BIOL 3130 LEC 9
1) What is the Mediator complex and where is it found?
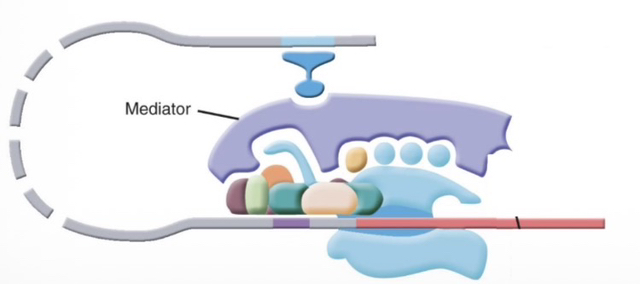
A large protein complex (20-30 subunits) associated with the PIC at active RNAP II promoters. It is vital for activated transcription in living cells (in vivo).
2) Is Mediator a GTF or a Coactivator?
It acts like a GTF (present at almost all active promoters).
It acts like a Coactivator (doesn't bind DNA, needed for activation, interacts with ssTFs/activators).
Note: It is not a "genuine" GTF because it is not strictly required for transcription in a test tube (in vitro).
3) How does the Mediator physically connect distant DNA elements?
It makes contacts with RNAP II (specifically the CTD), GTFs, and activators. It acts as a "molecular bridge," facilitating a DNA loop between enhancers and promoters.
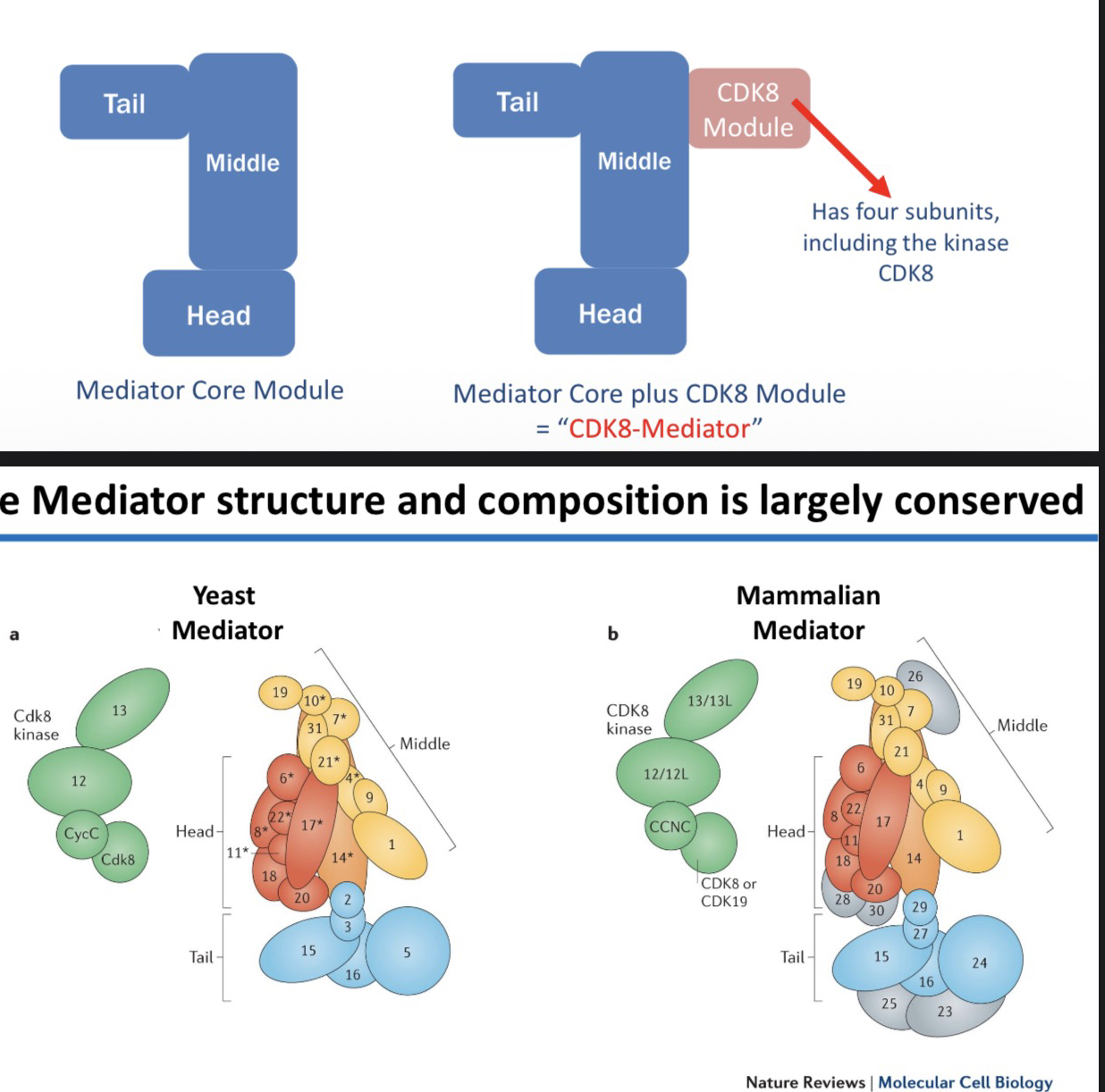
4) Describe the two main components of the Mediator complex.
Core Module (Head, Middle, Tail).
CDK8 Module (four subunits, including the CDK8 kinase).
Note: The exact composition can change depending on the gene/context.
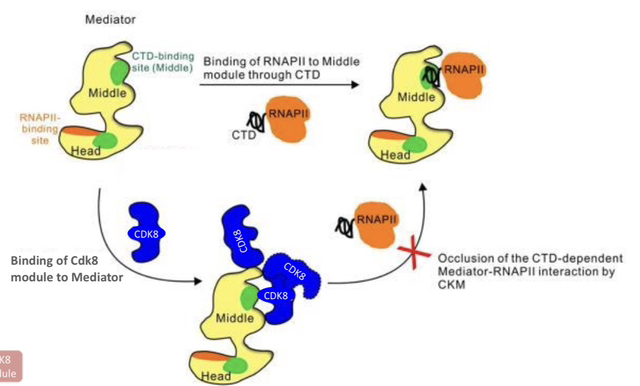
5) What is the key interaction rule between Mediator Core, the CDK8 Module, and RNAP II?
The Mediator core associates with either the CDK8 Module OR RNAP II, but never both at the same time. Binding is mutually exclusive.
6) What are the proposed functions of the CDK8 module after RNAP II has cleared the promoter?
Helping RNAP II overcome pausing during elongation.
Helping to reset the promoter for transcription re-initiation.
7) What must happen for transcription initiation and promoter clearance to occur?
RNAP II must be released from the Mediator complex. Mediator helps recruit and position TFIIH, whose kinase phosphorylates the CTD, causing this release.
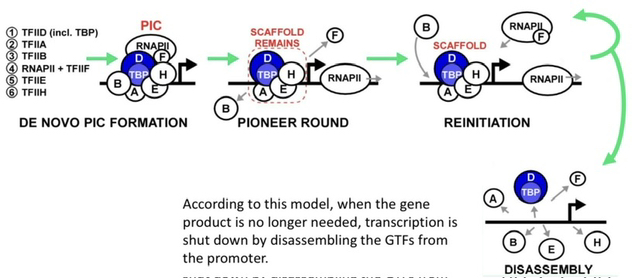
8) How does the cell efficiently perform multiple rounds of transcription without starting from scratch every time?
A "scaffold" of some GTFs remains at the promoter after RNAP II clears. This scaffold (TFIID, TFIIA, TFIIE, TFIIH) enables faster reassembly of a new PIC (transcription re-initiation).
9) Where does Promoter-Proximal Pausing occur, and which two factors are responsible for establishing the pause?
It occurs shortly downstream of the promoter (usually the +20 to +60 region). The pause depends on the association of negative elongation factors: NELF and DSIF.
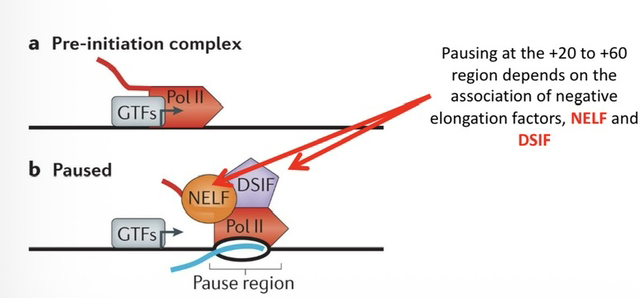
10) What complex is responsible for releasing the pause, and what is its active component?
P-TEFb (Positive Elongation Factor b). It contains a kinase called CDK9.
11) What are the three specific targets phosphorylated by CDK9 to relieve pausing?
NELF: Phosphorylation causes it to be released from the complex.
DSIF: Phosphorylation converts it into an elongation-promoting factor.
RNAP II CTD: Specifically targets Ser2 of the heptad repeats.
12) How does "poising" (pausing) RNAP II help with Synchronous Gene Activation?
It allows a population of cells to turn on a gene at the same time. Because the polymerase has already initiated, the gene can go from "Off" to "On" very rapidly once the signal to release the pause arrives.
13) What is the relationship between pausing and 5′ capping?
Capping occurs when the transcript is 20–60 nucleotides long. Pausing at this exact stage provides a "window" for the Capping Enzyme Complex (CEC) to associate and attach the 5′ cap to the pre-mRNA.
14) How does a paused RNAP II affect chromatin?
The paused molecule serves as a platform to recruit machinery (like remodellers or histone methyltransferases) that alter chromatin to a permissive (open/accessible) state. This makes future rounds of transcription more efficient.
15) What is the SEC, and how does it relate to the CDK8 module?
The SEC contains proteins that promote elongation, including P-TEFb. It is hypothesized that the CDK8 module helps overcome pausing by recruiting the SEC to the promoter after RNAP II has cleared it.
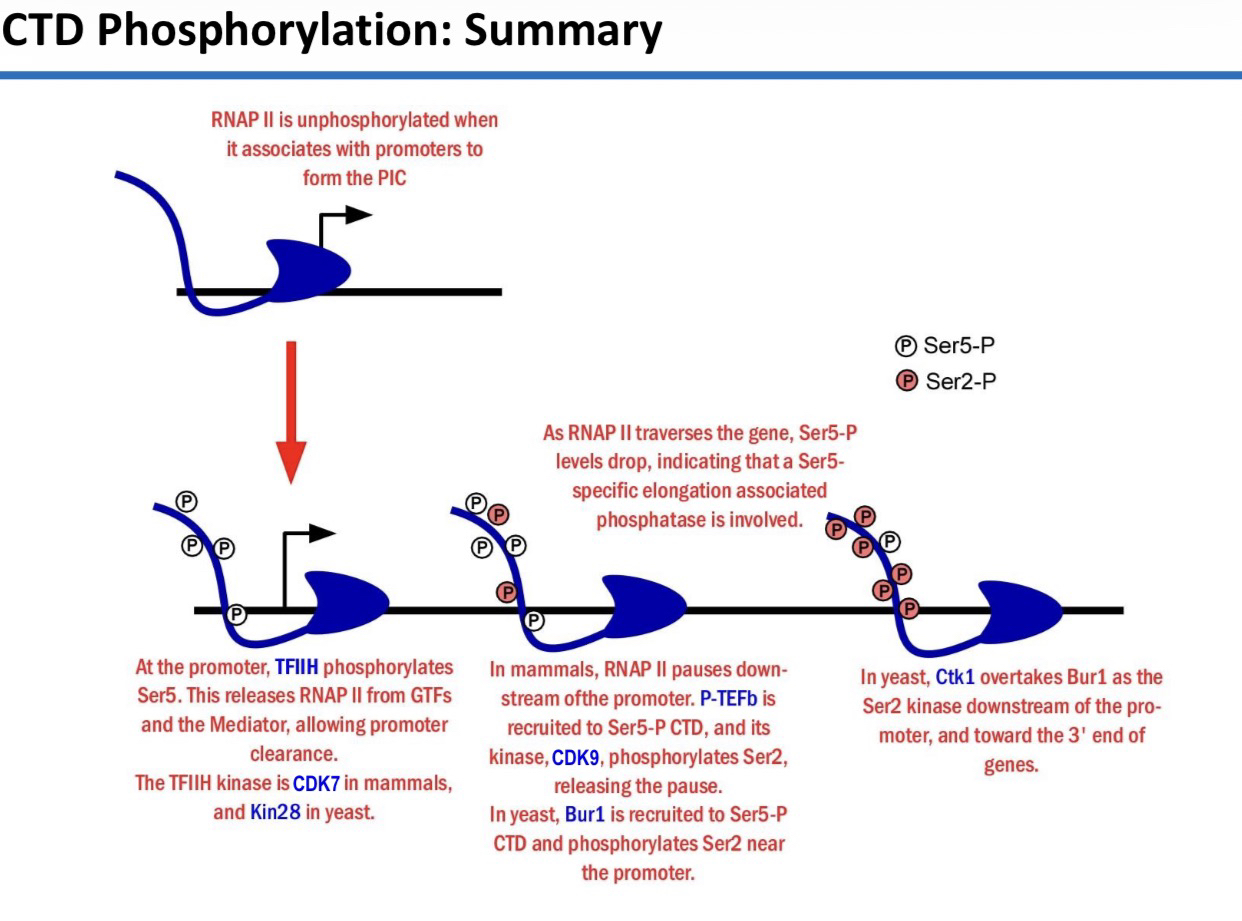
16) Compare the CTD phosphorylation performed by TFIIH vs. P-TEFb.
TFIIH (CDK7/Kin28): Phosphorylates Ser5; associated with initiation and promoter clearance.
P-TEFb (CDK9): Phosphorylates Ser2; associated with elongation and releasing the pause.
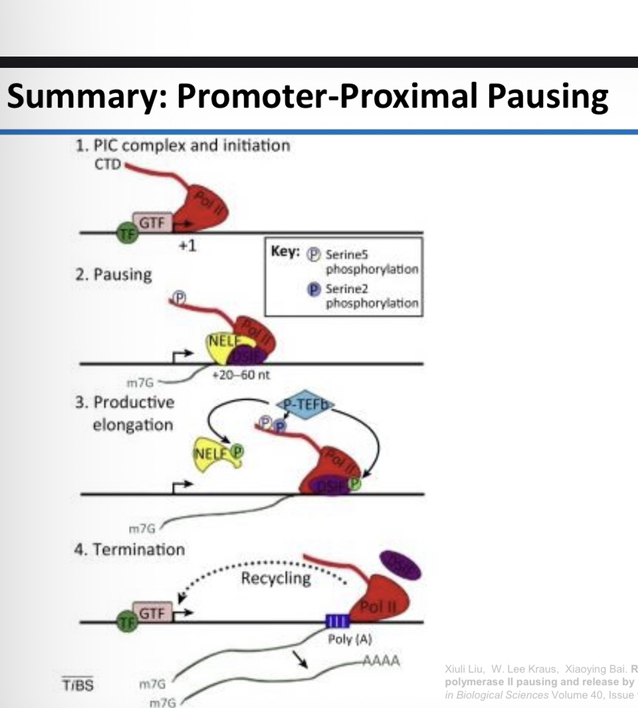
17) Compare the phosphorylation state of the CTD when RNAP II binds a promoter versus when it leaves the promoter.
At the Promoter (PIC): It is hypophosphorylated (little or no phosphorylation; RNAP IIa).
Leaving the Promoter: It is hyperphosphorylated (heavily phosphorylated; RNAP IIo).
18) How did scientists determine that the CTD pattern changes across a gene, and what were the two main "markers" found?
They used ChIP-seq with antibodies specific to each phosphorylated amino acid (Tyr1, Ser2, Thr4, Ser5, Ser7).
Ser5-P is concentrated at the 5′ end (start).
Ser2-P increases toward the 3′ end (finish).
19) Which complex phosphorylates Ser5, and what are the specific kinase names in mammals and yeast?
TFIIH.
Mammals: CDK7.
Yeast: Kin28.
Function: Allows RNAP II to break contact with GTFs/Mediator and clear the promoter
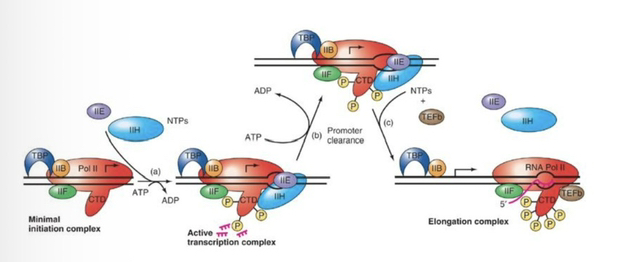
20) In mammals, which kinase phosphorylates Ser2, and what is its biological role?
CDK9 (part of the P-TEFb complex).
Role: It phosphorylates Ser2 to overcome promoter-proximal pausing, allowing RNAP II to engage in elongation.
21) How does the phosphorylation of Ser5 affect Ser2 in mammals?
The recruitment of P-TEFb (the Ser2 kinase) depends on the prior phosphorylation of Ser5. Therefore, Ser5-P is a prerequisite for Ser2-P.
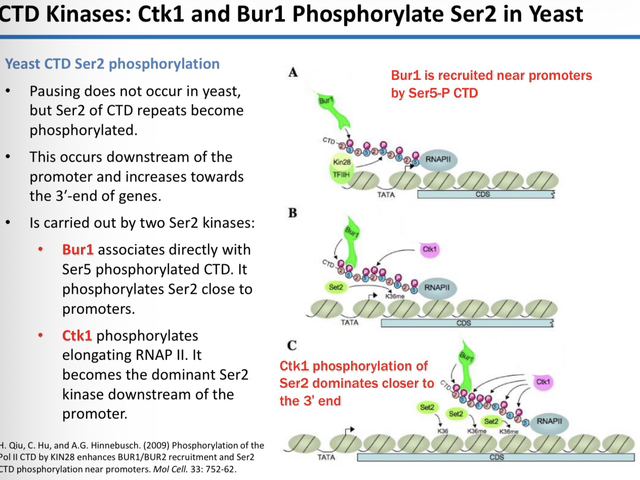
22) Name the two kinases responsible for Ser2 phosphorylation in yeast and their specific locations.
Bur1: Recruited by Ser5-P; phosphorylates Ser2 near the promoter.
Ctk1: The dominant Ser2 kinase; phosphorylates the elongating RNAP II further downstream toward the 3′ end.
23) Based on the lecture, what is a major difference in the elongation phase between yeast and mammals?
Promoter-proximal pausing occurs in mammals (requiring CDK9 to release it), but it does not occur in yeast.
24) Which amino acids in the Y1-S2-P3-T4-S5-P6-S7repeat can be phosphorylated?
All non-proline residues (Tyr1, Ser2, Thr4, Ser5, and Ser7) have been detected as phosphorylated forms.
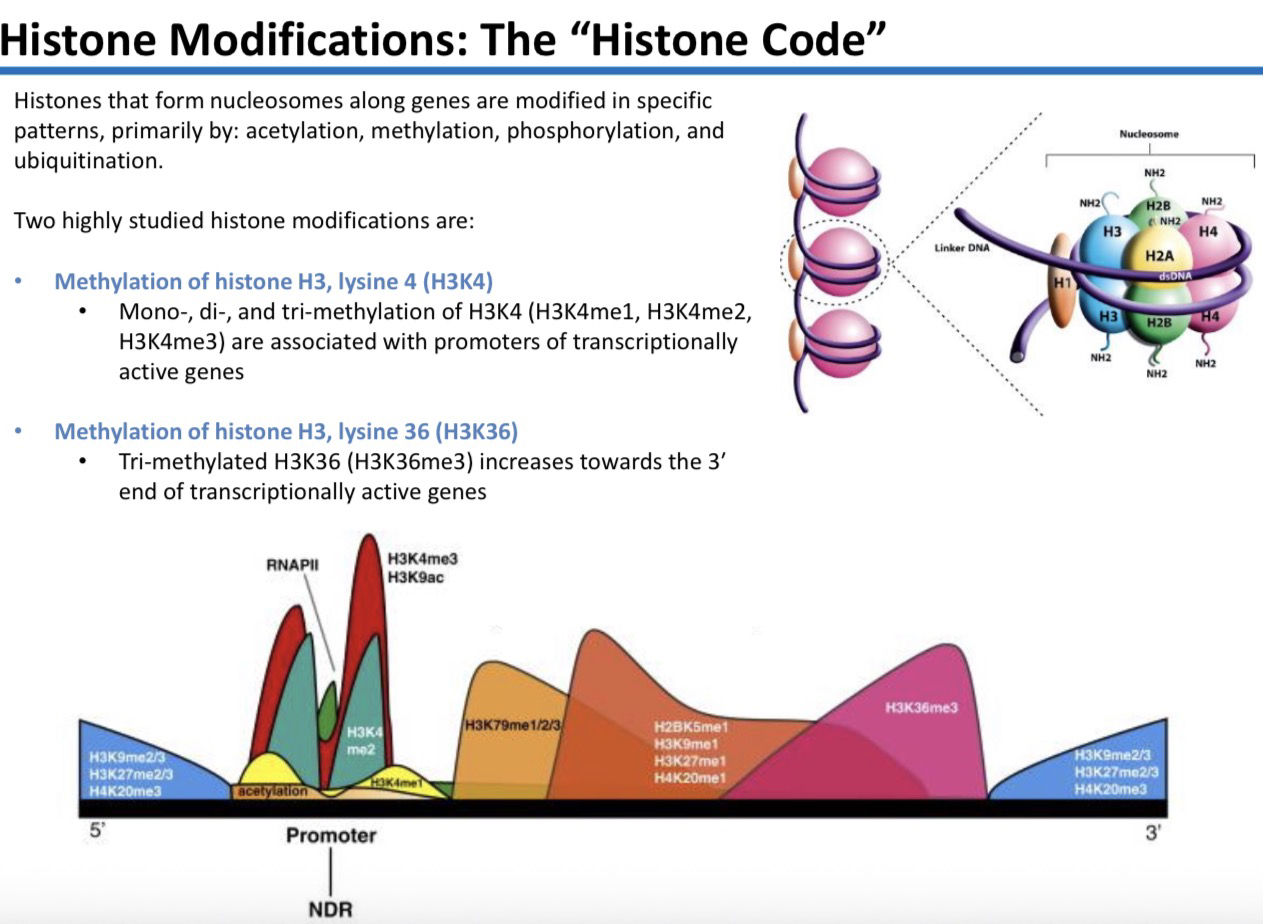
25) What are the four primary ways histones are modified to create the "Histone Code"?
Acetylation
Methylation
Phosphorylation
Ubiquitination
26) Describe the pattern and meaning of H3K4 methylation (mono-, di-, or tri-).
It is primarily associated with the promoters of transcriptionally active genes. It is placed at the 5′ end of the gene.
27) Describe the pattern of H3K36me3 (tri-methylation).
Its concentration increases towards the 3′ end of transcriptionally active genes. It marks the "body" and end of the gene.
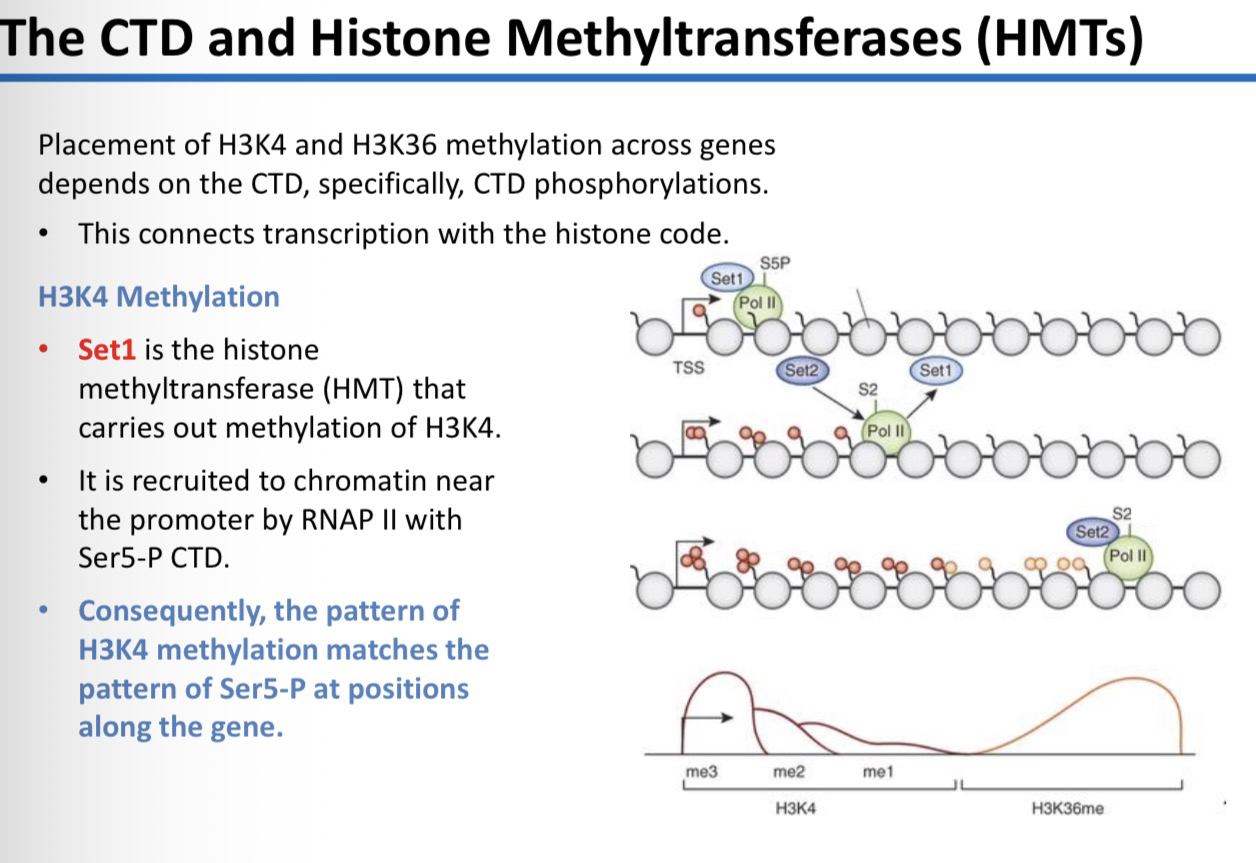
28) Which enzyme carries out H3K4 methylation and how is it recruited?
Set1 (a histone methyltransferase or HMT). It is recruited to chromatin near the promoter by RNAP II that has a Ser5-P CTD.
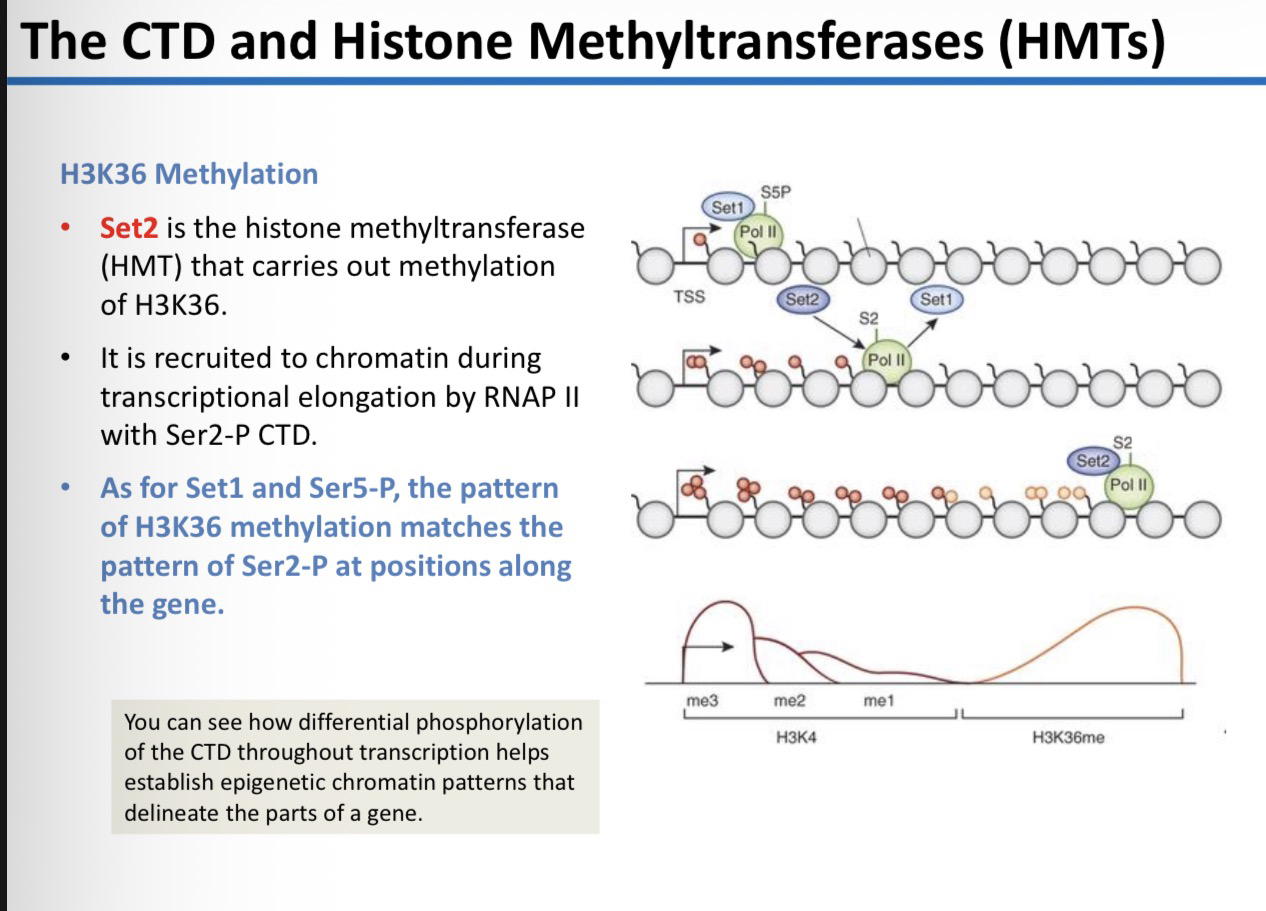
29) Which enzyme carries out H3K36 methylation and how is it recruited?
Set2. It is recruited to chromatin during transcriptional elongation by RNAP II that has a Ser2-P CTD.
30) How does CTD phosphorylation coordinate the physical processing of the RNA transcript?
Ser5-P: Recruits capping enzymes to the 5′ end of the pre-mRNA.
Ser2-P: Recruits cleavage and polyadenylation machineryto the 3′ end of the pre-mRNA.
31) How does the CTD help establish epigenetic chromatin patterns?
Differential phosphorylation (switching from Ser5 to Ser2) ensures that specific HMTs (Set1 and Set2) are recruited at the right time and place, creating a spatial map of histone marks that delineate the parts of a gene.
32) Name the kinases responsible for the Ser5 and Ser2 signals.
Ser5-P: TFIIH (CDK7 in mammals / Kin28 in yeast).
Ser2-P: P-TEFb (CDK9) in mammals; Bur1 and Ctk1 in yeast.