Transcription: Prokaryotic vs. Eukaryotic Mechanisms and Regulation
Prequel: Transcription, mRNA processing, and translation
DNA serves as the instruction manual for making proteins
Going from instructions to product occurs in 3 main steps:
Transcription
The information contained in the DNA is copied into a complementary strand of RNA (ribonucleic acid)
This RNA copy is called MRNA
Why is transcription necessary? (why copy the info)
Location issues
DNA is too valuable
MRNA Editing
The mRNA copy needs to be modified (cleaned-up)
before it leaves the nucleus
- Does not occur in prokaryotic cells
Translation
Ribosomes read the mRNA and make a protein
Activation of transcription
Transcription is gene specific
Each gene provides instructions for a single protein (usually
When turned on, only that one gene is transcribed (unlike DNA rep)
What tells a cell to start transcribing a specific gene?
Different signals for different genes in different cell types
Constitutive Expression: Some genes are transcribed continuously
independent of cell health or environmental conditionsUsually seen for those proteins that are needed 100% of the time
Example: ATP synthase, actin, tubulin
Regulated expression
Allows for much tighter control of gene
expressionSignal from outside or inside the cell
starts the whole processOften involves one or more signal transduction pathway ____________
Prokaryotic transcription
All genes (eukaryotic and prokaryotic) have a unique region of DNA sequence
upstream of (before) the transcriptional start site that serves as a binding site for
the RNA polymerase enzyme (makes RNA)
These regions are called promoters
General functions of promoters:
Provide specificity: Tells the cell where the gene of interest is
This is critical – transcribing the wrong gene could be deadly!!
Each gene has a unique promoter sequence
Tell RNA polymerase WHERE TO START (they are usually at the beginning)
Analogy: Front cover of a book
No promoter (or mutated promoter), no transcription
Upstream_____ pathways
Prokaryotic transcription: Initiation – Finding the gene
Although each promoter is unique, most share a few common sequence
elements called Consensus Sequences
These common elements are evolutionarily conserved
These common elements are extremely important for transcription
If mutated, transcription does not take place
Prokaryotic promoters share 2 main types of consensus sequences
TATA BOX (aka Pribnow box) – found 10 nucleotides upstream from the start site
(-10-15), sequence is usually TATAAT
TTGACA – found 35 nucleotides upstream from start (-35).
Exact sequence and spacing can vary!
RNA polymerase enzyme makes direct contact with the promoter at these
consensus sequences
• Unique seq. within and around promoters help determine rates of transcription
Prokaryotic transcription: Initiation – Binding of RNA polymerase to the promoter
Prokaryotic cells only produce 1 type of RNA polymerase
- Produces all of the mRNA, tRNA, and rRNA in the cellProkaryotic RNA polymerase structure:
- 2 alpha (α), 1 beta (β), 1 beta prime (β'), and 1 sigma (σ) subunitβ, β’, α subunits interact and form the core enzyme
- Together they have the basic catalytic function of producing new RNAσ factor controls binding to the promoter Promoter Recognition
After initial binding has been successful, the σ factor comes off the core
ANALOGY: σ factor is the general, the core enzyme is the troops
Bacteria make many Different Type of sigma factors
Different sigma factors recognize different promoter sequences
Examples: σ70 – controls transcription of general purpose genes
σ54 – controls transcription of genes only during nitrogen
starvation
Prokaryotic Transcription: Initiation and Elongation
Initiation and Elongation
Initiation Steps:
RNA polymerase (RNA pol) binds to a specific promoter sequence.
It positions itself over the transcriptional start site, chosen based on its distance from the consensus sequence
No specific "start signal" exists.
RNA pol unwinds the DNA near the start site, creating a transcription bubble.
It reads the first ~- DNA nucleotides and brings in complementary RNA nucleotides (inserting U across from A).
These nucleotides are connected via phosphodiester bonds.
Only one DNA strand is transcribed.
RNA pol moves 5’→3’ in the direction, initially slowly.
Change in RNA Polymerase Shape:
After transcribing ~ nucleotides, RNA polymerase changes shape.
This change causes it to lose its attraction for the consensus sequences, allowing it to move quickly along the gene.
It also causes the sigma factor to fall off.
Once the factor is free, RNA pol moves quickly along the rest of the gene (in the direction).
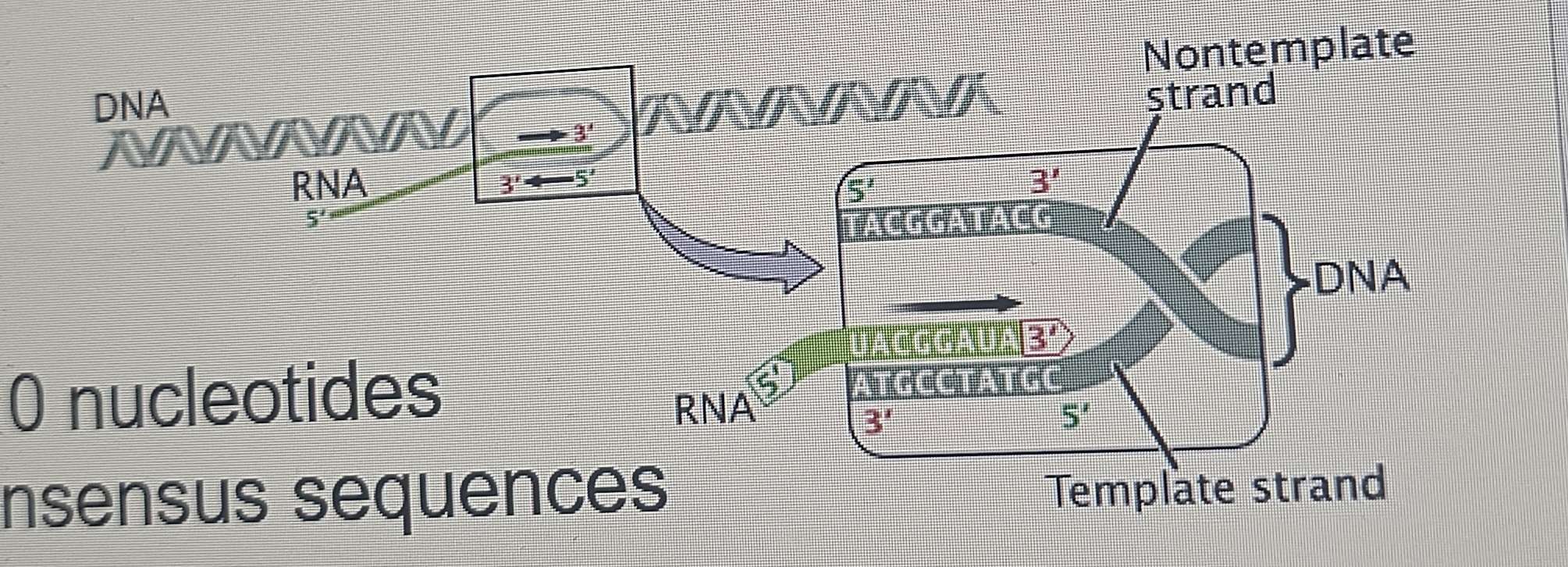
Prokaryotic Transcription: Termination
RNA pol continues producing complementary RNA until it transcribes a terminator signal within the sequence.
Analogy: This is not like a stop light; it's more like spikes on the road, forcing the vehicle to stop.
Two major prokaryotic transcriptional terminator signals exist.
Both mechanisms must:
Slow down the RNA polymerase.
Weaken the interaction between the DNA and RNA within the transcription bubble.
Properties of the two major prokaryotic terminator signals:
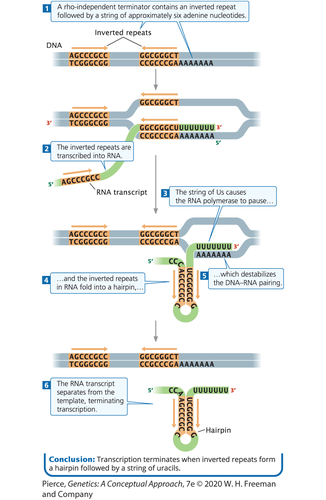
Rho-independent (Intrinsic Termination):
The DNA sequence contains an inverted repeat followed by a string of 6 adenines (poly-adenine).
Following their transcription, the inverted repeats within the newly synthesized RNA hydrogen bond with each other, forming a stable hairpin loop structure.
The formation of this loop causes the RNA pol to pause.
This pause, combined with the weak hydrogen bonding between the ~ straight A-U pairs (transcribed from the DNA's poly-A stretch) in the DNA-RNA hybrid, causes the RNA to totally fall off the DNA template.
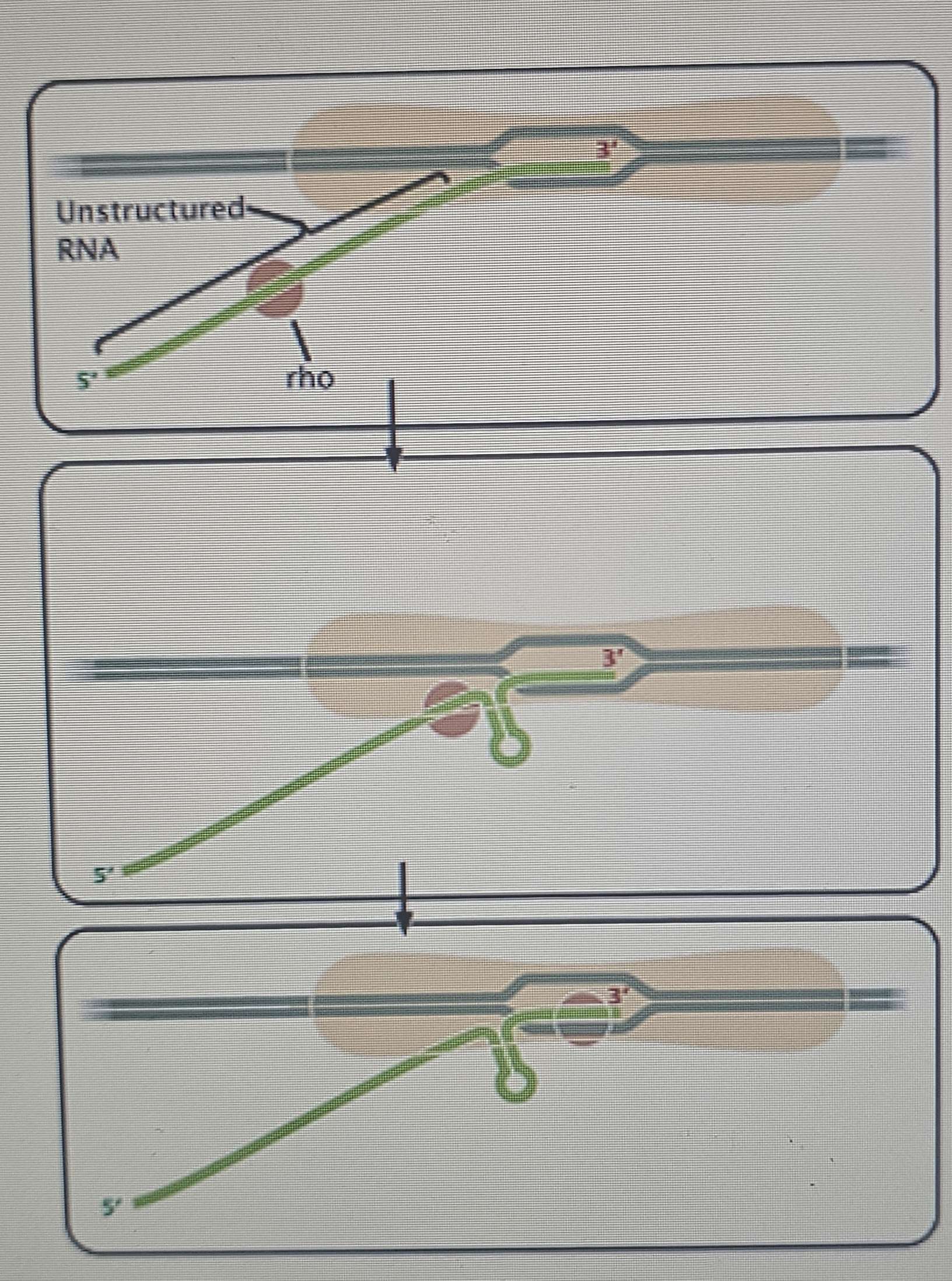
Rho-dependent Termination:
Also contains inverted repeats that cause hairpin loop formation in the new RNA, which slows down the polymerase.
The RNA contains a binding site for the Rho protein.
Rho moves towards the end of the nascent RNA.
It catches up to the stalled RNA polymerase and, acting as a helicase, unzips the RNA from the DNA template.
These terminators do not contain an adenine-rich region after the inverted repeat.
Nearly all bacterial genes have one of these two types of terminators.
Eukaryotic Transcription: Activation and Initiation
Eukaryotic transcription follows the same basic steps as prokaryotic transcription: Activation, Initiation, Elongation, Termination. However, major differences exist in how these steps are accomplished.
General features of transcriptional Activation are similar, with some genes constitutively expressed and others regulated.
The exact nature of activation signals can be very different (e.g., testosterone's effect in eukaryotes vs. bacteria).
Initiation – Making the DNA Accessible:
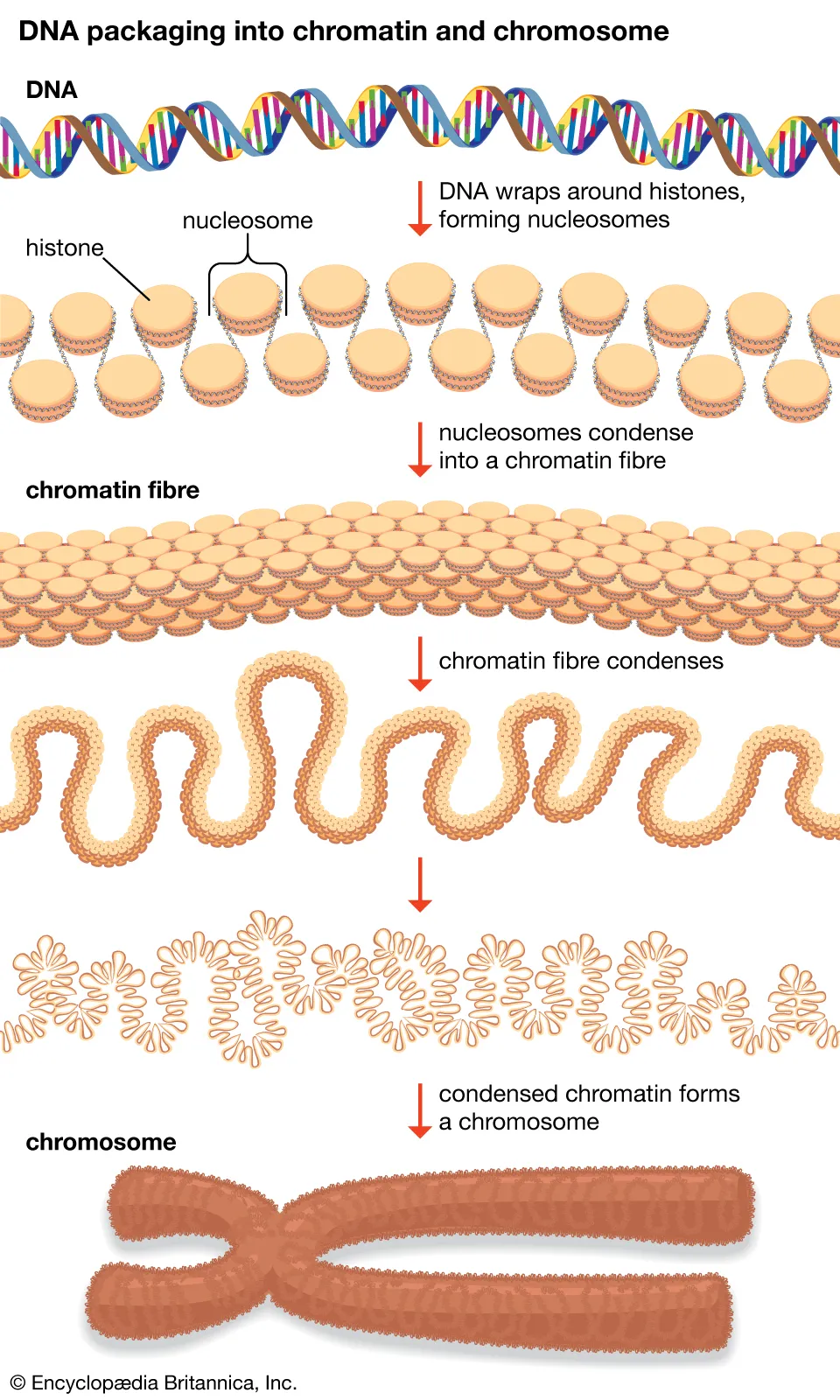
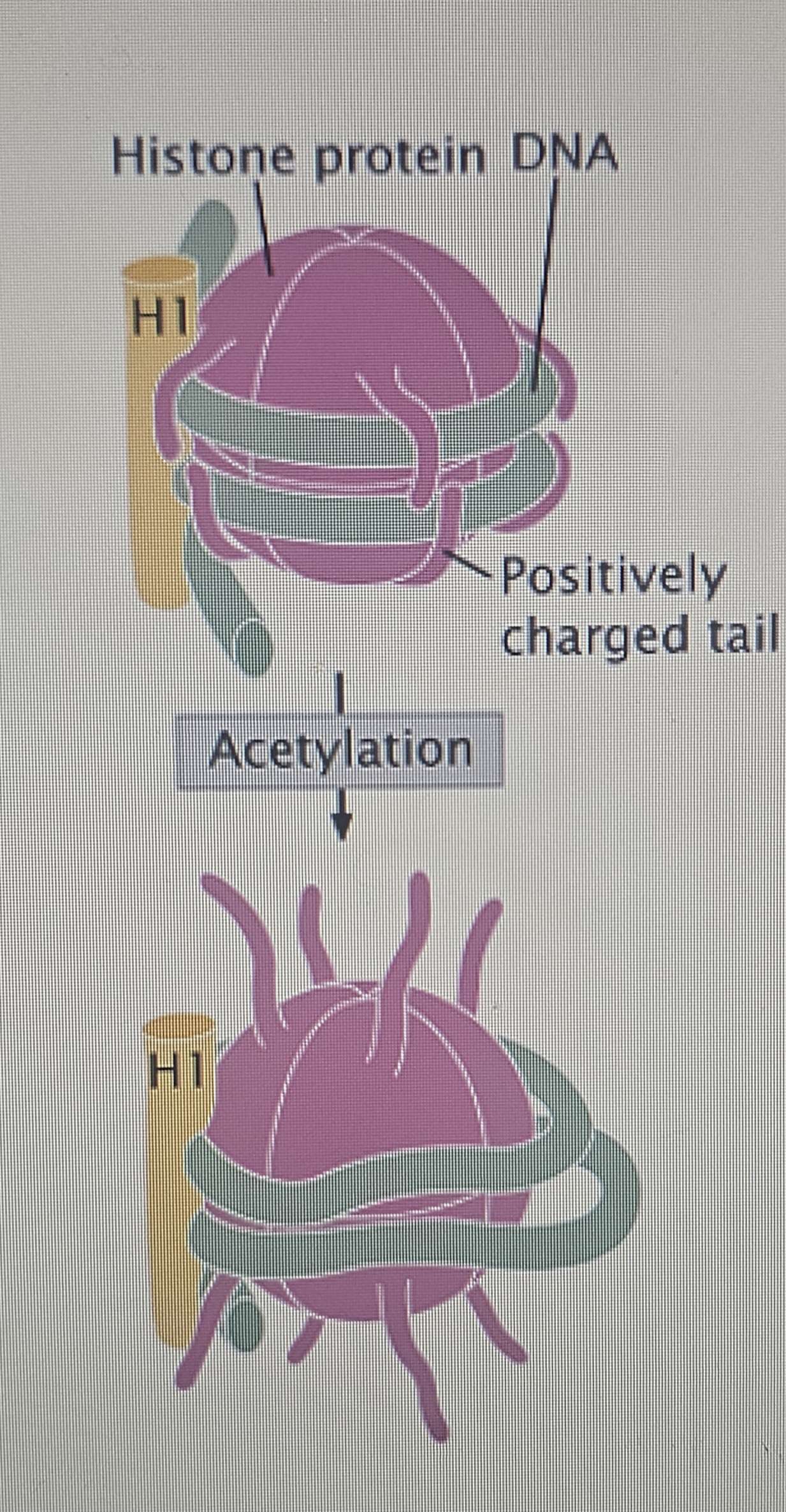
Eukaryotic DNA is tightly coiled around proteins called histones (and some non-histone proteins).
DNA is negatively charged (due to phosphate groups), while histones are very positively charged.
Every ~ base pairs of DNA are wrapped around a histone octamer (a complex of histone proteins), forming a nucleosome.
To initiate transcription, DNA in the gene area must be loosened/unwound from histones and other proteins.
DNA is freed from histones in two major ways:
Histone Acetylation:
Enzymes called histone acetyltransferases (HATs) add an acetyl group () to histones.
This neutralizes their positive charge, causing them to lose attraction for the negatively charged DNA.
This process is specific and occurs only in the area to be transcribed.
Deacetylases remove acetyl groups after transcription, allowing DNA to re-condense.
Chromatin Remodeling:
Some proteins actively move nucleosomes around, freeing up DNA without directly modifying histones.
Eukaryotic Transcription: Initiation – Finding the Gene (Promoters and Enhancers)
Eukaryotic Promoters:
Much more complex than bacterial promoters.
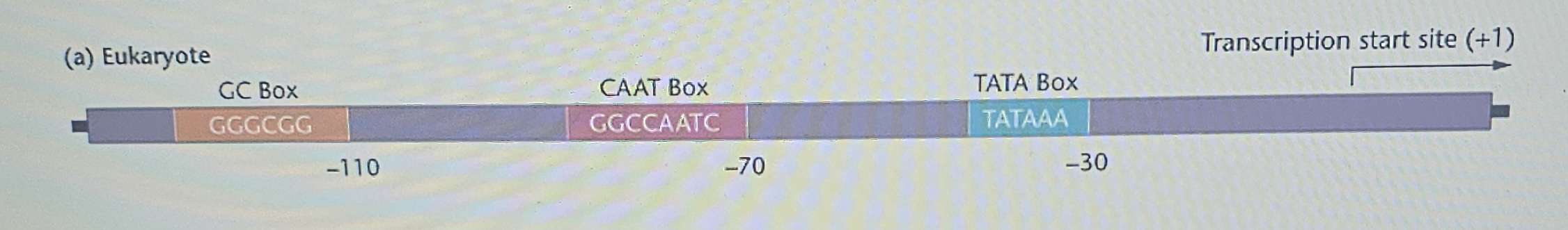
Major consensus sequences include:
TATA box: Similar to the prokaryotic TATA box, but with the sequence TATAAA and located at positions from relative to the transcriptional start site.
CAAT box: Located at , always containing either CAAT or CCAAT.
Mutations in this region usually significantly lower the rate of transcription.
GC box: Usually has the sequence GGGCGG.
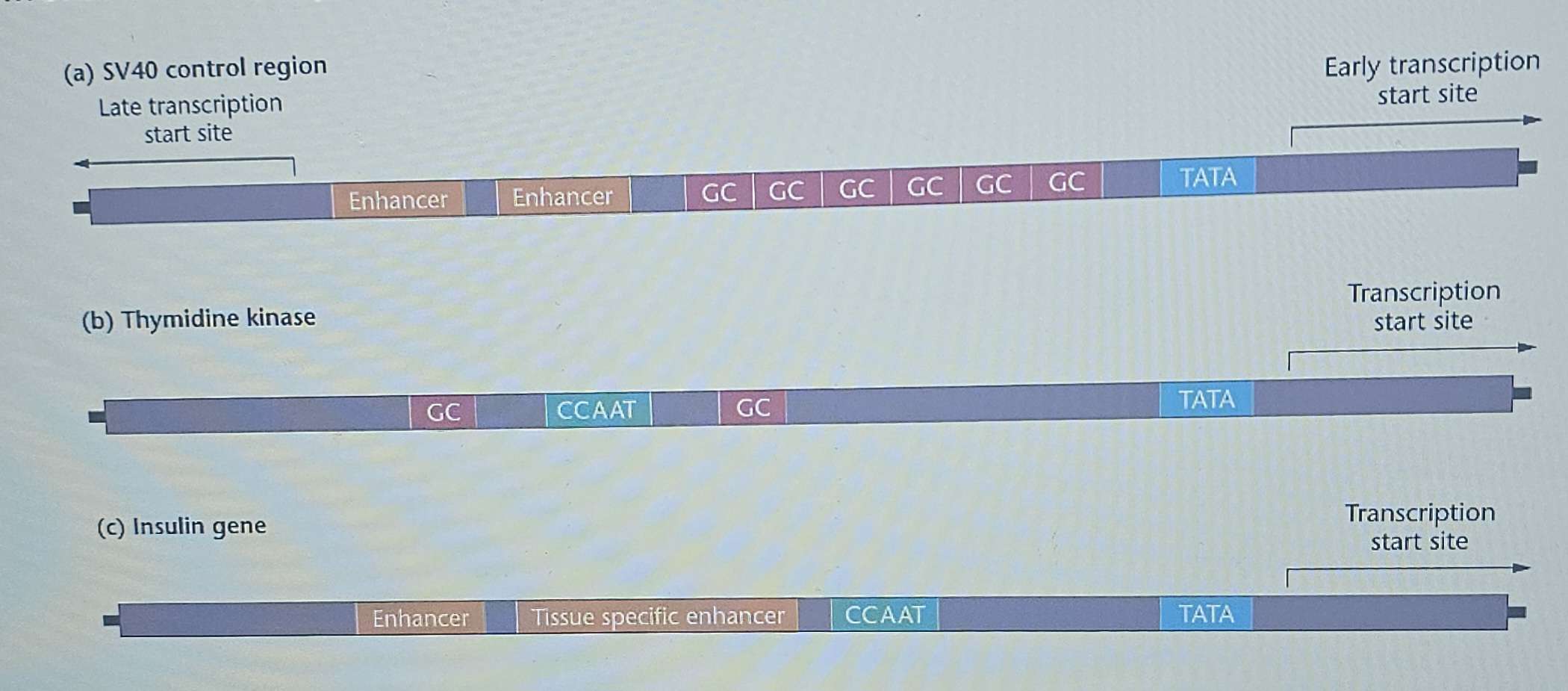
Promoters can contain multiple copies of each consensus sequence.
The regions found between consensus sequences are unique in each promoter, allowing the cell to recognize and distinguish different genes.
Eukaryotic Transcription: Initiation – Binding of RNA Polymerases to the Promoter
Prokaryotic Review: A single type of RNA polymerase recognizes and binds directly to consensus sequences.
Eukaryotic Complexity: Eukaryotic cells are more complicated, having multiple RNA polymerases, and none of them recognize, bind directly to promoter sequences.
Three Eukaryotic RNA Polymerases:
RNA polymerase I: Produces ribosomal RNA (rRNA).
RNA polymerase II: Produces messenger RNA (mRNA).
All mRNA is made by RNA pol II (the focus of discussion).
RNA polymerase III: Produces some types of transfer RNA (tRNA).
All three transcribe DNA into RNA (not just mRNA).
They are much larger and more complex than bacterial RNA polymerase, reflecting the diversity of eukaryotic genes (e.g., different genes).
Role of Transcription Factors:
Since eukaryotic RNA polymerases don't bind directly, they rely on transcription factors (TFs).
TFs are a class of proteins that bind to DNA and help recruit RNA polymerase enzymes to promoters, similar to the sigma factor in bacteria.
They regulate eukaryotic transcription.
Two main classes of TFs:
Basal transcription factors (General TFs): Common proteins needed to initiate transcription by all promoters, yielding low levels of transcription.
Regulatory transcription factors: Provide more specific transcriptional control, either activating or repressing transcription above or below basal levels.
Either activate or repress transcription
above/below basal levels
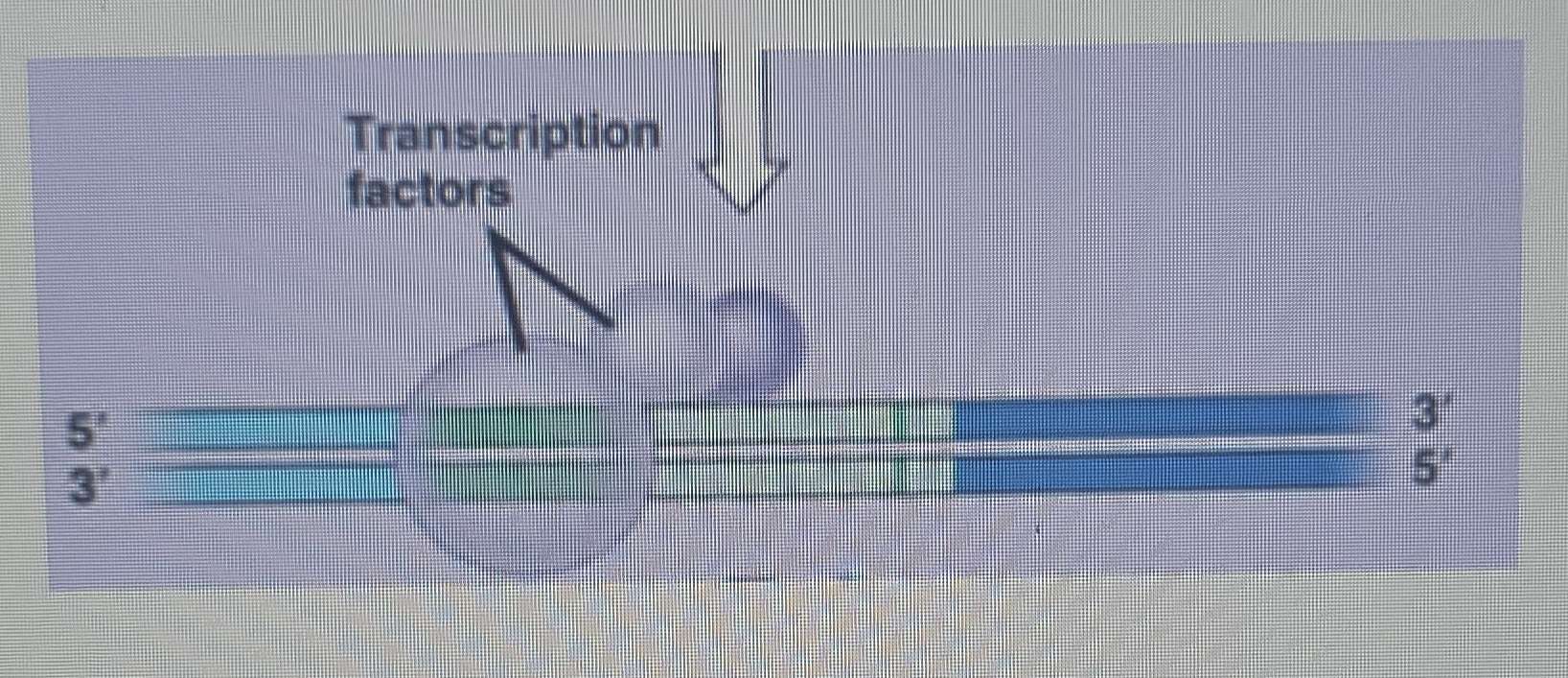
Binding of Basal Transcription Factors and RNA Pol II Recruitment:
The TFIID complex (specifically its TBP subunit, TATA-binding protein) binds to the TATA box.
This binding alters DNA shape, allowing TFIIA and TFIIB to bind.
TFIIA acts as a stabilizer interaction between TBP and the DNA.
TFIIB helps find the transcriptional start site.
RNA polymerase II, escorted by TFIIF, then interacts with the preassembled complex.
Other factors assist RNA Pol II in gaining direct access to the promoter.
TFIIH serves as a helicase to separate DNA strands during transcription.
Once this giant protein complex is assembled on the TATA box, RNA pol II will leave most other proteins behind and begin making mRNA.
These events occur during all eukaryotic gene transcription and provide just basal transcription.
To achieve higher or lower levels of transcription, cells utilize regulatory transcription factors.
Eukaryotic Transcription: Elongation and Termination
Elongation:
The basic mechanism is essentially the same as in prokaryotic cells.
Involves a transcription bubble and the addition of ribonucleotides to the end of the growing RNA strand.
Termination:
Each type of RNA polymerase uses a different termination mechanism, none as well characterized as in bacteria.
For RNA polymerase II, no clear termination signal typically follows the allosteric/torpedo model.
Transcription continues well beyond the actual end of the gene.
Termination is coupled to mRNA processing (discussed in later contexts).
Regulation of Transcription: Prokaryotic Cells
Prokaryotic and eukaryotic cells control how often gene transcription rates to regulate protein levels (e.g., liver-specific genes not transcribed in brain cells).
Mechanisms of Prokaryotic Transcriptional Regulation:
Promoters and Sigma Factors:
Bacteria contain many different types of sigma factors.
Different sigma factors recognize different promoters.
Some sigma factors promote high transcription levels, others low levels.
When a cell needs to transcribe specific genes, it swaps out sigma factor components to recognize the appropriate promoters.
add the appropriate sigma factor to the core components sends it off.
Gene Grouping (Operons):
Bacteria efficiently organize genes with related functions together into operons.
Example: Genes for eye color in humans may be on different chromosomes, but all genes for tryptophan (Trp) production in bacteria are found sequentially.
An operon contains:
A promoter region, which includes the operator region.
The operator is a binding site for a repressor protein, acting as an ON/OFF switch.
A set of related genes found in tandem (back-to-back).
Because they share a single promoter, if you turn one gene on, you turn all of them on.
Types of Operon Regulation:
a) Positive Regulation (Inducible Operons):
Transcription is normally turned OFF.
This is because a repressor protein is active when unbound and blocks transcription.
An inducer molecule binds to the repressor, inactivating it.
* With the repressor inactivated, transcription of all genes in the operon is allowed to occur.
* Example: The Lac operon (induced by lactose).
b) Negative Regulation (Repressible Operons):
Transcription is normally ON.
A co-repressor molecule binds to and activates the repressor protein.
The activated repressor then binds to the operator, shutting off transcription of all genes in the operon.
* Example: The Trp operon (repressed by tryptophan).
Regulation of Transcription: Eukaryotic Cells
Eukaryotic cells use much more complicated mechanisms for transcriptional control.
Mechanisms of Eukaryotic Transcriptional Control:
Regulatory Transcription Factors:
Basal TFs provide only low levels of transcription; their sole function is to recruit RNA polymerase to DNA.
Regulatory TFs alter transcription above or below basal levels, providing fine control.
Cells have hundreds of these.
Regulatory TFs bind to two types of DNA sequences:
Promoters (beyond just consensus sequences).
Enhancers: Function to increase transcription rates. They can be located far from the promoter (upstream or downstream).
TF binding causes DNA bending, bringing the enhancer and promoter into close proximity in 3D space.
How Regulatory TFs Work:
a) Directly altering RNA polymerase activity: Some regulatory basal TFs bind to DNA and directly alter the recruitment, binding, or movement of RNA polymerase. They may sterically block binding or stabilize it.
b) Altering chromatin structure: Others bind to DNA and recruit or block HATs thus alters chromatin structure, for example, by increasing or decreasing histone acetylation, which affects how well basal TFs can access the DNA.
End Result of Both: Either clear the way for basal TFs (enhancing transcription) or block them (reducing transcription).
DNA Methylation:
A methyl group is enzymatically added to cytosines (specifically, in CG dinucleotides) via specific methyltransferases.