DNA sequencing
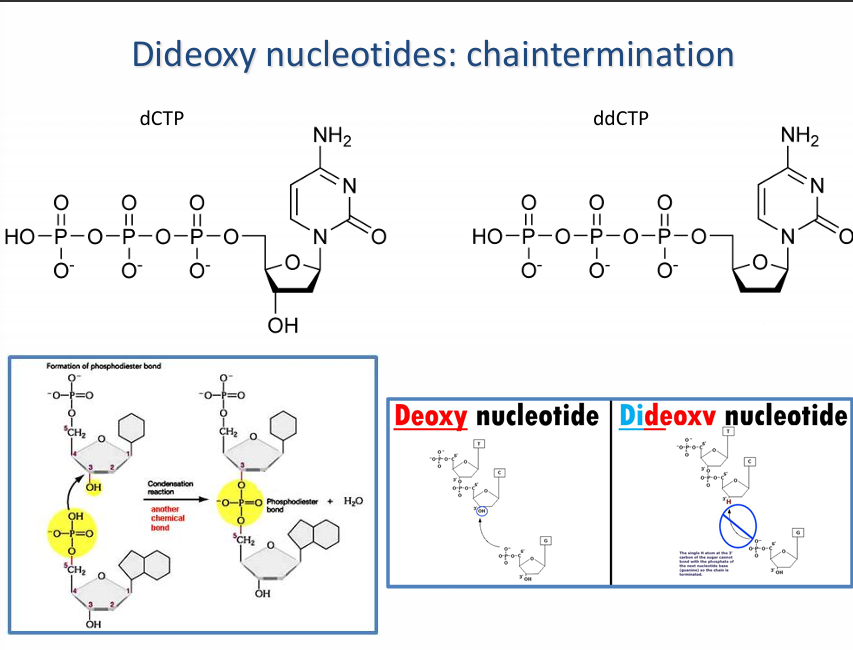
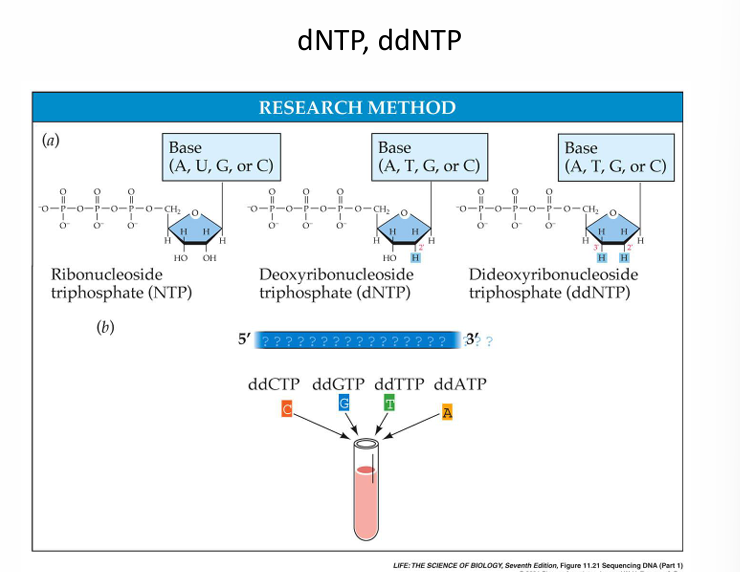
What is the difference between the deoxynucleotide and the dideoxynucleotide?
The deoxynucleotide is the building block of DNA. It has a 3′-OH group while the dideoxynucleotide lacks the 3′-OH needed to add another nucleotide.
Deoxynucleotide allows DNA synthesis, while dieoxynucleotide stops DNA synthesis.
The principle of chain termination sequencing
In chain-termination DNA sequencing, DNA synthesis is performed on a sample (template) using deoxy and dideoxy nucleotides.
Each reaction is PARTIAL; the chain-terminating dideoxy nucleotide is randomly incorporated into the newly synthesised DNA chain.
RESULT: DNA segments of different lengths with a common 5’ end (sequencing primer) and different 3 ‘ends (template DNA complement).
Components of the sequencing reaction
Template DNA (genomic, plasmid, phage, PCR product)
Sequencing primer (15-25 nucleotides in length determines the start point ).
dNTP (can be marked or unmarked).
ddNTP
Sequencing enzyme (DNA polymerase, e.g., T7, Taq).
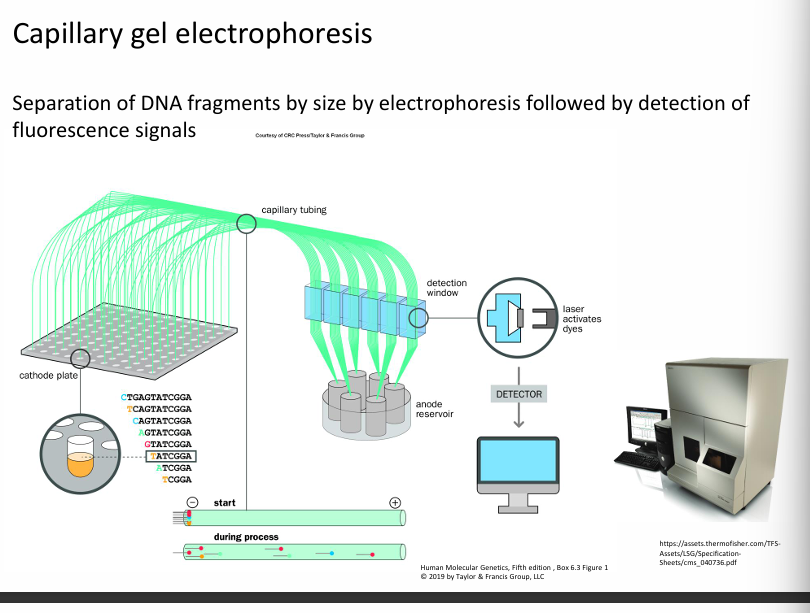
Electrophoresis
Different in one nucleotide from each other.
It is suitable for separating DNA fragments.
TWO METHODS OF ELECTROPHORESIS:
Denaturing Polyacrylamide gel electrophoresis.
Capillary electrophoresis.
Methods of detection- radioactive detection:

What the slide means
Your slide is about radioactive detection used in DNA sequencing (especially early Sanger sequencing).
So the roles are:
Step | What happens |
|---|---|
Electrophoresis | Separates DNA fragments by size |
Detection method | Allows us to see the DNA fragments after separation |
Radioactive detection process (what your slide lists)
Radiolabeled deoxynucleotide is incorporated
A nucleotide containing a radioactive isotope (often (^{32}P)) is added during DNA synthesis.
The DNA fragments become radioactively labelled.
Electrophoresis
DNA fragments are separated on a polyacrylamide gel.
Gel drying
The gel is dried onto filter paper to stabilise it.
Autoradiographic development
The gel is placed against X-ray film.
Radiation from the DNA exposes the film, producing bands.
Sequence reading
The band pattern is read from bottom to top to determine the DNA sequence.
Key clarification
These are methods of detection, not electrophoresis types.
Category | Examples |
|---|---|
Electrophoresis methods | PAGE, Capillary electrophoresis |
Detection methods | Radioactive labeling, Fluorescent labeling |
Modern sequencing mostly uses fluorescent detection instead of radioactive detection.
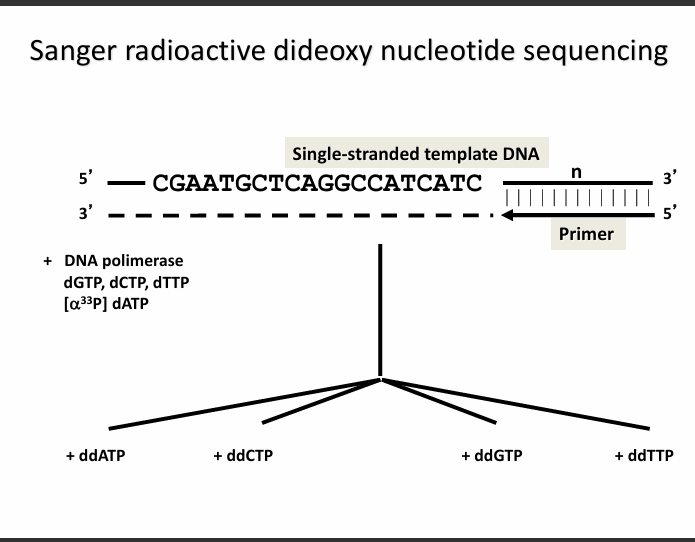
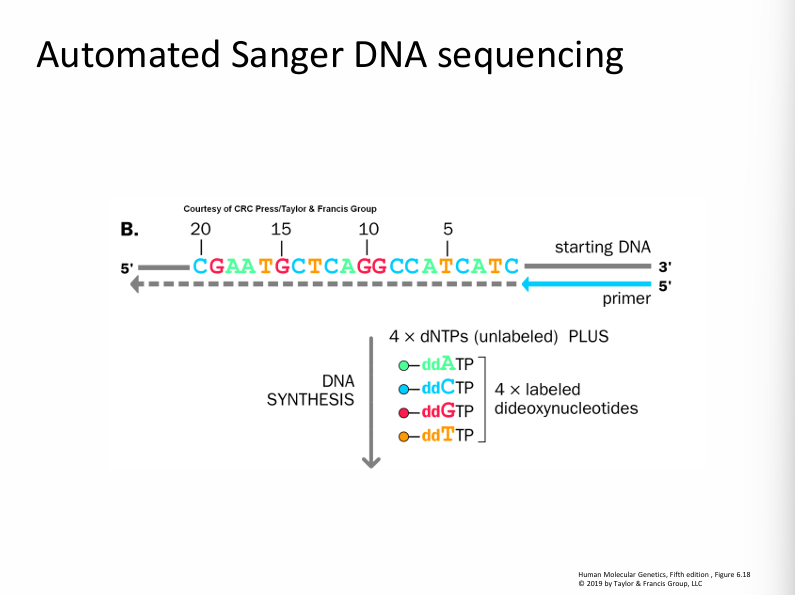
1. What this slide is showing (Setup of the experiment)


Components in the reaction
You start with:
1. Single-stranded template DNA
5' CGAATGCTCAGGCCATCATC 3'
This is the DNA sequence we want to determine.
2. Primer
A short DNA fragment that binds to the template.
DNA polymerase starts synthesis from the primer.
3. DNA polymerase
An enzyme that adds nucleotides to extend the new strand.
4. Normal nucleotides (dNTPs)
dATP, dCTP, dGTP, dTTP
Used to build the new DNA strand.
5. Radioactive nucleotide
Example: [α³³P] dATP
This labels the DNA so it can be detected later.
The key idea: four separate reactions
The mixture is divided into four tubes, each containing a different dideoxynucleotide:
Tube | Contains |
|---|---|
A reaction | ddATP |
C reaction | ddCTP |
G reaction | ddGTP |
T reaction | ddTTP |
Why ddNTPs matter
A dideoxynucleotide (ddNTP):
Stops DNA synthesis
because it lacks the 3'-OH group
So when a ddNTP is inserted:
➡ DNA cannot extend further
➡ The chain terminates
This produces DNA fragments of different lengths.
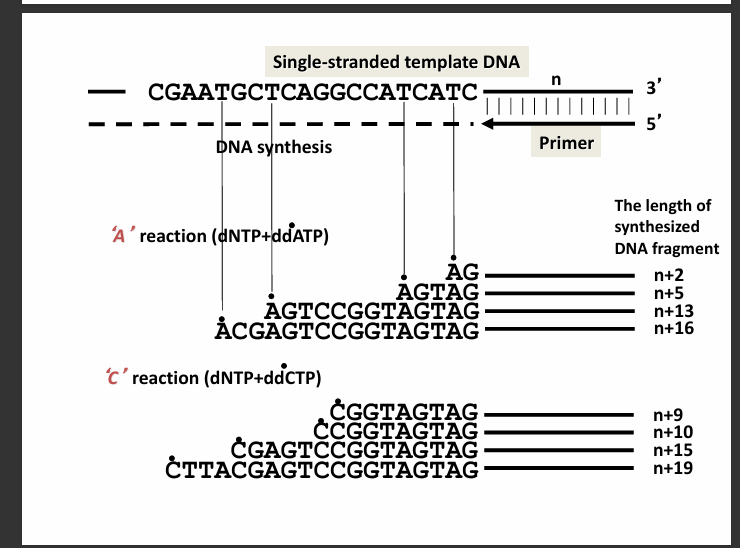
The slide shows what happens in two of the tubes.
A reaction (dNTP + ddATP)
Whenever an A should be added, sometimes ddATP is inserted instead of dATP.
When that happens:
➡ DNA synthesis stops at that position
So you get fragments ending at every A position.
Example fragments shown:
AG
AGTAG
AGTCCGGTAGTAG
ACGAGTCCGGTAGTAG
Each fragment ends with A.
C reaction (dNTP + ddCTP)
Same idea.
Whenever C should be added, ddCTP may terminate the chain.
Fragments end at C positions.
Example fragments:
CGGTAGTAG
CCGGTAGTAG
CGAGTCCGGTAGTAG
CTTACGAGTCCGGTAGTAG
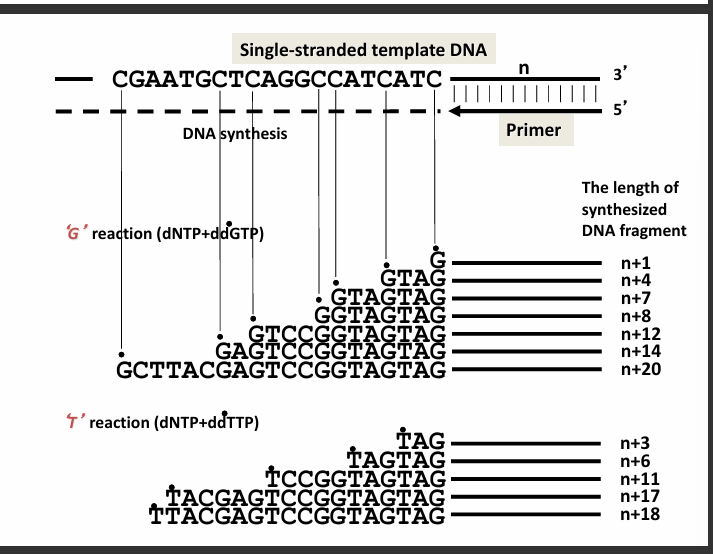
3. What this slide shows (G and T reactions)
The other two tubes work the same way.
G reaction (dNTP + ddGTP)
Fragments terminate at G.
Examples:
G
GTAG
GTAGTAG
GGGTAGTAG
T reaction (dNTP + ddTTP)
Fragments terminate at T.
Examples:
TAG
TAGTAG
TCCGGTAGTAG
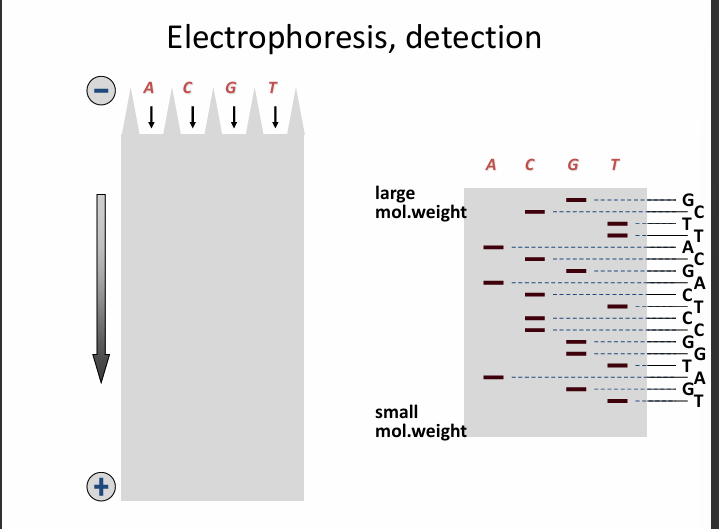
Electrophoresis and Detection:
1. Four lanes (A, C, G, T)
After the sequencing reactions, you have four tubes:
A reaction → fragments ending in A
C reaction → fragments ending in C
G reaction → fragments ending in G
T reaction → fragments ending in T
Each reaction is loaded into a separate lane of the gel.
A | C | G | T
2. DNA moves in the electric field
DNA is negatively charged because of its phosphate backbone.
So it moves:
from the negative electrode (top)
➡ toward the positive electrode (bottom)
3. Separation by size
During electrophoresis:
Small DNA fragments move faster
Large fragments move more slowly
So on the gel:
Position | Fragment size |
|---|---|
Top | large fragments |
Bottom | small fragments |
4. Bands appear after detection
After autoradiography, the gel shows bands where DNA fragments are located.
Each band corresponds to one DNA fragment length.
5. How the sequence is read
You read the bands from bottom → top.
Why?
Because:
Bottom = shortest fragment
Shortest fragment = first nucleotide added after the primer
Example reading:
Bottom band → T
Next band → G
Next band → A
Next band → T
So the sequence becomes:
TGAT...
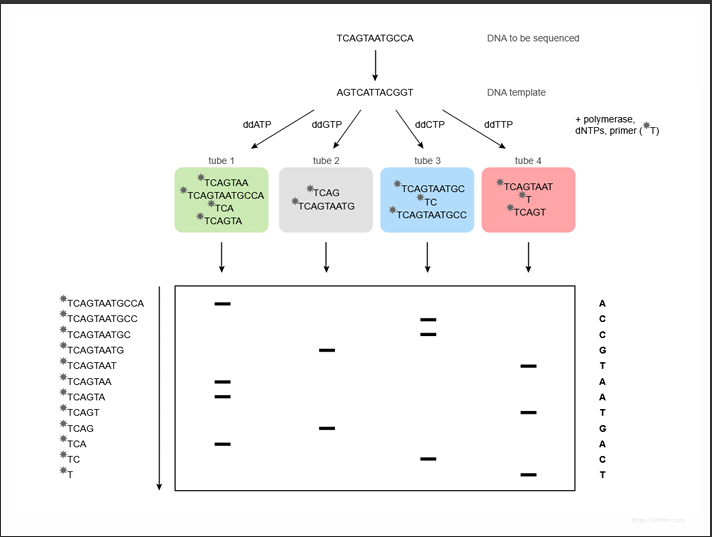
How the fragments were produced
This slide shows how those fragments in the gel were created.
1. DNA template
The DNA you want to sequence is:
TCAGTAATGCCA
DNA polymerase synthesises the complementary strand.
2. Four sequencing reactions
The reaction mixture is split into four tubes:
Tube | Contains |
|---|---|
Tube 1 | ddATP |
Tube 2 | ddGTP |
Tube 3 | ddCTP |
Tube 4 | ddTTP |
Each tube also contains:
DNA polymerase
primer
normal nucleotides (dNTPs)
3. Chain termination
Sometimes a ddNTP is incorporated instead of a normal nucleotide.
When this happens:
❌ DNA synthesis stops
because ddNTP lacks the 3'-OH group.
So fragments of different lengths are produced.
Example in the ddATP tube:
TCAGTAA
TCAGTAAT
TCAGTAATGCCA
All fragments end with A.
4. Running the gel
Fragments from each tube are loaded into the gel lanes:
A C G T
The gel separates fragments by size.
5. Reading the final sequence
Reading the bands from bottom → top gives:
A
C
C
G
T
A
A
T
G
A
C
T
So the sequence is:
ACCGTAATGACT
The key concept of the slides
Sanger sequencing works because:
ddNTP randomly stops DNA synthesis
This creates DNA fragments of different lengths
Electrophoresis separates them
The band pattern reveals the DNA sequence
✅ The most important rule to remember
Always read Sanger sequencing gels from bottom → top.
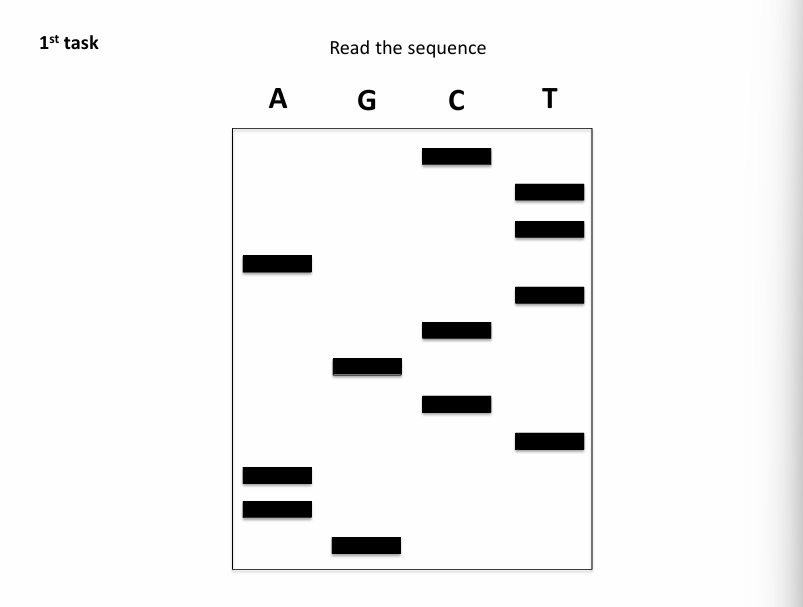
ANSWER:
GAATCGCTATTC
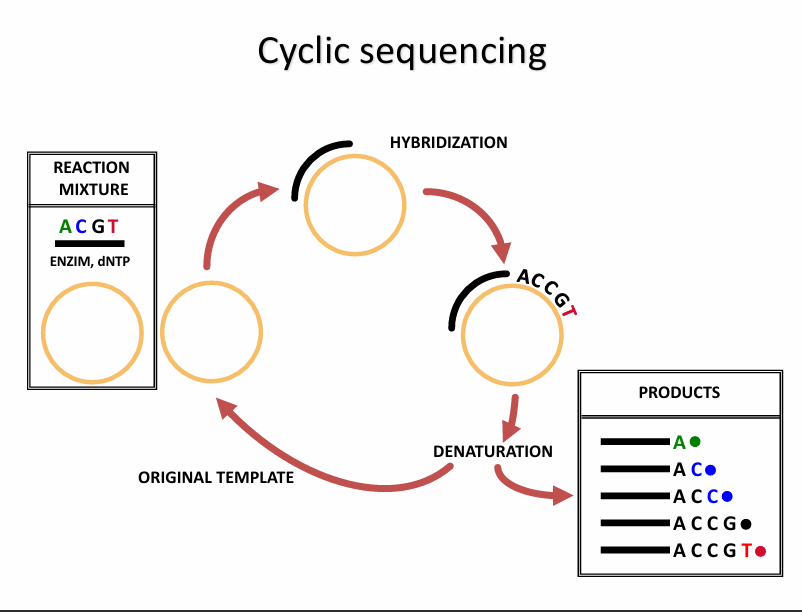
Cyclic Sequencing
This slide shows cycle sequencing, which is a modified Sanger method that works similarly to PCR cycling.
Reaction mixture
The mixture contains:
Template DNA
Primer
DNA polymerase
Normal nucleotides (dNTPs)
Fluorescent ddNTPs (ddA, ddC, ddG, ddT)
Each ddNTP has a different fluorescent colour.
Steps of Cyclic Sequencing (in order)
1. Denaturation
The double-stranded DNA separates into two single strands.
This happens at high temperature (~95 °C).
The template strand becomes available for the primer.
2. Primer Hybridisation (Annealing)
The primer binds to the complementary sequence on the template DNA.
This occurs at a lower temperature (~50–60 °C).
3. Extension (DNA synthesis + chain termination)
DNA polymerase extends the primer.
It adds:
normal nucleotides (dNTPs) → continue DNA synthesis
fluorescent ddNTPs → occasionally inserted and terminate the chain
A
AC
ACC
ACCG
ACCGT
Each fragment ends with a fluorescent base.
4. Repeat the cycle
The steps repeat 25–35 times:
Denaturation → Annealing → Extension → repeatEach cycle produces more terminated fragments of different lengths.
5. Fragment separation and detection
After cycling is finished:
DNA fragments are separated by capillary electrophoresis
A laser detects the fluorescent base at the end of each fragment
The computer reconstructs the DNA sequence
Each fragment ends with Radioactive vs Fluorescent Sequencing
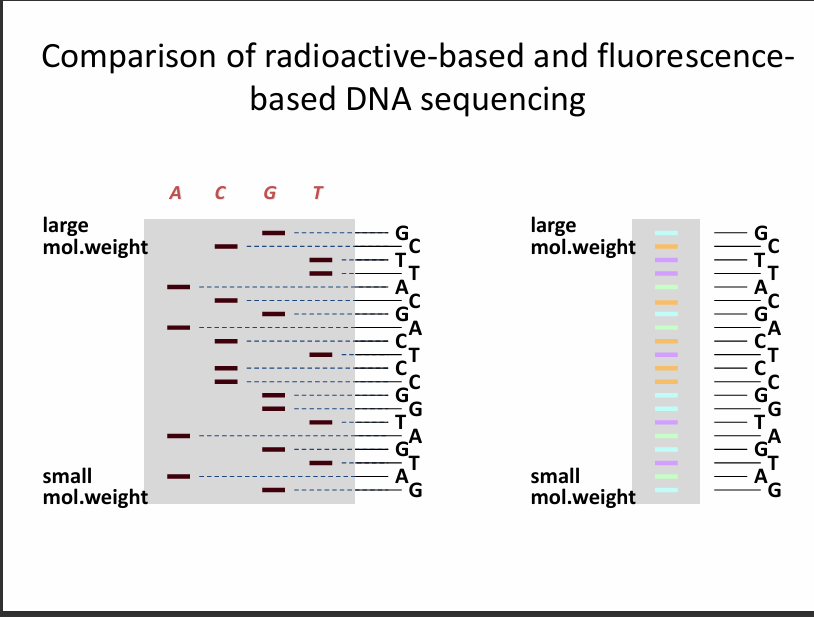
This slide compares the old method with the modern method.
Left side: Radioactive sequencing (old)
Features:
4 separate reactions
4 gel lanes
A | C | G | T
Each lane corresponds to a different ddNTP.
Detection method:
radioactive labeling
autoradiography (X-ray film)
The sequence is read manually from the gel.
Right side: Fluorescent sequencing (modern)
Features:
1 reaction instead of 4 (i.e., has 1 lane instead of 4).
All 4 ddNTPs in one tube
Each base has a different fluorescent colour
Example:
Base | Color |
|---|---|
A | green |
C | blue |
G | yellow |
T | red |
Fragments are separated by capillary electrophoresis, and a laser detects the colour.
The machine automatically converts the colours into the DNA sequence.
REMEMBER: The large weight fragments are on the top while the small weighted ones are located at the bottom
Fluorescent Fragment Detection
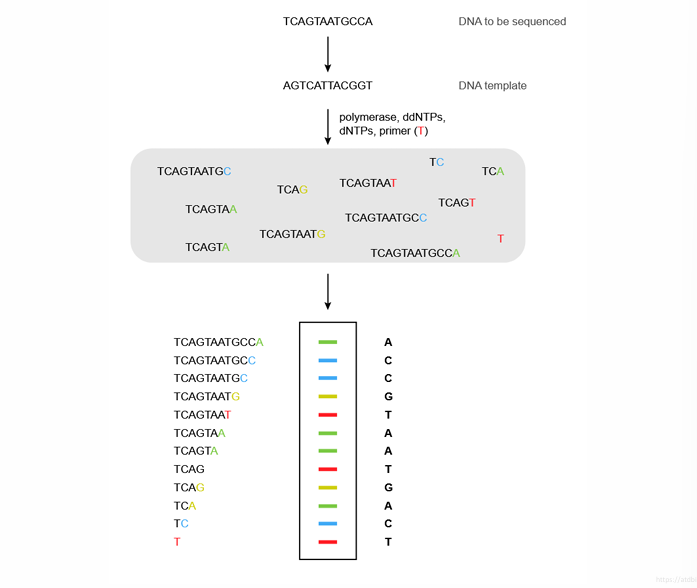
This slide shows how the fragments are read automatically.
Step 1: DNA synthesis
DNA polymerase produces fragments of different lengths.
Example fragments:
T
TC
TCA
TCAG
TCAGT
TCAGTA
Each fragment ends with a fluorescent ddNTP.
Step 2: Electrophoresis
Fragments move through capillary electrophoresis.
Important rule:
Small fragments arrive first
Large fragments arrive later
Step 3: Laser detection
A laser detects the color of the last base.
Example signal sequence:
green
blue
blue
yellow
red
green
green
red
yellow
green
blue
red
The computer converts colors → bases.
Final sequence produced
Example:
ACCGTAATGACT
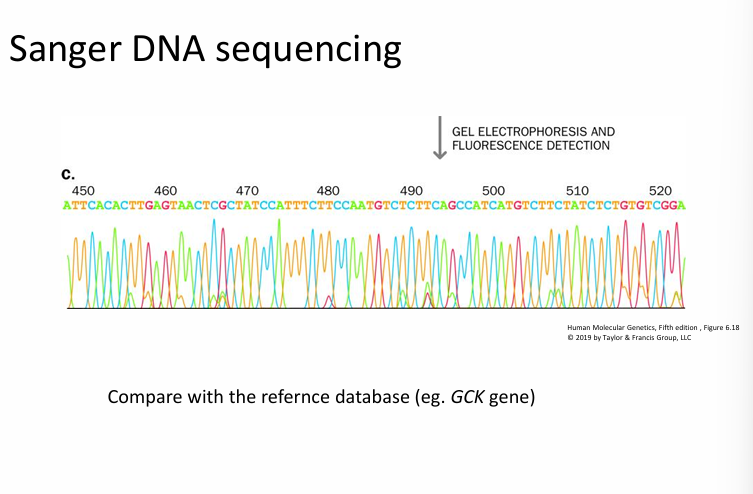
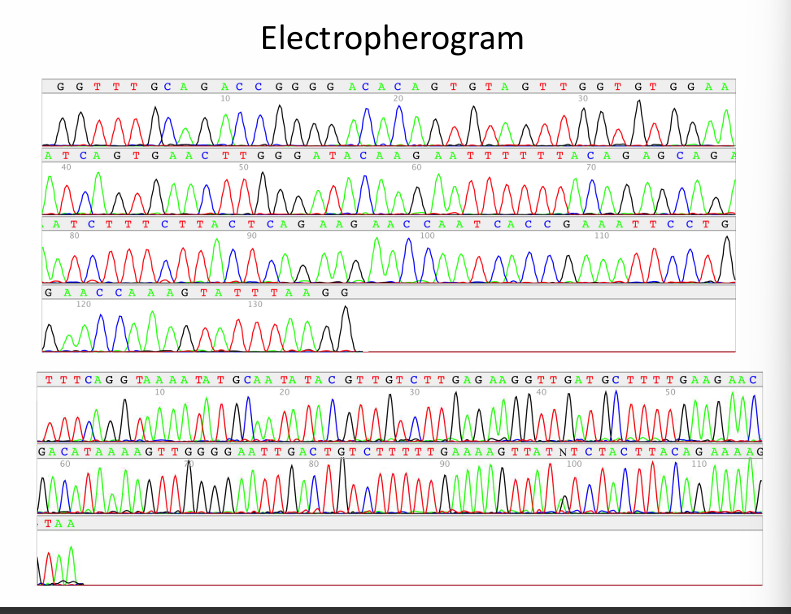
An electropherogram is a graph of colored peaks representing the DNA sequence detected during automated Sanger sequencing.
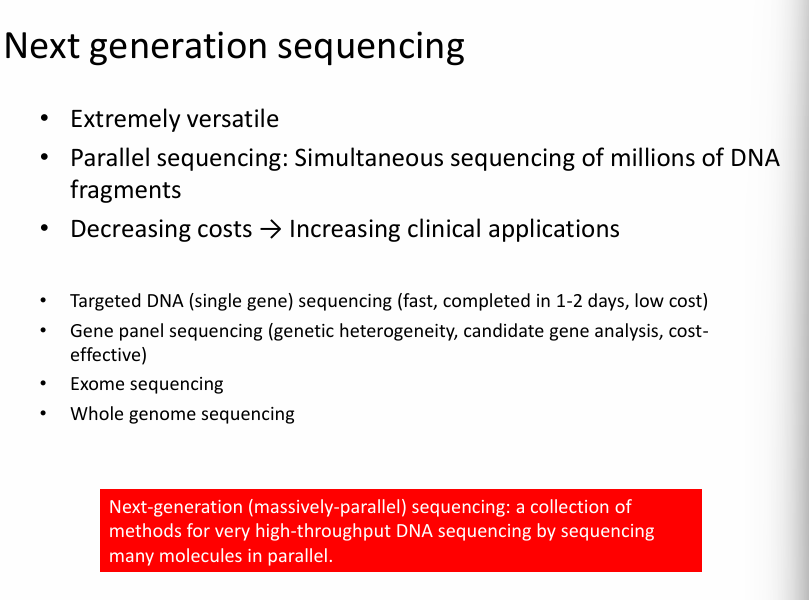

Key idea of Next Generation Sequencing:
Instead of sequencing one DNA fragment at a time (like Sanger sequencing), NGS sequences millions of fragments simultaneously.
What are the characteristics of Next Generation Sequencing:
Extremely versatile
Parallel sequencing: simultaneous sequencing of millions of DNA fragments.
Decrease costs—> increasing clinical applications.
Applications of NGS (from the slide)
Targeted DNA sequencing
Sequencing one specific gene.
Example:
checking for a mutation in a disease gene
Fast and cheap.
Gene panel sequencing
Sequencing a group of related genes.
Example:
cancer gene panels
hereditary disease panels
Used when multiple genes could cause a disease.
Exome sequencing
Sequences all exons in the genome.
Exons = protein-coding parts of genes
Only about 1–2% of the genome, but many disease mutations occur there.
Whole genome sequencing
Sequences the entire genome.
This gives the most complete genetic information.

Steps of Next-Generation Sequencing:
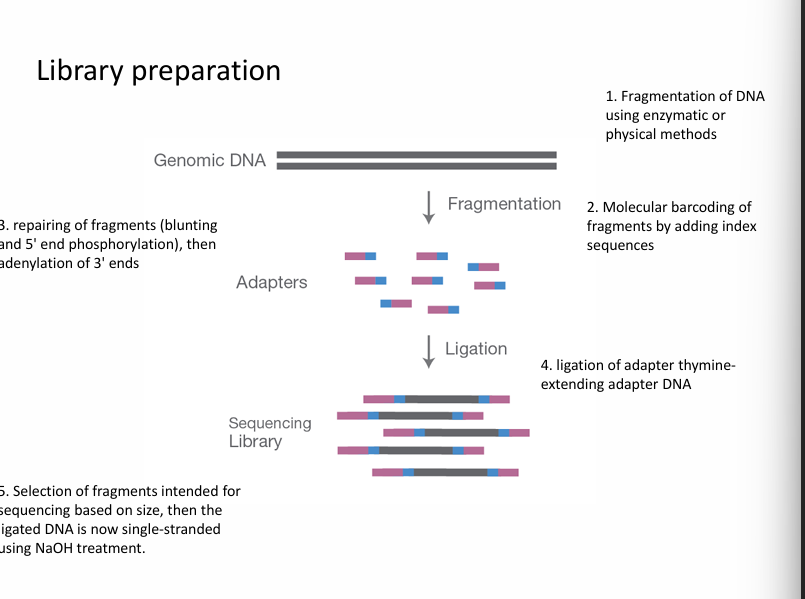
Step 1: Library preparation
The DNA sample is prepared for sequencing.
This involves:
cutting DNA into small fragments
attaching adapters (short DNA sequences)
Adapters allow fragments to bind to the sequencing surface.
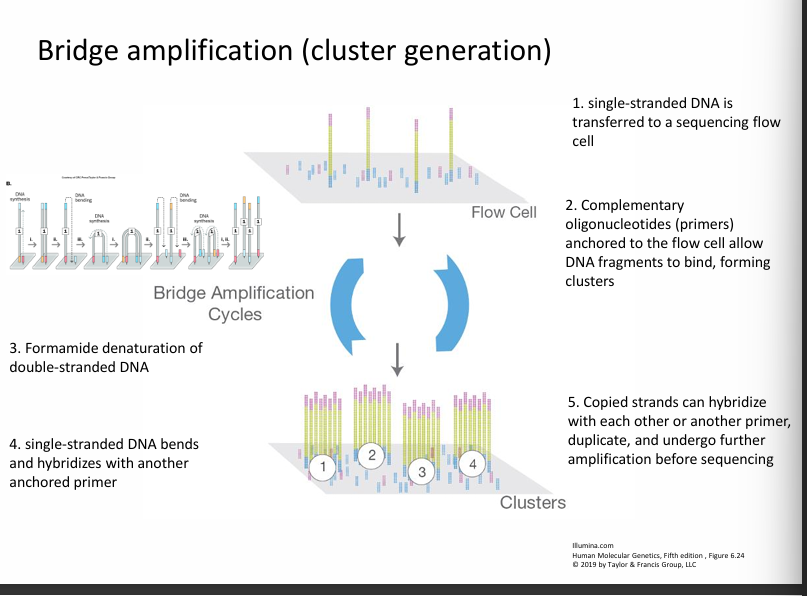
Step 2: Amplification (cluster generation)
Each DNA fragment is copied many times.
This produces clusters of identical DNA fragments.
Why?
Because many copies make the signal easier to detect during sequencing.
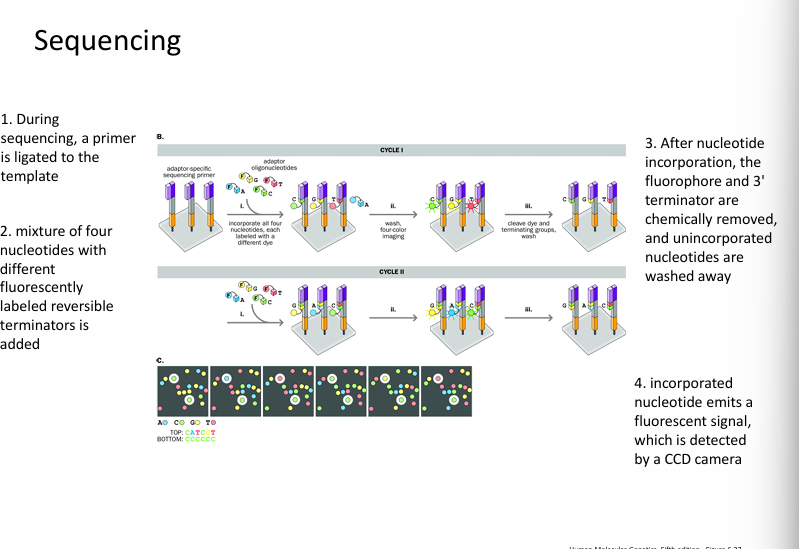
Step 3: Synthesis-based sequencing
DNA polymerase builds a new complementary strand.
Fluorescent nucleotides are added one at a time.
Each nucleotide emits a colour signal when incorporated.
This allows the machine to determine which base was added.
Step 4: Detection
A camera or laser detects the fluorescent signals.
Each colour corresponds to:
Base | Color |
|---|---|
A | one color |
C | another |
G | another |
T | another |
The system records the sequence base by base.
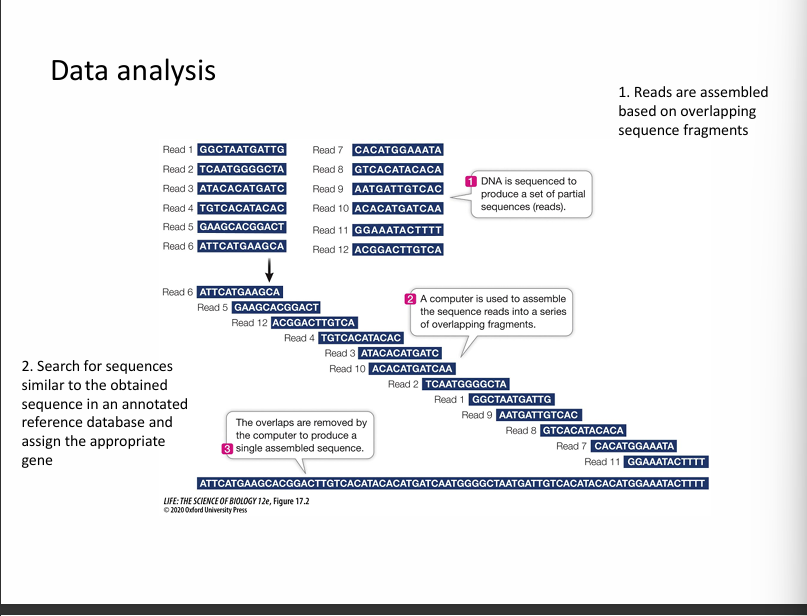
Step 5: Data analysis
Computers analyse the sequencing data.
They:
assemble DNA fragments
Align them to a reference genome
Identify mutations or variations.