Nucleic Acids
^^Nucleic Acids^^
- example of polymer
- storing information of living organisms
- biologically occurring polynucleotides
- poly - many
- nucleotides - monomeric/building block (repeating units of nucleotides)
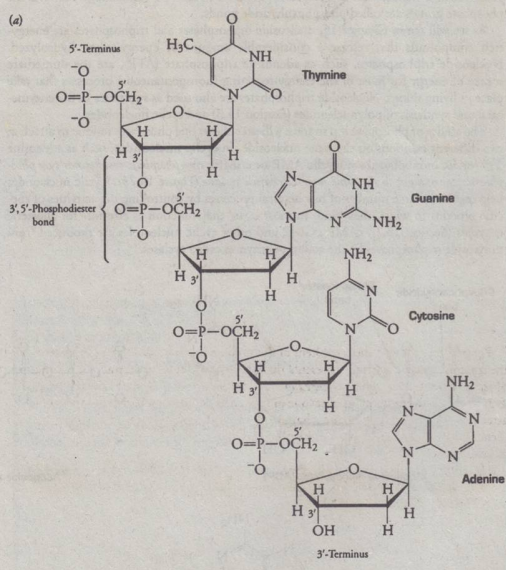
- nucleotide residues are linked in a specific sequence by phosphodiester bonds
- phospho - phosphate ion
- di ester bonds - 2 ester bonds
^^Monomeric unit^^
- nucleotide
^^Properties of Nucleic Acids^^
- High molecular weight
- Made up of CHONP (Carbon, Hydrogen, Oxygen, Nitrogen, Phosphorus)
- Upon hydrolysis; nucleotides
- Further hydrolysis; phosphoric acid and nucleosides
^^Nucleotides^^
- have their own building block
- 1 nucleotide = phosphate group, pentose and nitrogen containing base
- Component of Nucleotide: Phosphorylated Nucleoside
^^Nucleosides^^
- combination of pentose and base (hydrolize)
- pentose - 5 carbon sugar
- base - nitrogen containing base
^^Functions of Nucleic Acids^^
- serves as energy stores for future use in phosphate transfer reactions
- ATP is an example of nucleotides since it is composed of a phosphate group, pentose and nitrogen containing base
- Forming a portion of several important coenzymes
- -NAD+ (Nicotinamide Adenine Dinucleotide)
- NADP+ (Nicotinamide Adenine Dinucleotide Phosphate)
- FAD (Flavin Adenine Dinucleotide)
- Serving as mediators of numerous important cellular processes
- any energy that will be needed by the cell in any cellular reaction, it will make use of ATP
^^Basic Structure of Nucleic Acids^^
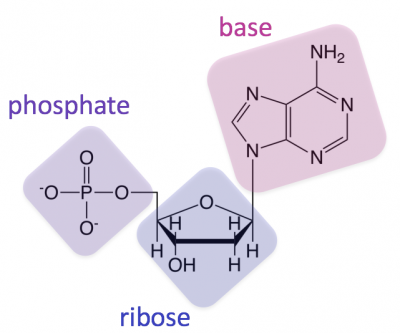
^^DNA or RNA?^^
- look at the pentose; locate the second carbon atom (encircled part)
- OH - RNA (ribose)
- H - DNA (deoxyribose)
^^Each nucleotide has 3 parts;^^
- Pentose (sugar)
- Phosphate group (phosphoric acid)
- organic nitrogen containing base (diff. bases found in DNA & RNA)
There are five different types of nucleotides
- the number of the types of nucleotide depends on the number of the bases
- DNA (A, G, C, T)
- RNA (A, G, C, U)
Information is encoded in the nucleic acid by different sequences of these nucleotides
- depends of what would be the sequence of amino acid
Structure of Nitrogenous bases
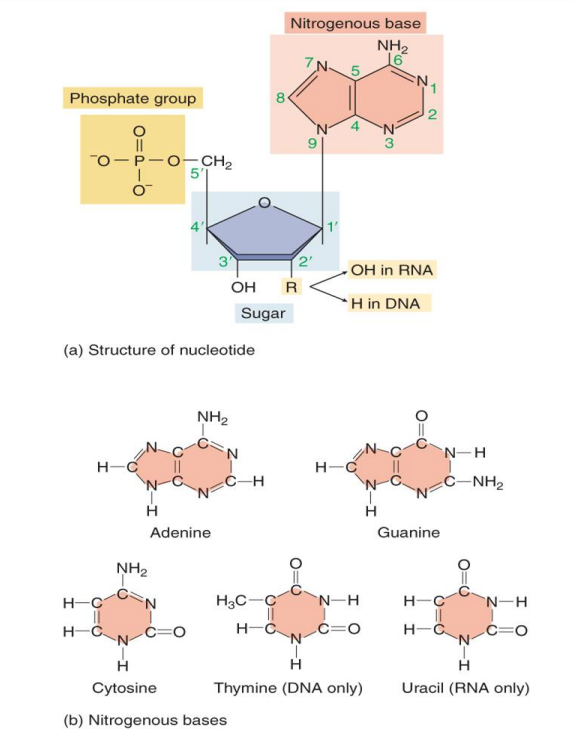
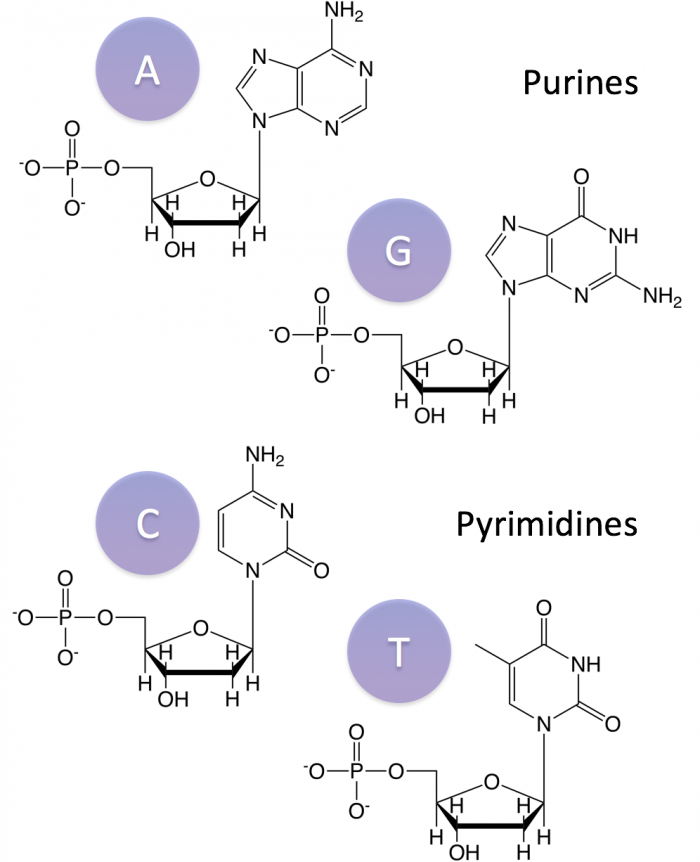
Purines
- short name but big structure
- 2 rings
- Adenine & Guanine (similar in structure)
Pyrimidine
- long name but small structure (1 structure)
- 1 ring
- Cytosine, Thymine, Uracil
Building Blocks of Nucleotide
- Pentose
- Phosphate group
- Organic nitrogen-containing base
Building blocks of Nucleotide: Monosaccharide
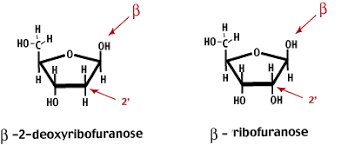
Difference of the two;
- Presence of OH in Ribose
- Presence of H in Deoxyribose
Furan
- 5 sides
- similar to pyran (6 sides)
- same shape during formation of cyclic form or ring form
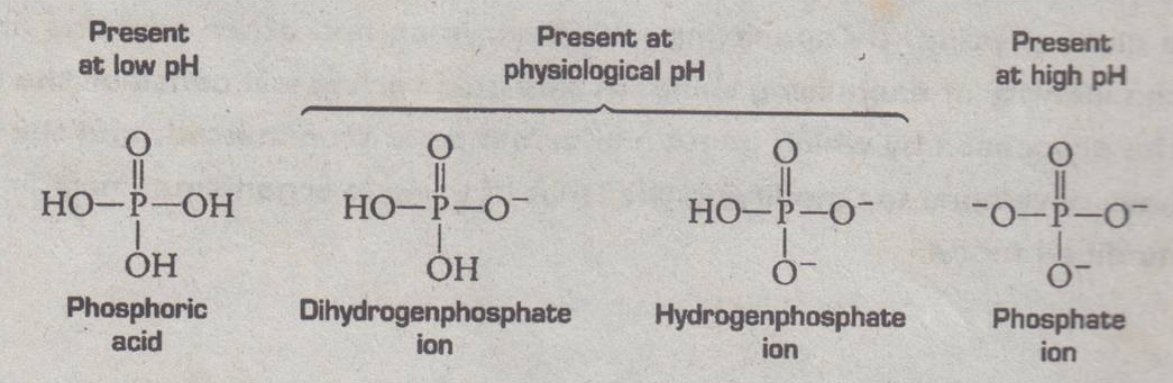
Building Blocks of Nucleotide: Phosphoric Acid
- not existing as phosphoric acid
- not occurring inside the body because of physiological PH that we have (7.35-7.45)
- existing either as hydrogen phosphate or 2 hydrogen atoms
- Proteins will be destroyed first
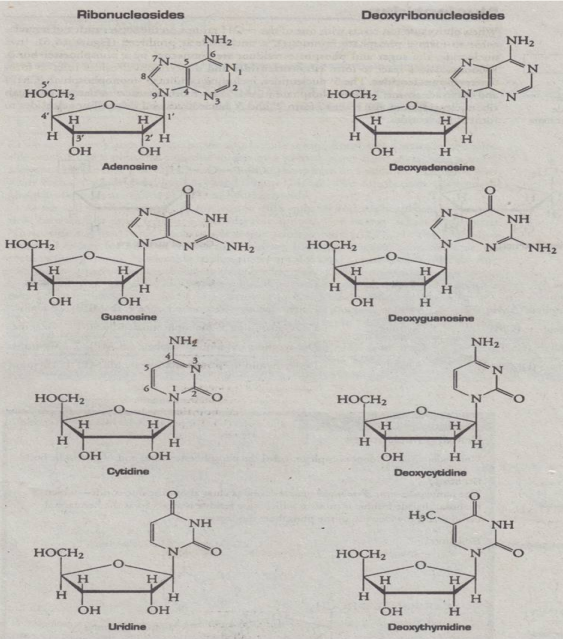
Nucleosides
Nucleotides
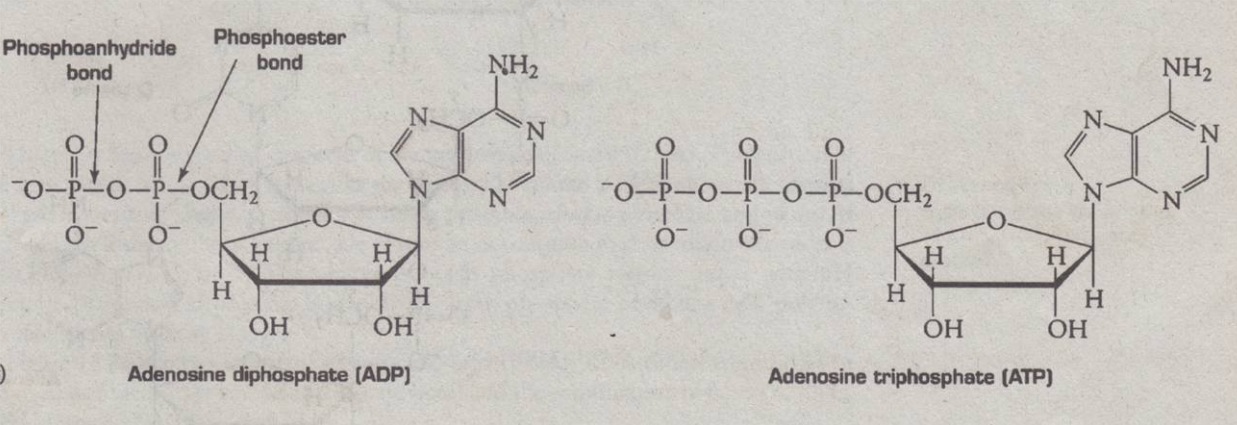
- P=O (Phosphorus double bond oxygen); it locate ester bonds
Phosphoester bond
- responsible for holding the phosphate group and the sugar
If several group is attached to a sugar
- it is being held by phosphoanhydride bond
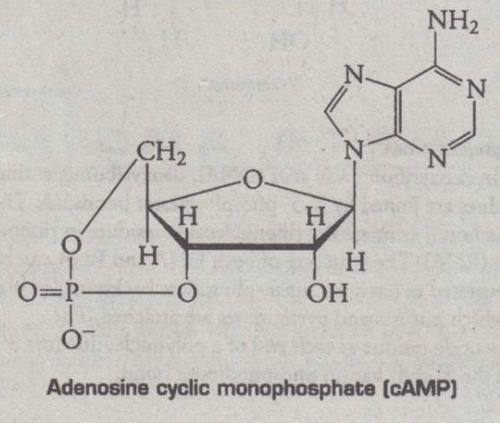
Nucleotides having more than 1 phosphate group
- used by cells as a source of energy (ATP)
Release of energy from ATP
- removal of phosphate group giving rise to Adenosine Diphosphate
- Phosphate group is connected to 2 carbon atoms of the pentose 3’,5’
Polynucleotides
many/multiple
- held between connection of phosphate group and sugar then bases are dangling lol
- P-OCH2 (already existing because it is part of 2nd nucleotide)
- backbone of nucleic acids
Significance of 3 prime / 5 prime
- number of carbon atoms that will be involve in the connection
3 prime (3’)
- 3rd carbon atom of a first nucleotide
5 prime (5’)
- 5th carbon atom of a sugar in a second nucleotide
O=P
- determining the location of phosphodiester bonds
Formation of 2 ester bonds
- 3rd & 5th connected to phosphate group
Types of Nucleic Acids
- Two types of nucleic acids; DNA & RNA
Deoxyribonucleic Acid (DNA)
- acting as an information molecule inside the nucleus
- more stable
- storage of information
- double helix (2 strands); interaction between the strands
- sugar deoxyribose
- A G C T
Ribonucleic Acid (RNA)
- protein synthesis / translation
- single strand (1 strand)
- inside & outside of nucleus
- sugar ribose
- A G C U
Complementary base pairing
- interaction between the base (own pair)
- A & T
- G & C
DNA Structure
- Primary structure - sequence of its nucleotide residues (simple sequence)
- Secondary structure - helix formed by the interaction of two DNA strands (hydrogen bonds are responsible for formation/folding)
- Tertiary Structure - arises from supercoiling, double helices being twisted into tighter, more compact shapes
phosphoanhydride
DNA Primary Structure
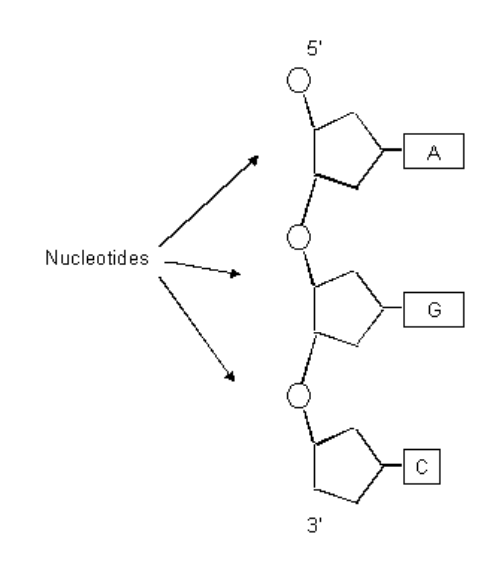
DNA Secondary Structure
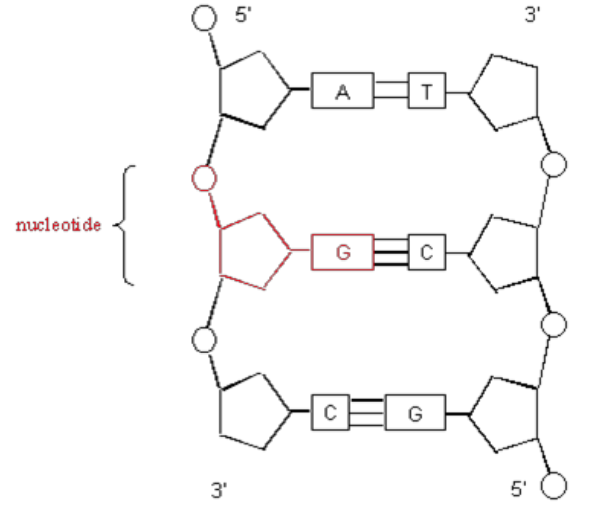
- antiparallel
DNA Tertiary Structure
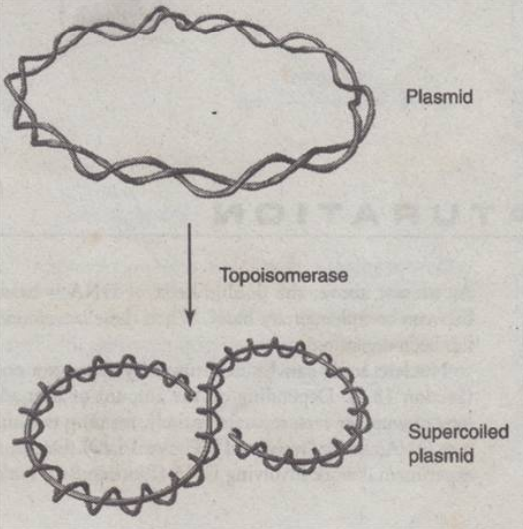
- Plasmid - exit of chromosomal structure; undergo supercoiling
- Supercoiled plasmid - compact, thicker
DNA Structure
- helps it to function
- hydrogen bonds of the base pairs can be broken to unzip the DNA so that information can be copied (can be replicated with appropriate enzymes)
- each strand of DNA is a mirror image so the DNA contains 2 copies of the information
- having 2 copies means that the information can be accurately copied and passed to the next generation (there’s still proofreading)
- DNA copied - RNA transcription (template will be copied; if there will be missing only one part, the sequence will change)
- Amino acids / protein produced will be dependent on the sequence of amino acid
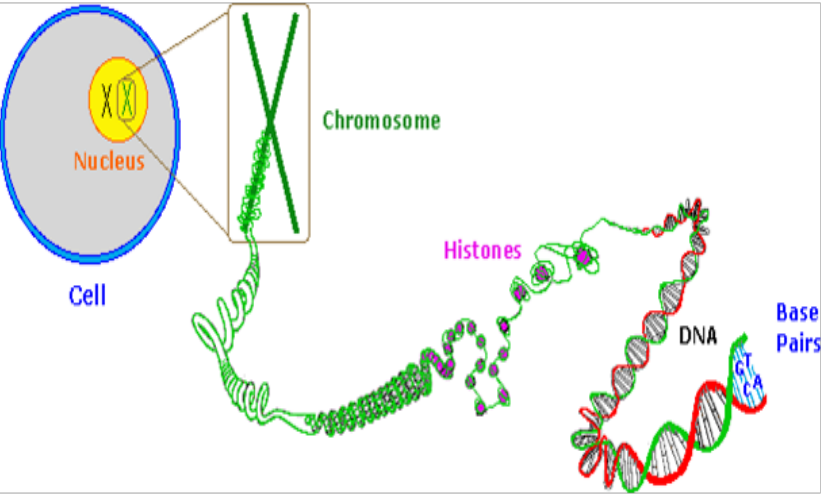
- chromosome - undergoes cell division
chromatin - not undergoing cell division
RNA
- contains different monosaccharide residue (ribose)
- DNA - inside the nucleus
- RNA - cytoplasm and nucleus
Types of RNA
- Transfer RNA (tRNA) - smallest of the three types of RNA, carry correct amino acid to the site of protein synthesis
- Messenger RNA (mRNA) - inside the nucleus; carry the information that specifies which protein should be synthesized or made
- Ribosomal RNA (rRNA) - combine with proteins to form ribosomes, multi subunit complexes in which protein synthesis takes place
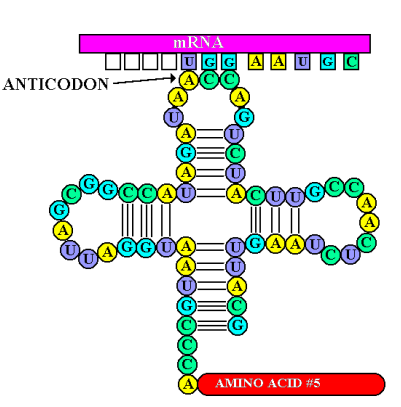
- being read by 3 bases= 1 codon
- anticodon - complementary base pairing
Nucleic Acid & Information Flow

\
- DNA cannot leave nucleus but it’s the source of information
- transcription - mRNA synthesis
- mRNA - can leave the nucleus; transcribed
- Translation - translate into a specific AA sequence
- Protein synthesis - cytoplasm
DNA Replication
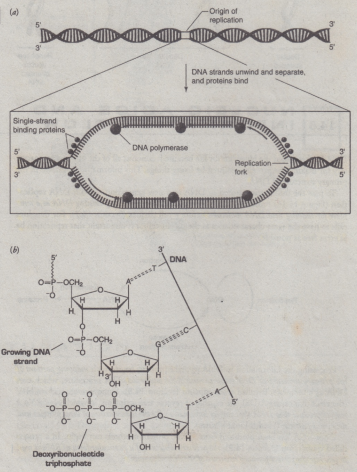
- the process by which the genome's DNA is copied in cells. Before a cell divides, it must first copy (or replicate) its entire genome so that each resulting daughter cell ends up with its own complete genome.
Step 1: Replication Fork Formation
- Before DNA can be replicated, the double stranded molecule must be “unzipped” into two single strands.
- DNA has four bases called adenine (A), thymine (T), cytosine (C) and guanine (G) that form pairs between the two strands.
- Adenine only pairs with thymine and cytosine only binds with guanine. In order to unwind DNA, these interactions between base pairs must be broken.
- This is performed by an enzyme known as DNA helicase. DNA helicase disrupts the hydrogen bonding between base pairs to separate the strands into a Y shape known as the replication fork. This area will be the template for replication to begin.
- However, the replication fork is bi-directional; one strand is oriented in the 3' to 5' direction (leading strand) while the other is oriented 5' to 3' (lagging strand). The two sides are therefore replicated with two different processes to accommodate the directional difference.
Step 2: Primer Binding
- simplest to replicate. Once the DNA strands have been separated, a short piece of RNA called a primer binds to the 3' end of the strand. The primer always binds as the starting point for replication. Primers are generated by the enzyme DNA primase.
Step 3: Elongation
- Enzymes known as DNA polymerases are responsible creating the new strand by a process called elongation.
- There are five different known types of DNA polymerases in bacteria and human cells. In bacteria such as E. coli, polymerase III is the main replication enzyme, while polymerase I, II, IV and V are responsible for error checking and repair.
- DNA polymerase III binds to the strand at the site of the primer and begins adding new base pairs complementary to the strand during replication.
- lagging strand begins replication by binding with multiple primers. Each primer is only several bases apart.
- DNA polymerase then adds pieces of DNA, called Okazaki fragments, to the strand between primers. This process of replication is discontinuous as the newly created fragments are disjointed.
Step 4: Termination
- Once both the continuous and discontinuous strands are formed, an enzyme called exonuclease removes all RNA primers from the original strands. These primers are then replaced with appropriate bases.
- Another exonuclease “proofreads” the newly formed DNA to check, remove and replace any errors.
- Another enzyme called DNA ligase joins Okazaki fragments together forming a single unified strand. The ends of the linear DNA present a problem as DNA polymerase can only add nucleotides in the 5′ to 3′ direction. The ends of the parent strands consist of repeated DNA sequences called telomeres.
- A special type of DNA polymerase enzyme called telomerase catalyzes the synthesis of telomere sequences at the ends of the DNA. Once completed, the parent strand and its complementary DNA strand coils into the familiar double helix shape. In the end, replication produces two DNA molecules, each with one strand from the parent molecule and one new strand.
Transcription 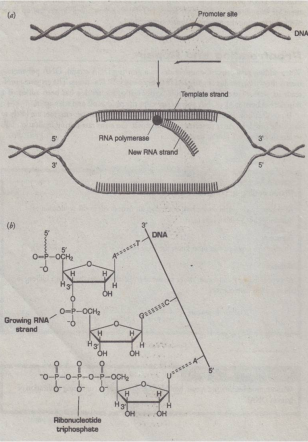
- the process of making an RNA copy of a gene's DNA sequence. This copy, called messenger RNA (mRNA), carries the gene's protein information encoded in DNA.
- DNA transcription is the process by which the genetic information contained within DNA is re-written into messenger RNA (mRNA) by RNA polymerase. This mRNA then exits the nucleus, where it acts as the basis for the translation of DNA. By controlling the production of mRNA within the nucleus, the cell regulates the rate of gene expression.
- follows complementary base pairing
- mRNA synthesis
Translation
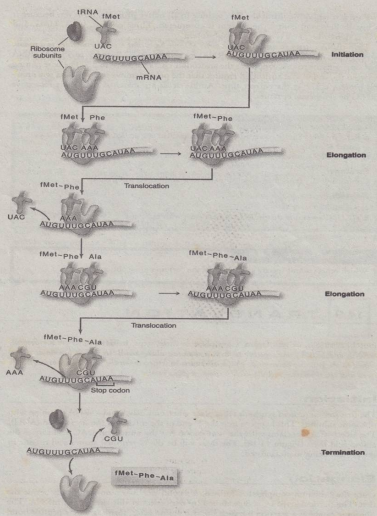
- also known as protein synthesis
- the process that takes the information passed from DNA as messenger RNA and turns it into a series of amino acids bound together with peptide bonds. It is essentially a translation from one code (nucleotide sequence) to another code (amino acid sequence).
Steps of translation:
- tRNA charging:
1. The process of attaching an amino acid to its respective transfer RNA (tRNA) is known as amino acid activation, also known as aminoacylation or tRNA charging
- Initiation:
1. In the initiation step, the charged tRNA attaches to the start codon (AUG), the small subunit of ribosome binds to the mRNA, and finally, the large ribosomal subunit binds to create the initiation complex.
- Elongation:
1. According to the codons found in the mRNA, the polypeptide chain keeps growing. 2. Each amino acid has a peptide bond attaching it to the growing chain. 3. Elongation continues till the whole gene is translated.
- Termination:
1. When the ribosome reaches a stop codon, such as UAA, UAG, or UGA, translation is finished since these codons lack tRNAs. 2. When this happens, the translation stops, and the newly produced polypeptide chain is released.
Genetic Code
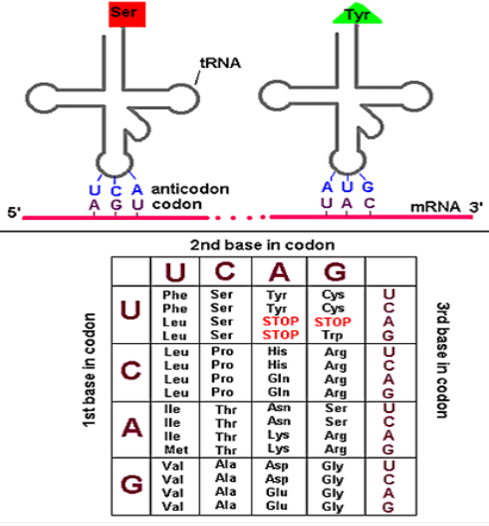
- AUG - Methionine
- UUU - Phenylalanine
- UAA - Stop (Signal molecule)
- CAG - Glutamine
- Stop codons in genetic code - UAG, UAA & UGA
\