminoacyl site (A): Where different charged tRNAs enter after the P-site recognizes AUG or after a new peptide bond is formed, moves charged tRNA into the P-site as the polypeptide forms
-Peptidyl site (P): Part of 70S, once the start codon is here the complex is poised for the insertion into the A-site of the second aminoacyl tRNA
-Exit site (E): Where tRNA enters after it loses the amino acid it was carrying and becomes uncharged, from the P-site to here. Releases empty tRNA
Initiation Factor 3 (IF3): Part of initiation of translation in prokaryotes, binds to 30S to prevent premature binding of 50S
-Initiation Factor 1 (IF1): Part of initiation of translation in prokaryotes, binds to 30S and prevents aminoacyl tRNA from binding to the A-site prematurely. Ensures fMet occupies the P-site first and that translation starts precisely at AUG
-Initiation Factor 2 (IF2): Along with GTP, facilitates the binding of the initiator tRNA (fMet) with mRNA, stabilizing it to the P-site
-GTP: When hydrolyzed causes the dissociation of all the initiation factors and allows the 30S to bind to 50S
70S: The structure formed when 30S binds to 50S, complete initiation complex
Shine-Dalgarno sequence (AGGAGG): In prokaryotes, what 30S binds mRNA to, precedes AUG start codon in bacteria
-16S: A part of 30S, base pairs with the Shine-Dalgarno sequence
-30S: The small ribosomal subunit (cap)
-50S: The large ribosomal subunit
-Elongation factors: Tu, Ts, and G. These transport charged tRNAs to the complex
-Peptidyl transfer: the process of moving the newly formed polypeptide from the P-site to the A-site for further elongation
Peptidyl transferase: A function of 23S rRNA, previously thought to have been a catalytic enzyme for reactions of moving sites, though it is actually just 23S rRNA
-Translocation: The process of the ribosome moving along the mRNA by one codon as it forms a polypeptide
-Tu (EF-Tu/EF-I): Elongation factor, bound with GTP and facilitates binding of the proper charged tRNA into the A-site. Becomes recharged with GTP hydrolysis
-EF-G/EF-II: Aids in translocation in the ribosomal subunits
-23S rRNA: Part of 50S, a ribozyme that catalyzes peptide bond formation between amino acids on tRNA at the A-site and the growing peptide chain bound to tRNA at the P-site
Termination: The process that ends translation, signaled by the codons UAG, UAA, and UGA in the A-site, which don’t signal any amino acids
GTP-dependent release factors: Stimulate hydrolysis of polypeptide from peptidyl tRNA, released from translation complex
-Release factors: During prokaryotic termination, bind to the stop codon in the A-site
-GDP: Formed via hydrolysis of GTP, causes a conformational change in the ribosome during termination
-Stop codons (UAG, UAA, UGA): A codon that doesn’t have a corresponding tRNA or associated amino acid, instead causing termination
Polysome: mRNA with several ribosomes translating it at once, as the mRNA passes through one ribosome it associates with another small subunit
-Cap-dependent translation: Eukaryotic translation, involves the m7G cap on the 7’ end of mRNA associating with a small ribosomal subunit
-Eukaryotic initiation factors (EIFs): Initiation factors for translation in eukaryotes
-Kozak sequence: Like the Shine-Dalgarno sequence but for eukaryotes, increases the efficiency of translation by interacting with initiator tRNA, physically is purines (A or G) three bases upstream from the AUG initiator codon, and a G following the AUG
-Rough endoplasmic reticulum: An organelle that contains some ribosomes
-Closed loop translation: The process that free-floating ribosomes undergo to be translated as polyribosomes, mRNA forms a closed loop with cap and tail together
-eIF4G: A cap binding protein that serves as a scaffolding protein, helps bind the cap and tail together in closed loop translation
-Alkaptonuria: Results from a mutation, leads to metabolic blocks. Stops individuals from metabolizing alkapton, which leads to an accumulation of and intermediate substance called homogentisic acid
-Phenylketonuria (PKU): Results from a mutation and leads to metabolic blocks, cause an individual to be unable to convert phenylalanine to tyrosine, as phenylalanine hydroxylase is inactive. Toxic levels of phenylalanine and low levels of tyrosine. Phenylalanine enters the cerebrospinal fluid and results in seizures and cognitive disabilities
-George Beadle: Showed the first evidence that genes are directly responsible for the synthesis of enzymes, showed nutritional mutations in bread mold caused a loss of enzymatic activity
-Auxotrophy: The inability of an organism to synthesize a particular organic compound required for its growth
-One-gene One-enzyme Hypothesis: Thought of by Beadle and Tatum though ultimately wrong, describes how each trait or gene is linked to one specific enzyme
-One Gene One Polypeptide Chair Hypothesis: Revision of one-gene one-enzyme hypothesis, due to not all proteins being enzymes, and some proteins having subunit structures with two or more polypeptide chains
-Sickle Cell Anemia: Caused by a point mutation in hemoglobin, HbA and HbS have one different amino acid resulting in different folding leading to a sickle shape of red blood cells
-Fingerprinting: A technique that uses the 2D separation of peptide fragments into distinctive patterns of spots
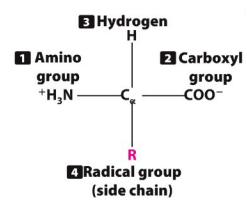
-Peptide: Precursor to proteins, assembled amino acid chains from N-terminus to C-terminus
-Proteins: Peptides that are folded up into functional 3D structures, have many diverse functions
-Peptide bond: Links amino acids together, formed between a carbon and a nitrogen through condensation and broken via hydrolysis
-Cotranslational folding: When a protein begins to fold into their 3D conformation based on chemical properties of the amino acid sequence as soon as it is translated
Chaperones: Proteins that mediate the folding of other non-spontaneously folding proteins
Ubiquitin ligase: Attaches ubiquitin to misfolded proteins in eukaryotes
-Proteosome: A large protease complex that recognizes proteins flagged by ubiquitin and degrades them
Denaturation: Happens when a protein loses its 3D structure, and can lead to loss of biogeological activity
-Prion: A misfolded protein in the brain that can cause aggregates that may be infectious. When they come into contact with normal proteins they refold the normal proteins and create clusters that eventually destroy the disease
-Immunoglobulins: Proteins that function in immune systems in vertebrates
-Transport proteins: Proteins that aid in the movement of molecules across membranes
-Hormones/Receptors: Proteins that regulate various types of chemical activity
-Histone: A kind of protein that binds to DNA in eukaryotes for tight packing
-Transcription factors: A kind of protein that regulates gene expression
-Hemoglobin/Myoglobin: Proteins that transport oxygen which is needed for cellular metabolism
-Collagen/Keratin: Proteins that are important for structural integrity in skin, connective tissue, and hair
-Actin/myosin: Contractile proteins found in muscle tissue
-Tubulin: The protein that is the basis of microtubule function in mitotic and meiotic spindle fibers
Enzymes: Proteins that will increase the reaction rate of a spontaneous reaction and lower the activation energy, most diverse group
Catabolism: Degrades larger molecules into simpler ones and releases chemical energy
Anabolism: The synthetic phase of metabolism, building molecules from smaller ones and using chemical energy
Requirements of a prokaryote for translation initiation are the small and large ribosomal subunits, an mRNA molecule, GTP, a charged initiator tRNA, Mg2+, and initiation factors
Elongation in prokaryotes requires 30S, 50S, mRNA, GTP, a charged initiator tRNA, Mg2+, Tu, Ts, and G
Generally elongation in prokaryotes involves fMet in the P-site bound to AUG, followed by the next charged tRNA entering the complex at the A-site. A peptide bond is formed between the amino acids in the two tRNAs. Now, the polypeptide is transferred to the A-site. The ribosome translocates, and shifts the empty tRNA from the p-site to the e-site and it is released
During termination in prokaryotes, release factors RF1 or RF2 bind to the stop codon in the A-site of the ribosome, causing the release of the polypeptide from the P-site with the use of hydrolysis. RF3 with GTP binds the ribosome, hydrolysis of this into GDP causes a conformation change in the ribosome, and the ribosome separates releasing mRNA, tRNA, and release factors
Posttranscriptional modifications of peptides include removal or modification of the N-terminus enzymatically, some radical groups undergoing conformational changes or phosphorylation (for tyrosine), the occassional attachment of carbohydrate side chains (for glycoprotein synthesis), chain trimming of mostly signaling sequences, and the addition of metals