A3.1 Diversity of organisms Notes
A3.1.1 variation between organisms as a defining feature of life
No Two Organisms Are Identical In All Their Traits
Definition
VariationVariation refers to differences in traits among individuals within a population, encompassing physical characteristics, behaviors, and genetic sequences.
Variation can manifest as differences in:
Morphological traits: e.g., fur color in cats, height in plants.
Behavioral traits: e.g., bird mating displays, migration patterns.
Biochemical traits: e.g., enzyme variants, antibiotic resistance in bacteria.
Note
Even closely related organisms, such as siblings, exhibit variation because each inherits a unique combination of alleles during reproduction.
Causes of Variation
1. Genetic Causes
Every individual of a species has a unique combination of alleles, except in the case of identical twins or clones.
Genetic variation is introduced by:
Mutation: Permanent changes in DNA sequences create new alleles.
Meiosis: Processes such as crossing over (exchange of DNA between homologous chromosomes) and independent assortment of chromosomes create novel genetic combinations in gametes.
Fertilization: Random fusion of gametes during sexual reproduction ensures that each zygote is genetically unique.
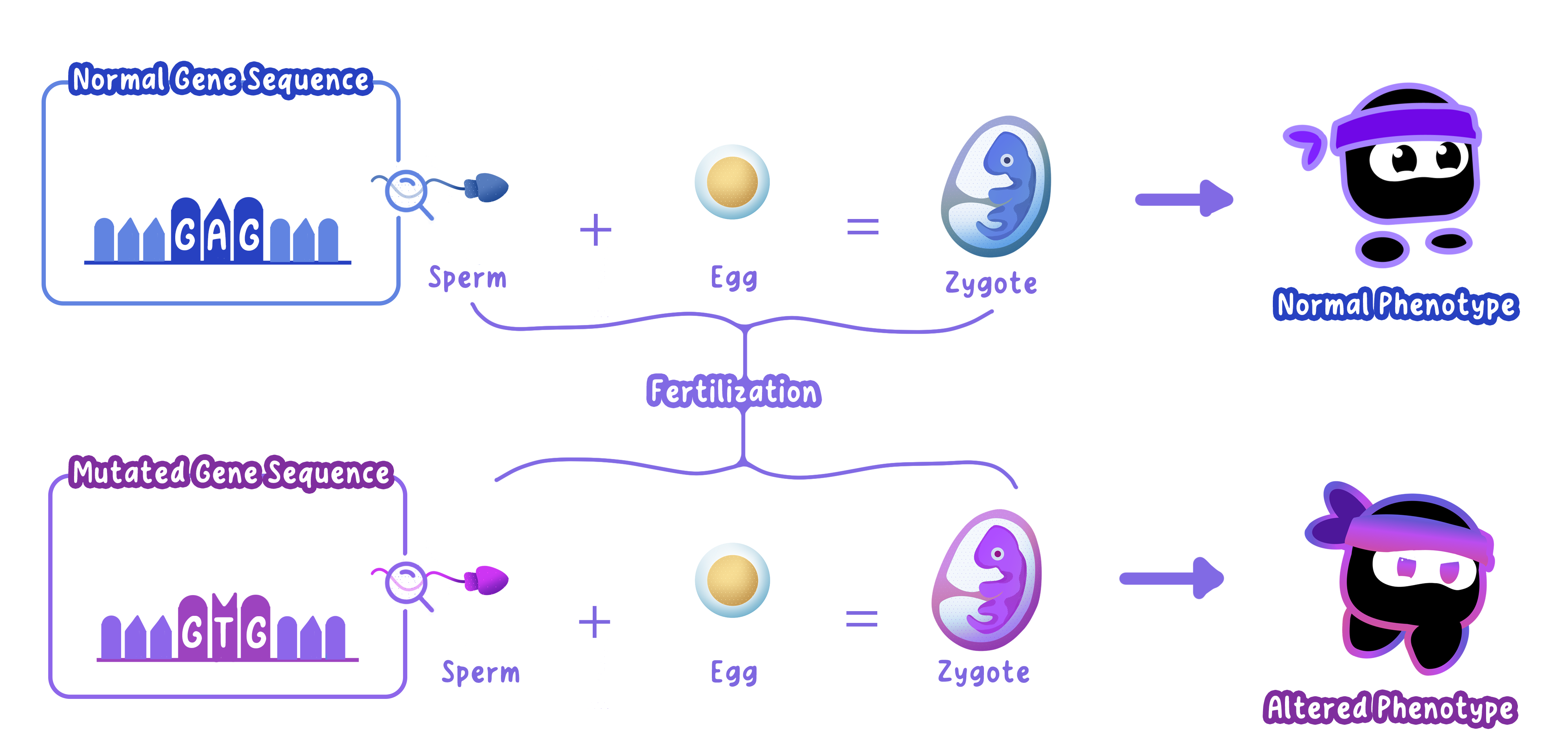
Example
A mutation in a gene coding for melanin can result in different skin pigmentation.
2. Environmental Causes
Even genetically identical organisms show variation due to environmental influence.
Environmental factors can switch genes on or off or affect the degree to which they are expressed.
Nutrient availability, temperature, light, oxygen concentration, and pH all affect phenotype.
Analogy
Genes act like a blueprint, while the environment is the construction site.
A perfect blueprint may result in different buildings if the construction conditions differ.
Example
Monozygotic twins (identical twins) are an excellent example of how variation arises.
These twins start with identical genetic material, but as they grow, environmental differences and random mutations cause them to diverge.
They may differ in height or weight because one had greater access to nutrition during development.
Patterns of Variation
1. Continuous Variation
Traits exhibit a range of values without distinct categories.
Characteristics:
Measured on a continuous scale.
Represented by a bell-shaped curve (e.g., height distribution).
Often influenced by polygenic inheritance and environmental factors.
Example
Human height, leaf length in plants, fur color in animals.
Tip
Traits showing continuous variation are controlled by multiple genes working together.
2. Discontinuous Variation
Traits fall into distinct categories with no intermediates.
Characteristics:
Categorical traits (e.g., yes/no).
Represented by bar graphs.
Typically controlled by a single gene or a few genes, with minimal environmental influence.
Example
Blood type in humans (A, B, AB, O) or flower petal presence.
Tip
Continuous traits show a range (e.g., height), while discontinuous traits fall into specific categories (e.g., blood type).
Variation Between Species vs. Within Species
Between species: Structural differences allow classification.
Within species: Members of a species share a common genetic basis but differ due to allelic combinations.
Example
Between species: presence of chloroplasts in plants vs. absence in animals.
Within species: humans all have the same genes for blood type, but different alleles (A, B, O) produce different phenotypes.
Tok
How does our understanding of variation challenge the idea of fixed categories in nature? For example, consider the ethical implications of classifying human populations based on genetic differences. Are these classifications scientific, or do they risk reinforcing stereotypes?
Self Review
Can you explain the difference between continuous and discontinuous variation?
Why is variation considered a defining feature of life?
List three genetic and three environmental causes of variation in organisms.
Explain why monozygotic twins are not completely identical, despite sharing the same genome.
A3.1.2 species as groups of organisms with shared traits
The Morphological Species Concept (Historical)
The morphological species concept defined a species as a group of organisms that share similar morphological characteristics (form and structure), distinguishing them from other groups.
In other words, this approach classifies organisms by what they look like, rather than by how they reproduce or how related they are genetically.
Analogy
Like grouping books by their cover design rather than their content.
Simple, but often misleading.
Why Morphology Was Used
Ease of observation: Early scientists lacked microscopes or genetic tools, physical features were all they could study.
Universality: Every organism has morphological traits that can be described and compared.
Comparative simplicity: Allowed systematic grouping based on visible patterns (shape, structure, symmetry).
Example
Linnaeus grouped whales with fish because both had fins, a conclusion we now know to be evolutionarily inaccurate.
Strengths of the Morphological Species Concept
Simple and visual: Classification possible using only external traits.
Useful for fossils: The only information available in many extinct species is morphology.
Field identification: Practical for ecologists and naturalists working without genetic tools.
Hint
The morphological species concept is historically significant and still has limited modern use.
For example, when only physical remains exist.
Limitations and Why It Was Replaced
Cryptic species: Genetically distinct species can appear identical.
Phenotypic variation: Males, females, or developmental stages may look very different.
Subjectivity: Different scientists may classify the same organism differently.
No genetic basis: It cannot explain relatedness or evolutionary descent.
Note
Because morphology alone is unreliable, this concept has been replaced by biological and genetic definitions that consider reproductive isolation and shared ancestry (more in A3.1.13 next).
Self Review
What is the main idea behind the morphological species concept?
Why was this concept useful in the 18th century?
Who developed the hierarchical system of classification still used today?
Why is morphology alone not sufficient to define a species?
What are cryptic species, and how do they challenge this concept?
Why is this no longer the definition of species used in biology?
A3.1.3 binomial system for naming organisms
Two Parts To The Binomial System
Definition
Binomial nameA binomial name is a two-part Latinized name that uniquely identifies a species. The first word represents the genus, and the second word represents the specific species within that genus.
The binomial system gives every species a unique two-part name, ensuring consistent identification.
It consists of:
Genus name: The first part, which groups closely related species.
Species name: The second part, which specifies the exact species.
Example
Homo sapiens:
Homo: The genus grouping humans with extinct relatives like Homo neanderthalensis.
sapiens: The species name identifying modern humans.
Formatting Rules
Italicize when printed, or underline when handwritten.
After first mention, the genus can be abbreviated (e.g., E. coli).
Names are Latin or latinized, ensuring international understanding.
Exam_technique
Genus = capitalized; species = lowercase.
Both italicized.
Functional Value
Consistency: The same species has the same name worldwide.
Clarity: Prevents confusion caused by common names.
Evolutionary meaning: Species within a genus share structural and genetic traits.
Self Review
What does the first part of a binomial name represent?
How should scientific names be formatted in print?
Why is Latin used in binomial nomenclature?
Give one reason the system prevents confusion.
A3.1.4 biological species concept
The Biological Species Concept (Modern Definition)
Definition
Biological Species Concept (BSC)The biological species concept (BSC) defines a species as a group of organisms capable of interbreeding and producing fertile offspring, emphasizing reproductive isolation.
This is the primary definition used in modern biology, unlike the morphological concept.
This view focuses on reproductive compatibility and shared gene pools, not just appearance.
Example
Lions (Panthera leo) and tigers (Panthera tigris) are separate species because they do not interbreed in the wild.
Hybrids like ligers (male lion X female tiger) are typically sterile, preventing gene flow.
Key Principles of the BSC
Interbreeding ability: Members of the same species can mate and produce fertile offspring.
Shared gene pool: All individuals exchange genes within the same population, maintaining genetic continuity and diversity.
Reproductive isolation: Different species are separated by barriers, geographic, behavioral, physiological, or genetic, preventing gene flow and ensuring independent evolution.
Tip
Reproductive isolation is the mechanism that maintains distinct species boundaries.
When the BSC Cannot Be Applied
Other classification methods are used when reproductive isolation cannot be tested:
Alternative Concept | Basis for Classification |
|---|---|
Morphological | Observable physical traits |
Genetic / Molecular | DNA or protein similarity |
Ecological | Distinct ecological roles or niches |
Evolutionary lineage | Ancestral relationships in fossils |
Biochemical | Differences in metabolism or chemical pathways |
Hint
No single definition fits all organisms.
Species concepts are tools, chosen based on context and available evidence.
Self Review
What is the main definition of a species under the biological species concept?
What role does reproductive isolation play in maintaining species boundaries?
Why is this concept more consistent with evolutionary theory than the morphological one?
Name two types of barriers that prevent interbreeding.
Why can't the BSC be applied to bacteria or fossils?
A3.1.5 difficulties distinguishing between populations and species
From Populations To Species
When populations of the same species become geographically or reproductively isolated, they start to accumulate:
Genetic differences through mutation and genetic drift
Phenotypic differences through natural selection
Over time, these differences may result in reproductive isolation, the key step in speciation.
Note
More on speciation in A4.1.6.
The Gray Area in Classification
In many cases, it is difficult to decide when diverging populations have become distinct species.
Populations may not interbreed simply because they live apart, not because they can't.
If separated populations remain genetically and phenotypically similar, they are still considered the same species.
Over many generations, isolation allows mutations, selection, and drift to cause divergence in traits such as size, color, or behavior.
Eventually, these changes may lead to complete reproductive isolation, producing two distinct species.
Tip
Speciation is a process, not a single event.
The point at which populations become separate species is therefore often subjective, and may be an arbitrary decision.
Self Review
How does a population differ from a species?
What mechanisms cause populations to diverge over time?
Why is it difficult to determine when two populations become distinct species?
A3.1.6 diversity in chromosome numbers of plant and animal species
Diversity in Chromosome Numbers Across Species
A species is partly defined by its characteristic chromosome number, which remains stable in normal individuals.
Chromosome number is conserved across all diploid cells within a species, ensuring accurate inheritance during reproduction.
Occasional mutations or chromosomal rearrangements may alter this in individuals, but the species norm remains stable.
Across species, chromosome number varies dramatically--from just two chromosomesin some nematodes to hundreds in ferns.
Chromosome Numbers Are Shared Traits Within a Species
Definition
ChromosomesChromosomes are the carriers of genetic information in all living organisms. They consist of two essential components: deoxyribonucleic acid (DNA) and proteins.
One of the defining characteristics of a species is that its members typically share the same number of chromosomes.
This uniformity is essential for successful reproduction.
Diploid Cells and Even Chromosome Numbers
Most plants and animals are diploid, meaning their body cells contain two sets of chromosomes, one from each parent.
Humans: Somatic cells have 46 chromosomes (23 pairs).
Each pair consists of homologous chromosomes that carry the same sequence of genes.
Haploid Gametes: During fertilization, sperm and egg cells (haploid) combine to restore the diploid number.
Hint
Remember, the diploid chromosome number is always even because it represents two complete sets of chromosomes, one from each parent.
Chromosome Numbers Are Usually Even
Diploid organisms inherit two sets of chromosomes: one paternal, one maternal.
During gamete formation, chromosomes must pair during meiosis; this is only possible if the diploid number is even.
After fertilization, the zygote is restored to diploid status.
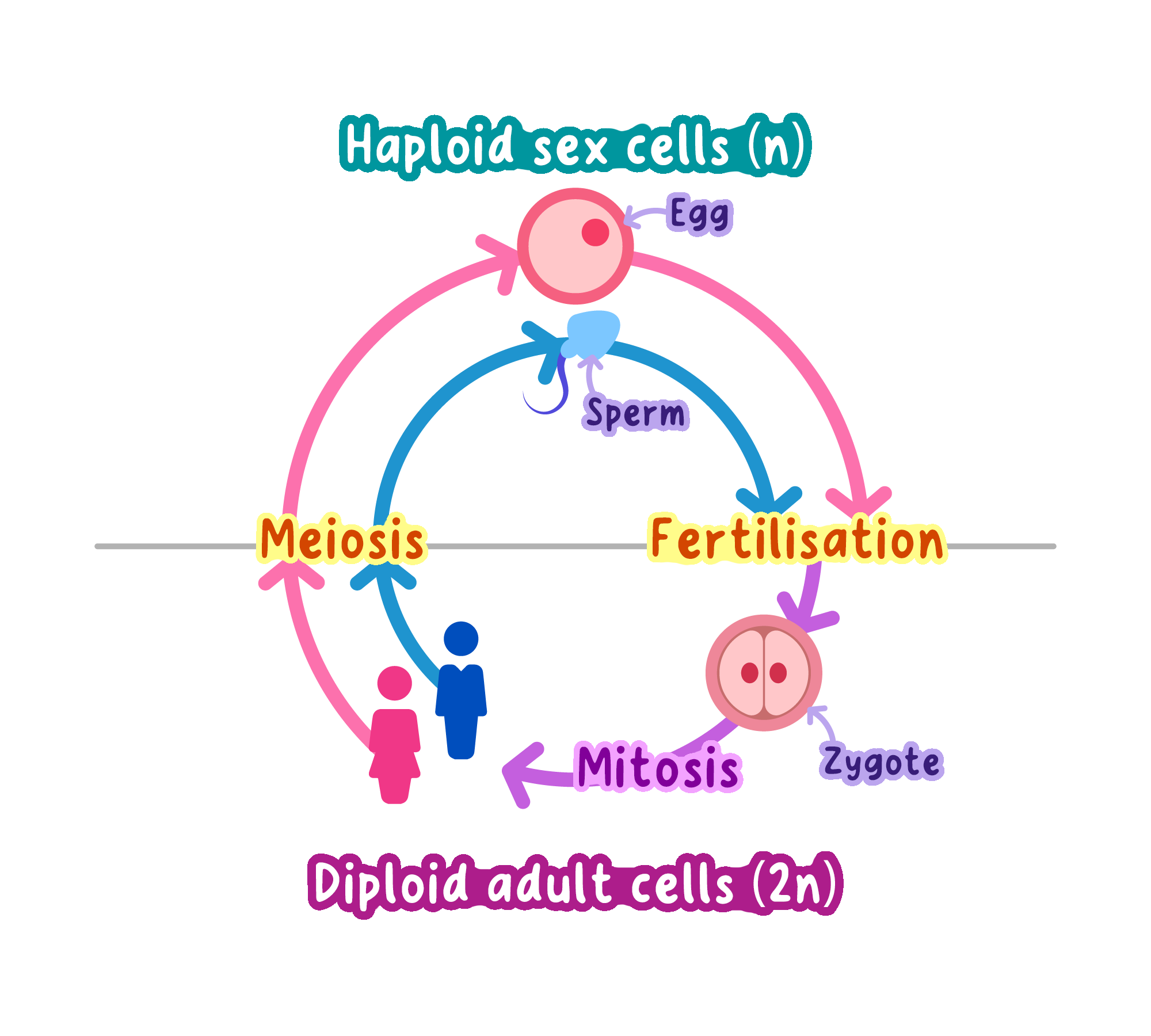
Exam_technique
In IB exams, always state whether you are referring to the haploid (n) or diploid (2n) number.
Saying "humans have 23 chromosomes" is incorrect.
It should be "23 pairs in diploid cells" or "23 chromosomes in haploid gametes."
How Chromosome Numbers Change Over Time
Changes in chromosome numbers are rare events, but when they do occur, they can be caused by:
1. Chromosome Fusion
Two chromosomes join together end-to-end.
This reduces the chromosome number.
Example
Human chromosome 2 is the result of a fusion of two ancestral ape chromosomes.
This explains why humans have 46 chromosomes while chimpanzees, gorillas, and orangutans have 48.
2. Chromosome Fission
A single chromosome splits into two smaller chromosomes.
This increases the chromosome number without increasing the DNA content.
3. Polyploidy
Entire sets of chromosomes duplicate.
Common in plants (e.g., wheat is hexaploid, 6n).
Polyploidy can lead to instant speciation because polyploid individuals cannot breed with diploid members of the same species.
Example
Wheat is hexaploid (six sets of chromosomes).

Example
A horse has 64 chromosomes, while a donkey has 62.
Their offspring, a mule, inherits 63 chromosomes.
This odd number prevents proper chromosome pairing during meiosis, leading to infertility.
Diversity in Chromosome Numbers Across Species
Chromosome numbers vary dramatically among species, ranging from as few as 2 to over 100.
Humans share close genetic similarity with chimpanzees, yet differ in chromosome number (46 vs 48) due to a fusion event in human ancestry.
Example
Humans vs. Chimpanzees
Humans: 46 chromosomes.
Chimpanzees: 48 chromosomes.
Cause of Difference: Research suggests that human chromosome 2 resulted from the fusion of two ancestral chromosomes after humans and chimpanzees diverged.
Polyploidy in Plants
Unlike animals, plants frequently exhibit polyploidy (more than two sets of chromosomes).
Polyploidy increases genetic diversity and often leads to new species.
Note
Polyploidy is rare in animals but common in plants, as plants can reproduce asexually, bypassing the challenges of uneven chromosome numbers during meiosis.
Chromosome Size vs. Number
The same total DNA content can be arranged in a few large chromosomes or many small chromosomes.
Laboratory experiments in yeast show that reducing chromosomes from 16 to just 2-4 still allowed survival, proving that chromosome number itself is not the limiting factor.
Instead, it is the presence of all essential genes.
Example
Some plants have very high chromosome numbers but are not genetically more complex than humans.
Tok
How does the study of chromosome numbers challenge traditional definitions of species? Consider hybrids like ligers or polyploid plants that blur the boundaries.
Self Review
Define chromosome number, diploid, and haploid, using humans as an example.
Why are diploid chromosome numbers nearly always even?
Describe two mechanisms that can increase and one mechanism that can decrease chromosome numbers.
State the diploid chromosome numbers of humans and chimpanzees, and explain the difference.
A3.1.7 karyotyping and karyograms
Three Main Criteria Are used To Classify Chromosomes
Banding Patterns
When stained with dyes like Giemsa, chromosomes show light and dark bands unique to each chromosome.
These act like a barcode, helping identify specific chromosomes and detect deletions, duplications, or translocations.
Length
Chromosomes are arranged in a karyogram from longest to shortest.
In humans, chromosome 1 is the longest and chromosome 21 is among the shortest.
Centromere Position: The centromere divides the chromosome into two arms (p = short, q = long).
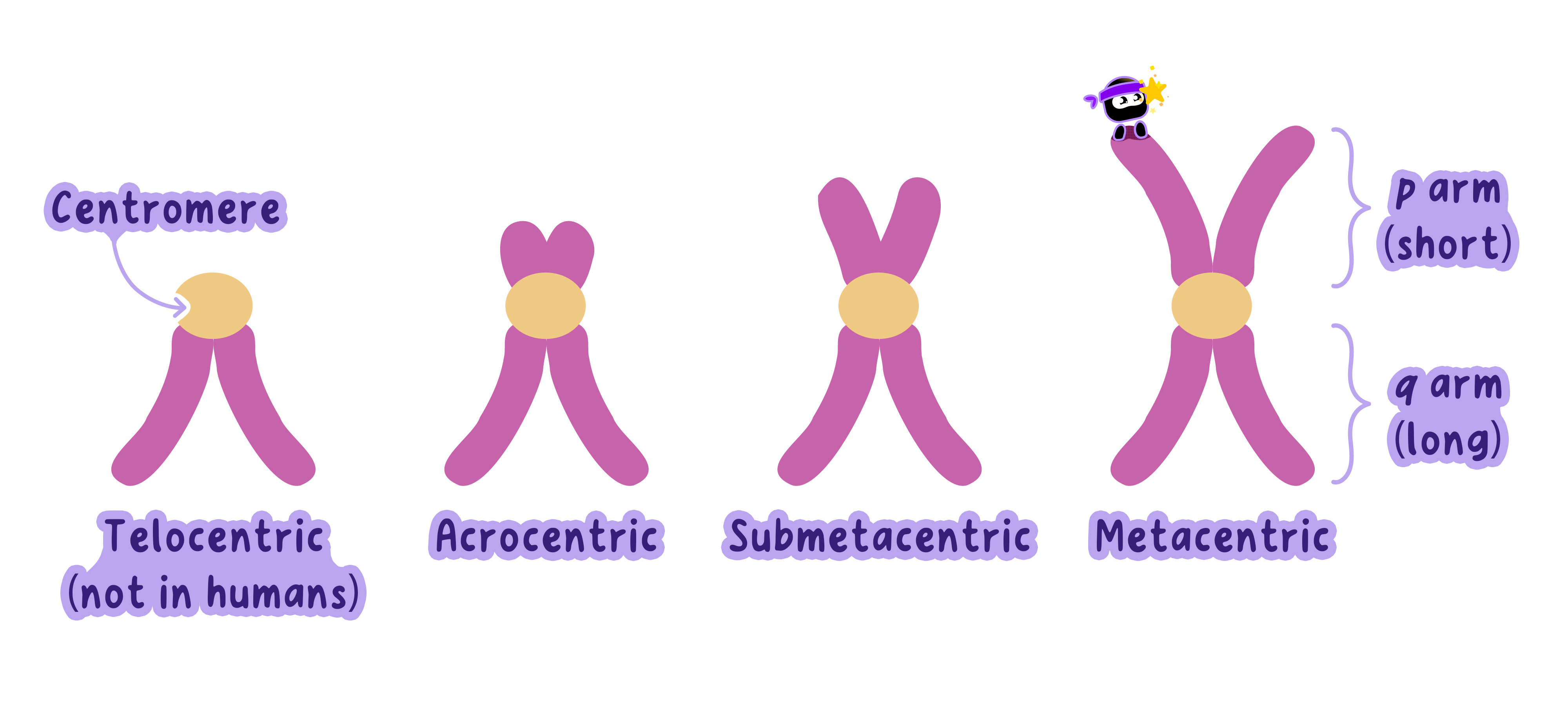
Karyotypes and Karyograms
A karyotype describes the complete chromosome set of an organism (number, size, and structure).
A karyogram is the visual arrangement of those chromosomes in homologous pairs, typically ordered by decreasing size.
Humans have 46 chromosomes (23 pairs):
22 pairs of autosomes
1 pair of sex chromosomes (XX or XY)
Tip
Remember that sex chromosomes (pair 23) often break the size order (e.g., X is large, Y is small).
Evidence for the Fusion of Human Chromosome 2
Humans have 46 chromosomes, while chimpanzees, gorillas, and bonobos each have 48.
This led scientists to hypothesize that human chromosome 2 formed through the fusion of two ancestral ape chromosomes (chimpanzee chromosomes 12 and 13).
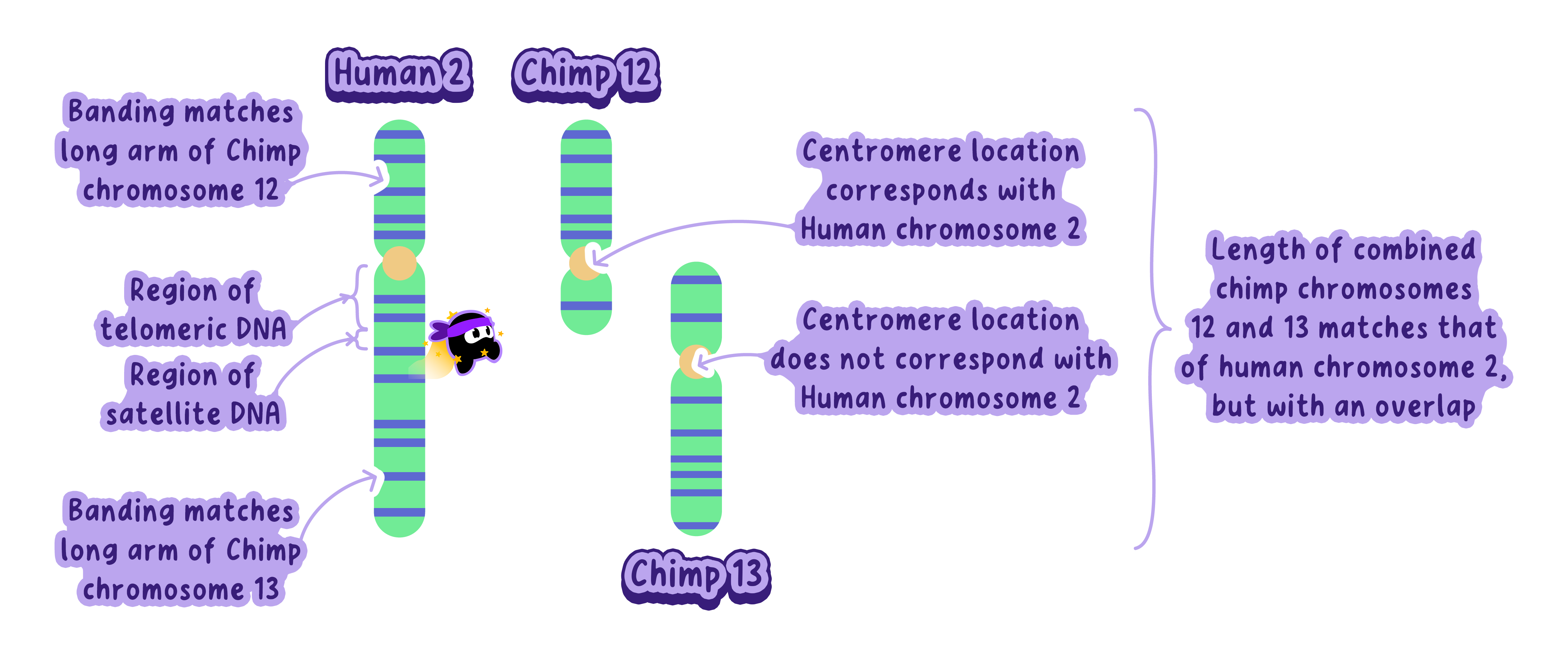
Example
The active centromere in human chromosome 2 corresponds to the centromere of chimpanzee chromosome 13,
While remnants of the second centromere align with chimpanzee chromosome 12.
Nature of Science (NOS): Testable Hypotheses
The origin of human chromosome 2 is a testable hypothesis because it can be supported or refuted by observable evidence, such as banding, centromere structure, and DNA sequence data.
Testable hypotheses must:
Make specific, evidence-based predictions.
Allow for acceptance or rejection through data.
Avoid vague or unverifiable claims.
Example
Testable: "Human chromosome 2 formed by fusion of two ancestral ape chromosomes."
Non-testable: "The ancestor of humans and chimpanzees preferred singing." (no evidence possible)
Self Review
What three criteria are used to classify chromosomes?
Why is the chromosome 2 fusion hypothesis considered testable?
Give one example of a non-testable statement and explain why it fails the criteria.
How does the fusion of chromosome 2 support the idea of shared ancestry among primates?
A3.1.8 unity and diversity of genomes within species
The Genome Allows for Genetic Diversity within Species
Definition
GenomeThe genome is the complete set of genetic instructions that determines the structure, function, and traits of an organism.
The genome is the instruction manual for building and maintaining an organism, written in the four DNA bases A, T, C, and G.
It includes:
Coding DNA: Sequences that encode proteins.
Non-coding DNA: Sequences that regulate gene expression or have other functions.
Analogy
Think of the genome as a comprehensive instruction manual where:
Chapters = Chromosomes, long strands of DNA wrapped around proteins.
Sentences = Genes, specific sequences coding for proteins.
Spaces = Non-coding regions, essential for timing and regulation.
Unity Within a Species
All members of a species share the same set of genes arranged in the same order on their chromosomes.
This shared structure ensures that organisms can reproduce successfully, allowing homologous chromosomes to pair, align and cross over correctly during meiosis.
Example
All humans have the hemoglobin gene on chromosome 11, regardless of which allele version they inherit.
Tip
Understand that if the chromosomes don't pair correctly, sexual reproduction would fail.
Genetic Diversity within a Species
While genomes are largely uniform within a species, small variations exist, creating individual uniqueness.
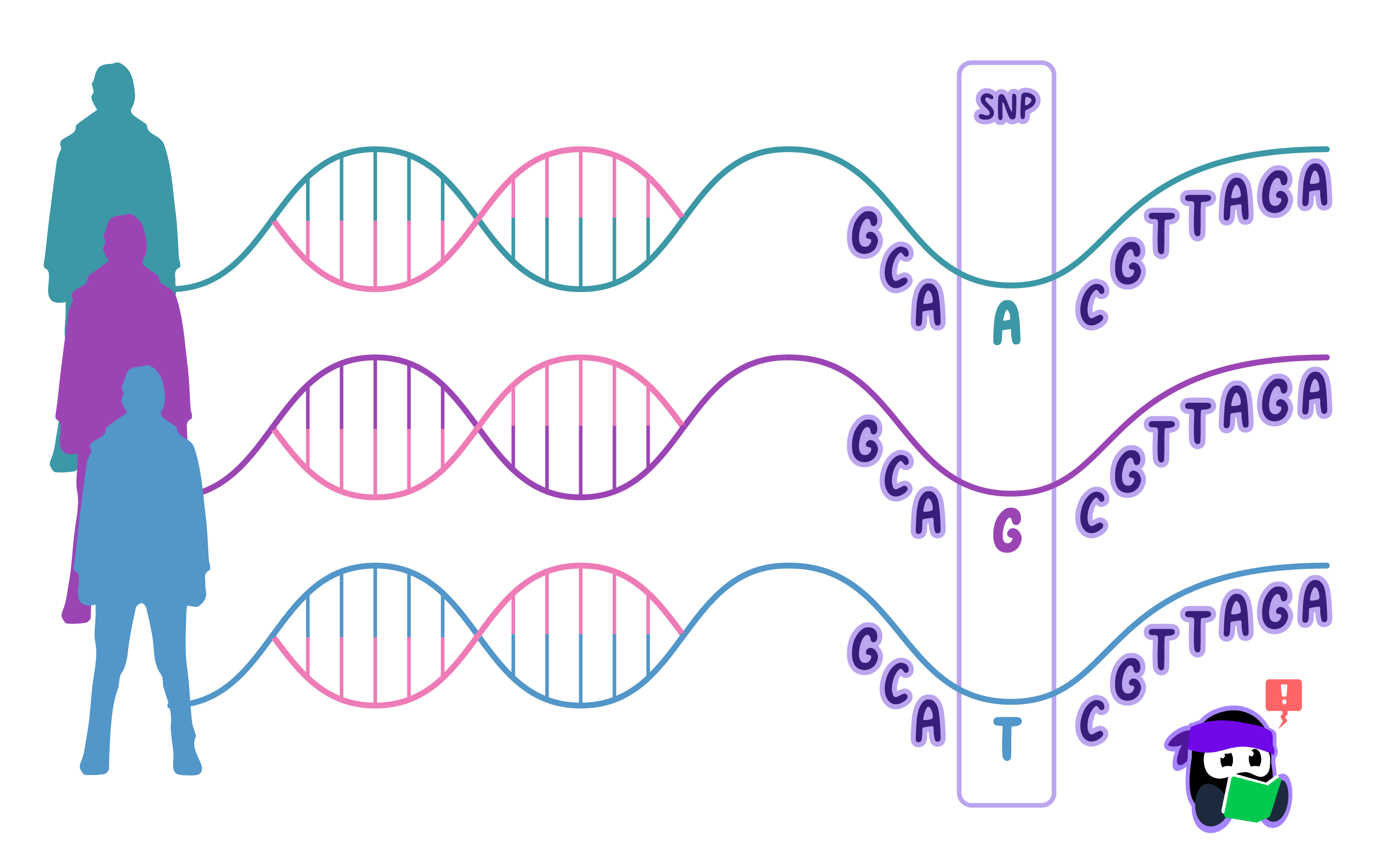

Single-Nucleotide Polymorphisms (SNPs)
Definition
Single-Nucleotide Polymorphism (SNP)SNPs are single base-pair changes in the DNA sequence.
The most common source of variation is the single-nucleotide polymorphism (SNP).
SNPs occur roughly once every 650,000 base pairs in humans, resulting in 4,000-5,000 SNPs per individual.
Example
Eye color
A single SNP might alter one base in this gene, resulting in blue eyes instead of brown.
This demonstrates how small genetic changes can lead to visible differences within a species.
Why Are SNPs Important?
Trait Variation: SNPs contribute to differences in height, fur color, or disease resistance.
Evolutionary Insights: They act as markers to trace inheritance patterns and evolutionary relationships.
Individual Uniqueness: SNPs drive the subtle variations that differentiate individuals within a population.
Tip
Keep in mind that most SNPs are neutral.
They don't affect an organism's traits or survival.
However, some SNPs can have significant impacts, such as increasing the risk of certain diseases or providing resistance to infections.
Unity and Diversity Across Species
Within species: Humans share approximately 99.9% of their DNA with each other. This explains why all humans belong to the same species and can reproduce together.
Between species: Humans share approximately 99% of their DNA with chimpanzees, reflecting evolutionary relatedness.
SNPs and other variations account for the 0.1% difference between individual humans, explaining why people look and behave differently despite being the same species.
Example
Lactose tolerance in adults is caused by SNPs in regulatory regions of the LCT gene.
Note
Even identical twins, who begin life with the same genome, accumulate genetic differences over time due to mutations and environmental influences.
This highlights the dynamic nature of genetic diversity.
Self Review
Why do all members of a species share the same gene order on chromosomes?
How does shared genome structure support successful meiosis?
What is a single-nucleotide polymorphism (SNP)?
What percentage of DNA do humans share with each other?
How do SNPs contribute to trait variation?
Why is genetic diversity important for a species?
A3.1.9 diversity of eukaryote genomes
Variation in Eukaryotic Genome Size and Sequence
Definition
GenomeThe genome is the complete set of genetic instructions that determines the structure, function, and traits of an organism.
All eukaryotes share the universal genetic code, but their genomes vary greatly in total size and base sequence composition.
Variation between species is far greater than variation within a species.
Variation in Eukaryotic Genomes
The genome of an organism is its complete set of DNA, including all coding (genes) and non-coding sequences.
In eukaryotes, this includes:
Nuclear DNA
Mitochondrial DNA
Chloroplast DNA (in plants)
Tip
Although the structure and code of DNA are universal, the total amount and sequence of DNA vary widely between organisms.
Why Genomes Differ
Differences in eukaryotic genomes arise from two main factors:
Variation in total DNA content (genome size)
Larger genomes contain more non-coding DNA, such as introns and repetitive sequences.
These regions contribute to genome size but not necessarily to gene number.
Variation in base sequence composition
Differences in nucleotide order accumulate through mutation, gene duplication, and deletion over evolutionary time.
These sequence differences explain genetic diversity both within and betweenspecies.
Variation in Genome Size
Genome size is measured in base pairs (bp).
Among eukaryotes, genome size varies by hundreds of thousands of times, mostly due to non-coding regions rather than differences in gene count.
Example
Human genome ≈ 3.2 X 10⁹ bp
Some plants (e.g., lilies) > 1 X 10¹¹ bp
Yet both have roughly similar numbers of functional genes.
Note
Genome size does not correlate with organism complexity or gene count.
Variation in Base Sequence
Within a species
Most individuals share nearly identical base sequences.
Minor differences, such as single nucleotide polymorphisms (SNPs), create genetic variation between individuals.
Between species
Base sequence differences are far greater, reflecting long-term evolutionary divergence and independent ancestry.
Tip
Variation within species underpins natural selection, while variation between species explains evolutionary relationships.
Self Review
What is included in a eukaryotic genome?
What are the two main causes of genome variation?
Why does genome size vary so widely among eukaryotes?
Why doesn't genome size correlate with the number of genes?
How does variation in base sequence differ within and between species?
A3.1.10 comparison of genome sizes
Genome Size and Complexity
Larger genomes do not necessarily indicate greater biological complexity.
Key contributors to variation include:
Non-coding DNA: repetitive sequences, introns, and transposable elements.
Polyploidy: whole-genome duplication, especially common in plants.
Gene duplication and intron variation: expansion of non-essential DNA regions.
Example
Humans (~3.2 X 10⁹ bp) have a smaller genome than some amphibians or plants (>10¹¹ bp) despite higher organismal complexity.
Summary Table (Typical Genome Sizes)
Group | Approximate Genome Size | Notable Feature |
|---|---|---|
Bacteria | 1-10 Mbp | Compact, mostly coding DNA |
Fungi | 10-100 Mbp | Moderate non-coding regions |
Plants | 100 Mbp - 100 Gbp | Polyploidy, high non-coding DNA |
Animals | 100 Mbp - 10 Gbp | Large variation; no link to complexity |
Self Review
Why might genome size increase without added complexity?
A3.1.11 current and potential future uses of whole genome sequencing
Advances in Speed and Cost
Whole genome sequencing (WGS) is now rapid and affordable, transforming biological and medical research.
It currently underpins studies of evolutionary relationships and genomic variation.
Future uses in personalized medicine, environmental monitoring, and agriculture are expanding rapidly.
Year | Approx. Cost per Genome | Time Required |
|---|---|---|
2003 | >$3 billion | 10+ years |
2025 | < $1,000 | < 24 hours |
Definition
Whole Genome Sequencing (WGS)Whole Genome Sequencing (WGS) is a DNA sequencing technique that determines the order of all or most of an organism's DNA bases.
Current Applications of WGS
Comparative genomics: Aligning DNA sequences identifies conserved genes (essential functions) and divergent genes (adaptive traits).
Molecular clocks: Mutation rates estimate divergence times between species.
Horizontal gene transfer (HGT): Reveals genes transferred between unrelated species, such as antibiotic resistance genes in bacteria.
Tip
WGS allows scientists to trace shared ancestry and evolutionary innovation with high precision.
Potential Future Applications
Personalized Medicine
Disease risk prediction: Identifies variants linked to inherited disorders or predisposition to disease.
Pharmacogenomics: Matches drug choice and dosage to individual genetic profiles.
Gene therapy: Guides genome-editing treatments (e.g., CRISPR) to correct faulty genes.
Environmental and Agricultural Uses
eDNA monitoring: Detects organisms from trace DNA in water, soil, or air--useful for tracking biodiversity and invasive species.
Precision agriculture: Identifies genes for drought tolerance, pest resistance, and higher yields.
Preventive Genomic Health
Routine genome sequencing at birth could identify risk alleles early, supporting preventive healthcare and lifelong monitoring.
Self Review
What does whole genome sequencing determine?
How has the cost and speed of WGS changed since 2003?
What are three current uses of WGS in evolutionary biology?
How does WGS contribute to personalized medicine?
What ethical challenges are associated with genomic data use?
A3.1.12 difficulties applying the biological species concept (hl)
Why the Biological Species Concept Faces Challenges
Definition
Biological Species Concept (BSC)The biological species concept (BSC) defines a species as a group of organisms capable of interbreeding and producing fertile offspring, emphasizing reproductive isolation.
BSC works well for sexually reproducing organisms.
But, it faces challenges when applied to organisms that reproduce asexually or exchangegenetic material across species boundaries.
Asexual Reproduction Challenges The BSC
Many organisms reproduce entirely asexually, meaning there's no interbreeding so the BSC cannot be applied.
Offspring are genetically identical clones of the parent, produced by mitosis rather than fertilization.
Without recombination of genetic material, species boundaries are difficult to definebecause all individuals are essentially genetic copies.
Definition
Clone A group of genetically identical organisms or cells derived from a single parent.
Example
Plants:
Dandelions (Taraxacum officinale) reproduce asexually by forming seeds without fertilization.
Many other plants reproduce vegetatively (e.g., runners, tubers).
Animals:
Some reproduce by parthenogenesis, where females produce offspring from unfertilized eggs.
Seen in aphids, some reptiles (e.g., Komodo dragons), and even sharks.
Parthenogenesis: A form of asexual reproduction where an egg develops into an individual without fertilization.
Exam_technique
If asked in exams, highlight that parthenogenesis challenges the BSC because no interbreeding takes place, yet viable lineages persist.
Horizontal Gene Transfer in Bacteria
Definition
Horizontal Gene TransferHorizontal gene transfer involves the direct exchange of genetic material between unrelated organisms, bypassing traditional parent-offspring inheritance. It is especially common in prokaryotes (bacteria and archaea).
Since bacteria reproduce by binary fission (strictly asexual), the BSC cannot be applied.
This is because bacteria frequently exchange genes across strains and even speciesthrough HGT.
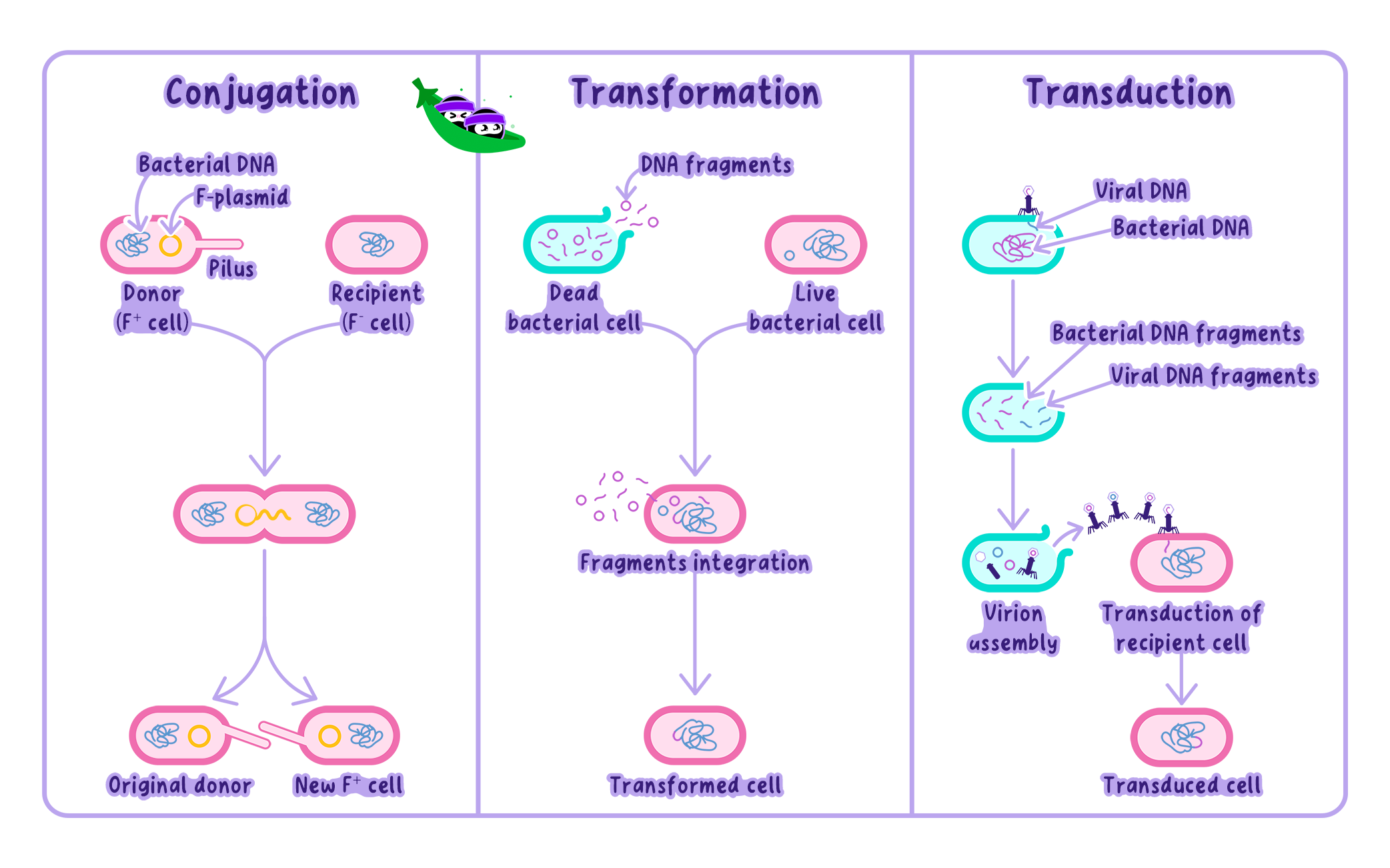
Example
Antibiotic resistance genes can spread between bacterial species, altering their genetic makeup independently of reproduction.
Why The BSC Fails in These Cases
Asexual Organisms
No interbreeding occurs, so the core definition of species under the BSC cannot be applied.
Leads to arbitrary classification into many clones or microspecies.
Horizontal Gene Transfer
Species boundaries are blurred as genetic material flows across unrelated taxa.
In bacteria, "species" boundaries are almost meaningless because of widespread gene sharing.
Exam_technique
Always state both asexuality and HGT as reasons why the BSC does not work universally.
Note
The biological species concept is useful for sexually reproducing organisms but has significant limitations when applied to asexual organisms and bacteria.
Competing species definitions exist to address these challenges.
Self Review
Define the biological species concept and explain its key limitation in organisms that reproduce asexually.
Why are Taraxacum officinale (dandelions) and Rubus fruticosus (blackberries) problematic for the BSC?
Define parthenogenesis and give one animal example.
What is horizontal gene transfer, and why does it prevent the BSC from being applied to bacteria?
A3.1.13 chromosome number as a shared trait within a species (hl)
Chromosome Mismatch Causes Hybrid Sterility
Chromosomes are the carriers of genetic information, organized in pairs in sexually reproducing organisms.
During reproduction:
Gametes (sperm and egg cells) are formed via meiosis, which reduces the chromosome number by half.
For meiosis to succeed, homologous chromosomes (matching pairs inherited from each parent) must align and segregate properly.
When two species with different chromosome numbers interbreed, their hybrid offspring may inherit an uneven chromosome set.
Example
A classic example is the mule, a hybrid between:
Horses: 64 chromosomes.
Donkeys: 62 chromosomes.
Mules inherit 63 chromosomes, which cannot form homologous pairs.
As a result:
Meiosis fails.
Functional gametes cannot form, rendering mules sterile.
Warning
It's a common misconception that all hybrids are sterile.
While this is often true, there are exceptions.
For example, some female ligers (lion X tiger hybrids) can produce offspring.
Plants Are a Special Case
Unlike animals, many plants can bypass these issues through polyploidy.
This is where entire sets of chromosomes are duplicated.
Polyploidy allows:
Homologous pairing of chromosomes.
Fertility in hybrids.
Definition
PolyploidyPolyploidy is the duplication of an organism's entire chromosome set.

Tip
Polyploidy in animals is rare because their cells are more sensitive to changes in chromosome number, whereas plant cells are more adaptable.
Why Chromosome Number is a Shared Trait
Within a species, having the same chromosome number ensures:
Viable gamete formation through homologous pairing in meiosis.
Genetic stability across generations.
Fertility of offspring, maintaining the species as a coherent unit.
Speciation and Chromosome Number
Over evolutionary time, chromosome number can change through fusion, fission, or duplication events.
When populations diverge and accumulate differences in chromosome number, interbreeding becomes impossible.
This acts as a mechanism of reproductive isolation, driving speciation.
Tip
When explaining why hybrids are infertile, always mention failure of homologous pairing in meiosis.
This is the key mechanism expected in IB answers.
Self Review
Why do all members of a species typically share the same chromosome number?
Explain why hybrids with mismatched chromosome numbers are often infertile.
How can changes in chromosome number contribute to the process of speciation?
A3.1.14 engagement with local plant or animal species to develop a dichotomous key (hl)
Developing a Dichotomous Key Using Local Plant or Animal Species
Definition
Dichotomous KeyA dichotomous key is a tool that simplifies the identification of organisms. It operates through a series of paired statements that describe contrasting traits. Each decision point leads to another pair of choices until the correct identification is achieved.
A dichotomous key is a tool used by biologists to identify unknown organisms by progressing through a series of paired, contrasting statements.
Each step provides two choices (_yes/no_ or present/absent), directing the user to the next step until the organism is identified.
This process is essential in taxonomy, fieldwork, and biodiversity studies, as it allows systematic and reliable identification of species without needing advanced molecular tools.
Example
Leaves are needle-like ... Go to Step 2 Leaves are broad and flat ... Go to Step 3
Needles are in clusters ... Pine tree Needles are single ... Spruce tree
Leaves are lobed ... Oak tree Leaves are not lobed ... Maple tree
Tip
When creating a dichotomous key, focus on traits that are easily observable and measurable, such as leaf shape, bark texture, or flower color.
Avoid subjective traits like "large" or "beautiful."
Why Use a Dichotomous Key?
Dichotomous keys are valuable for:
Fieldwork: Identifying organisms in natural habitats.
Biodiversity Studies: Cataloging species in an area.
Education: Teaching observation, classification, and critical thinking.
By creating and using a dichotomous key, you enhance your understanding of local ecosystems and sharpen scientific observation skills.
Analogy
Think of a dichotomous key as a "choose-your-own-adventure" story.
Each decision you make leads you closer to the final answer.
In this case, the species you're identifying.
Features of a Good Dichotomous Key
When constructing a dichotomous key, certain rules must be followed to ensure accuracy and usability:
Mutually exclusive statements: At each step, only one statement can be true.
Observable traits: Features should be visible without special equipment (e.g., colour, shape, structure).
Consistent characteristics: Avoid traits that vary with age, season, or environment (e.g., leaf colour in autumn).
Progressive narrowing: Each pair should split organisms into smaller groups until species are identified.
Clear outcomes: Every path should end in one species name.
Steps to Develop a Dichotomous Key
Step 1: Select a Group of Organisms
Choose a group of organisms to study. Examples:
Trees in a forest or park.
Insects on a flowering plant.
Birds visiting a feeder or wetland.
Note
Choose a group that is diverse enough to require multiple steps in the key but not so large that it becomes overwhelming.
Step 2: Observe and Record Traits
Study the organisms and record their distinguishing features. For example:
Plants: Leaf shape, bark texture, flower type, or seed structure.
Animals: Size, coloration, movement patterns, or behavior.
Example
Suppose you are studying local trees.
You might record traits like:
Leaf type: needle-like or broad
Leaf arrangement: opposite or alternate
Bark texture: smooth or rough
Presence of fruit or cones
Warning
Avoid relying on traits that are difficult to observe in the field, such as microscopic features or internal anatomy, unless you have access to specialized tools.
Step 3: Group and Organize Traits
Organize the organisms into categories based on shared traits. For instance:
Trees with needle-like leaves vs. trees with broad leaves.
Further divide these into subgroups using additional traits, like bark texture.
Further subdivide these groups using additional traits, like bark texture or fruit type.
Tok
Consistency is key.
Use traits that are stable and reliable within each species.
For example, leaf shape is a better trait than leaf color, which can vary seasonally.
Step 4: Write Paired Statements
Create pairs of contrasting statements to guide identification. Each pair should be:
Clear: Avoid ambiguous language.
Mutually Exclusive: Each option should cover all possibilities.
Warning
Steer clear of vague descriptions, such as "leaves are large" or "bark is dark."
Instead, use measurable traits, like "leaves are longer than 5 cm."
Step 5: Test and Revise
Test the key by using it to identify organisms in your study area.
Revise unclear or overlapping statements.
Ensure the key works for individuals unfamiliar with the species.
Self Review
What are the advantages of dichotomous keys compared to modern methods like DNA barcoding?
Define a dichotomous key and explain how it is structured.
Why must statements in a key be objective and mutually exclusive?
A3.1.15 identification of species from environmental dna in a habitat using barcodes (hl)
Barcoding Biodiversity: DNA Barcodes and Environmental DNA (eDNA)
Definition
DNA BarcodesDNA barcodes are short, standardized DNA sequences unique to each species, much like product barcodes in a store. These genetic "fingerprints" enable species identification, even from small or incomplete samples.
Traditional identification of species often depends on morphological features (e.g., flower shape, body markings).
Many organisms are hard to identify: plants without flowers, insect larvae, or rare animals that leave only traces behind.
DNA barcoding and environmental DNA (eDNA) provide a modern method for rapid and accurate species identification.
Example
A feather found in a forest can be analyzed for its DNA.
By amplifying the CO1 gene using polymerase chain reaction (PCR) and comparing it to a reference database, scientists can identify the bird species, regardless of visual evidence.
Environmental DNA (eDNA)
Definition
Environmental DNA (eDNA)Environmental DNA (eDNA) refers to genetic material shed by organisms into their surroundings through skin cells, mucus, feces, or other biological materials. It enables species detection without direct observation.
How Does eDNA Work?
Imagine investigating a pond's biodiversity:
Sample Collection: Collect water, soil, or air samples from the environment.
DNA Extraction: Use chemical and physical methods to isolate DNA.
Amplification: Amplify barcoding regions using PCR to increase detectable DNA.
Sequencing: Sequence the amplified DNA to read its genetic code.
Comparison: Match sequences to a reference database to identify species.
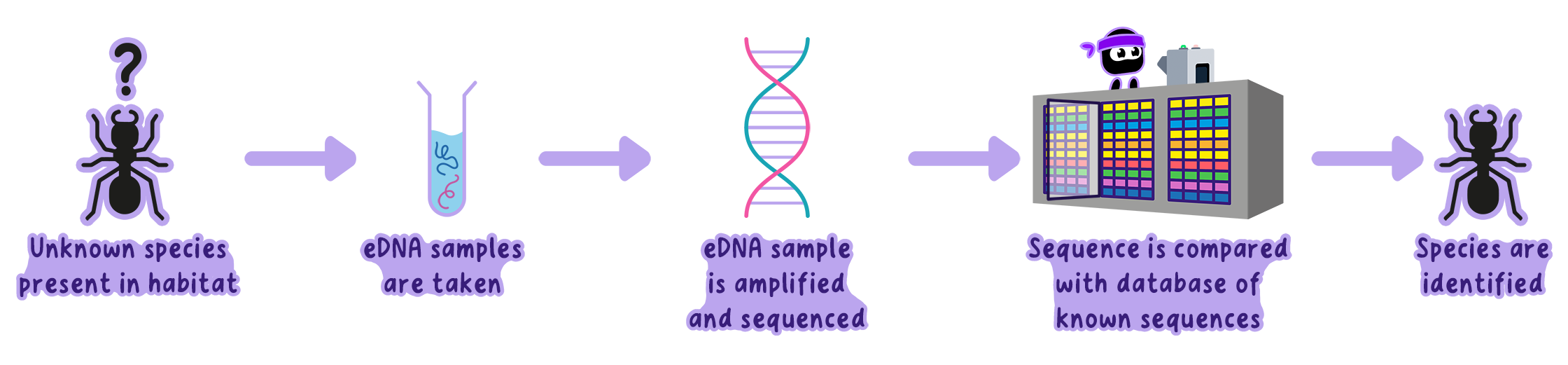
Note
Its advantages are:
Allowing detection of elusive, nocturnal, or rare species.
Non-invasive, preserving species and habitats.
DNA Barcodes: The Genetic "Fingerprint"
DNA barcodes are chosen from genes conserved across species, but which also show enough variation between species.
Animals: Cytochrome oxidase subunit I (COI) in mitochondrial DNA is widely used.
Plants: Chloroplast genes rbcL and matK.
Prokaryotes: 16S ribosomal RNA genes.
Applications of DNA Barcoding and eDNA
Rapid Biodiversity Assessments: Barcoding and eDNA enable quick surveys of biodiversity, which can inform conservation priorities.
Monitoring Invasive Species: Early detection of invasive species prevents ecosystem disruption.
Tracking Endangered Species: DNA-based methods confirm the presence of endangered species without disturbing them.
Disease Surveillance: eDNA can track pathogens in the environment, providing early warnings of outbreaks.
Advantages of eDNA
Non-invasive: eDNA can be collected without harming organisms.
Efficient: Barcoding and eDNA allow for rapid data collection and analysis.
Broad Scope: These methods can detect a wide range of species, including those that are rare or difficult to observe.
Cost-Effective: Compared to traditional surveys, eDNA analysis requires fewer resources and less field time.
Warning
One common mistake is assuming eDNA can always provide a complete picture of biodiversity.
Factors like sampling method, environmental conditions, and database quality can all affect results.
Limitations of eDNA
Degradation: eDNA breaks down quickly, requiring timely sample collection.
Database Dependence: Accurate identification relies on comprehensive, up-to-date reference databases.
False Positives/Negatives:
Contamination may cause false positives.
Degraded or low-quality DNA can lead to missed detections.
Taxonomic Resolution: Barcodes may not distinguish between closely related species.
Example
Sampling water for eDNA in a heavily polluted area could lead to degraded DNA, making species identification difficult or incomplete.
Self Review
Define DNA barcoding and explain how it differs from full genome sequencing.
What is environmental DNA and why is it especially useful in detecting rare or elusive species?