Class 1: Chlamydomonas - a Model System
Chlamydomonas Reinhardtii (Clammy)
Unicellular, eukaryotic (green algae with a flagella, a giant chloroplast (chlamy is photosynthetic), and an eyespot (a primitive ey e)
It has a simple structure: plasma membrane, cell wall, and an eyespot which controls phototaxis (eyespot detects light, directs flagella to move towards or away from it)
Prokaryotic vs. Eukaryotic Cells
Prokaryotic cells are approximately 1 micron in size.
Eukaryotic cells like Chlamy are about 10 microns.
Eukaryotic DNA is contained in the nucleus while in prokaryotes it is found in the cytoplasm.
Growth and Division of Clammy
Chlamy grows by binary fission (so binary fission is not limited to prokaryotes)
Chlamy is usually haploid which is advantageous for genetic studies; expresses mutations directly (either mutated or not mutated, no masking possible with a WT).
Divides approx every 10 hours, slower than bacteria (20 minutes).
Nutritional Requirements
Grown in a specific medium called TAP, which contains essential nutrients:
Macronutrients: Sulfur, Nitrogen and Phosphorus in higher amounts
Micronutrients: Molybdenum and Copper in lower amounts (usually 1000x less)
Sulfur:
In humans (and algae like Chlamy): Needed to make amino acids cysteine and methionine, needed for protein synthesis and function. Cysteiene residues also form disulfide bonds that stabilize protein structure.
In Chlamy: Sulfur is part of many coenzymes and is involved in synthesizing sulfolipids, needed for the membranes of chloroplasts
Copper:
In humans (and Chlamy): Copper is in enzymes like cytochrome c oxidase (COX) for mitochondrial energy production and superoxide dismutase (SOD), an antioxidant.
In Chlamy: Copper is needed as a cofactor for electron shuttle enzymes due to its redox properties. Plastocyanin is a protein made from copper that transfers electrons during light reactions of photosynthesis.
Molybdenum:
In humans: Needed for detoxing enzymes like xanthine oxidase (purine metabolism), aldehyde oxidase and sulfite oxidase.
In Chlamy: Needed as a cofactor in molybdoenzymes (e.g. nitrate reductase).
Growth Curves
Chlamy doesn’t always divide at a growth rate of 10 hours - that is the growth rate during the exponential phase
Chlamy stops growing when all nutrients in TAP media are exhausted (stationary cells will grow again when placed in new TAP media)
Unique Characteristics of Chlamydomonas
Has cell wall and chloroplast, although not a plant.
Although not an animal, chlamy also have flagella, plants lost flagella but chlamy retained it. (we also have flagella - sperm)
Prokaryotic (bacterial) flagella spins and has a hook while eukaryotic flagella waves and has microtubules (made from alpha-beta tubulin dimers)
They are homologs (flagella being analogous) but they are nothing like each other.
Dynein is a motor protein that uses ATP to "walk" towards negative end of microtubule, causing it to bend and the flagella to move like a whip. Therefor there are also many genes to code for dynein
So, flagella across eukaryotes is homologous (plants, fungi, animals), however prokaryotic flagella is analogous (bacteria)
Complexity of Flagella
Flagella structure in both eukaryotes and differences from prokaryotic flagella.
Motility importance and dynein protein contributing to movement in both species.
Identifiable similarities in proteins between clammy and humans indicate a common evolutionary heritage.
Cilia vs. Flagella
Understanding the structure and function of cilia, often confused with flagella.
Non-motile cilia role in sensory functions in humans (eyesight, olfaction, hearing).
Understanding defects leads to identification of ciliopathies (diseases linked to cilia dysfunction).
Model Organism Significance
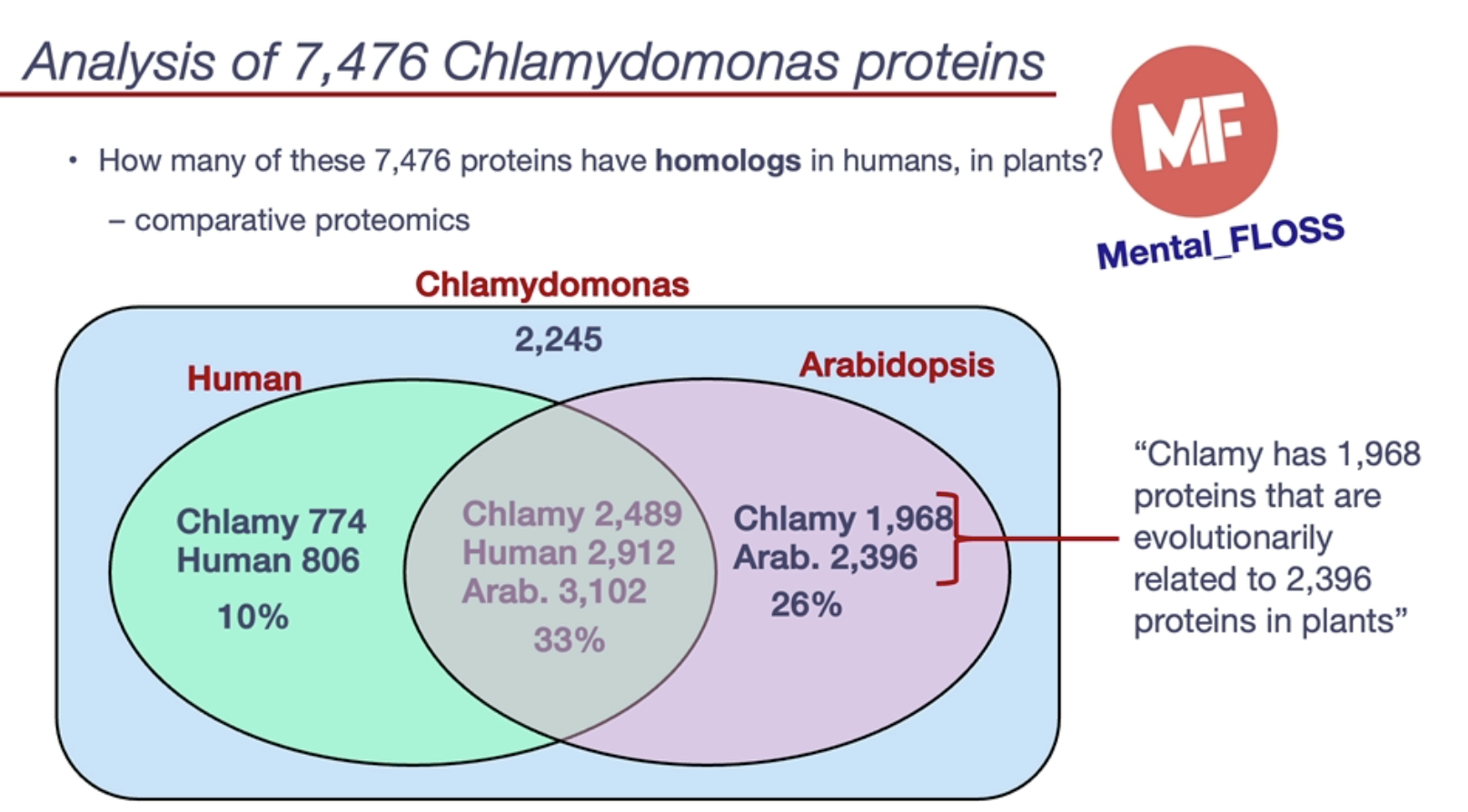
Chlamydomonas has its genome sequenced, allowing for extensive comparative studies in proteomics.
Around 26% of its proteins are homologous to Arabidopsis (plant model) proteins.
Only 10% homologous to human proteins, revealing evolutionary divergence.
What’s common? (“like fundamentals of eukaryotes in the centre”) U2 protein - a part of U2 snRNP which is used with U2 spliceosomal RNA