Genetic Diseases and Mutations
Principles of genetic Variation
Genetic variation is the difference between DNA sequences that are being compared
specific ones were found after the human
composed of library with large amount of people including twins
mutations are a large source of genetic variation and mutations are generally thought to be caused by external sources but it’s mostly the endogenous cell environment
most are not inherited unless early dev. of embryo or in germline cells
genome projects typically used a number of DNA donors with Craig Venten’s was the 1st whole genome
there are 2 main categories of genetic variation
changes that don’t affect the DNA content
there is no net gain or loss of nucleotides think single nucleotide replaced with another
changes in the copy number
think insertion or deletion of single or a small number of nucleotides
most common type of changes are small scale by larger variations (>50bo) have a larger affect
SNV (single nucleotide variation) → changes in a single nucleotide frequency <0.01; SNP Frequency>0.01
most common type of change in the genome
Indel →nucleotide is present of absent ie insertion or deletion variation (polymorphism) most of which are not new and come from evolutionary changes or ancestry
Variable tandem repats →sequences is repeated after another and generally unstable
DNA variants that differ in the number of tandem repeats
aka variable number of tandem repeats (VNTR) polymorphism
Satellite DNA: Array of 20 kb – hundreds of kb
Located at centromeres and other heterochromatic regions
Minisatellite DNA: Array of 100 bp – 20 kb
Primarily at telomeres and subtelomeres
microsatellite DNA: array of ~100 bp or >
repeat of ~2-4 nucleotides and is widely distributed
aka shor tandem repeats (STRs)
highly variable with several different alleles present in the population
used in forensics and high output PCR
Strand slippage→ happens during DNA replication where an STR is inserted or deleted during DNA replication
aren’t caught by the double checking mechanism cause of they are repeats
Balanced structural variation→ Same DNA content but the sequences are located in different positions
think translocations, inversions
SNP is an example of balanced genetic change
Unbalanced changes→ changes the copy numbers can result in diseases or metabolism phenotypes
can also change drug dosing
think CNV (copy number variations) and deletion or duplication
most variations has neutral affect cause proteins have multiple loci; 2 x amount of chromos; hard to actually change the amino acid that has an affect; only small amount is non coding and there is a large amount of redundancy in the genome
variation is beneficial when it increases survival or increases the fitness
becomes prevalent
think brain devleopment, cognitive function etc. for humans vs. gorillas
negative/ purifying selection happens when is leads to a lower fitness
different geographic regions, exposure to pathogens, and altered environments shaped the genome through selection
Infectious disease are considered to be one of the strongest selective pressures in human evolution
think the black plague those with a certain gene mutation were more likely to survive than those without it →Gene ERAP1/ ERAP2 has a C
positive change tracked with increase in those survived and increase in those who died
in gene study was ERAP 1 and 2 which helps T cell recognize toxins and their anitigens
changes in regulators is the most deferable in apes and humans are expectation is myostatin heavy chain 16→ changes jaw structure which increases skull capacity
context matters for natural selection
example people with higher HLPC expression that were to protect against animal pathogens but now people don’t have to worry about that so it’s not selected for now
can also be the reason why people allergies
UV radiation stimulates a phot lytic reaction in dermis that make vitamin D3 but also causes DNA damage (sun burn= cell death) so melaninated skin was needed
when people migrated north there was less sun which means less vitamin D3 this “lead” to the mutation in SLC24A5 gene a calcium transporter that regulates melanin that reduced skin pigmentation and become prevalent in Europeans
selective sweep→ positive selection for advantageous DNA can leave a signature in the genome
population variation is reduced at sequences immediately next to the gene variant
occurs when a mutation that is subject to positive selection increases in frequency to become the common allele and the adjacent variant hitchhike with it
chromosome with mutation is not passed down as a unit cause of recombination
also seen in SLC24A5 gene
changes to diet →high starch and consuming milk life long → ancestorial trait to digest lactose for nutrition till a kid a change in the promoter region of the gene (c→T) so that it turns on and makes lactose all the time
amylase in saliva is made by AMYIA→ breaks down starch and theres a copy number variations between chimps have 2; 2-15 diploid copy number CNV
more copies higher the enzyme concentration
used high res FISH to see how many copy og the genes there are
gene duplication →generations may have slightly different forms of a protein
positive selection means copy number stays the same over generations
largest protein coding gene is olfactory receptor family in humans which are used for food and not the environment
olfactory receptors are different in everybody each person makes about 500 variants this impacts food taste and preference
Single Gene Disorders: inheritance patterns, phenotype variation, and allele frequencies
a single locus can be primary determinant → like Mendel or Monogenic genetics
a particular genotype at a single locus is necessary and sufficient for a particular phenotype under normal circumstances
traits can be dominant or recessive
although rare single gene disorders are important contributors to disease and can be visualized using a pedigree
pedigree symbols to know
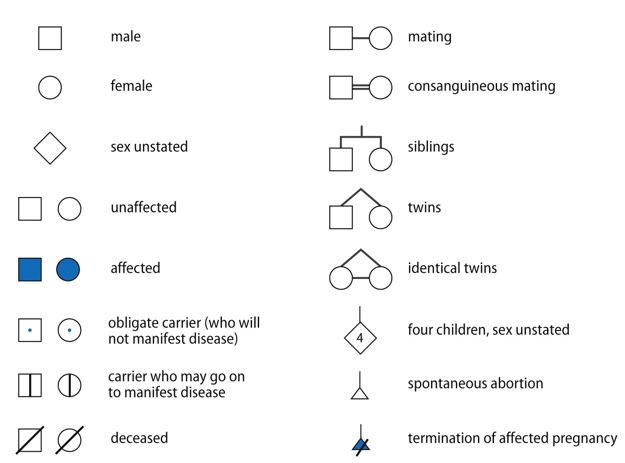 highlight: consanguineous mating → 1st or 2nd cousins married; diamond means unknown state
highlight: consanguineous mating → 1st or 2nd cousins married; diamond means unknown statecarriers may not always be shown in a pedigree
autosomal dominant →almost always heterozygous people are affected; unaffected individuals usually don’t transmit the disease and both sexes are affected equally
default assumption cause two means dead or in super serious conditions
think 2 people with dwarfism has kids if homozygote
usually parent is affected seen in every gen
achondroplasia is the most common cause of dwarfism
caused by mutation in FGFR3 that cause liganand independent activation of FGFR3 ie it is overactive
normally it reduces bone growth
typcially G→A
homozygous is lethal if heterozygous you got it and the other allele is compensating for the dysfunctional one
de novo mutations → new mutations in the parents and almost always come from the dad
occur in paternal germline and are associated with a high paternal age
comes from mutation of stem cells cause of age
autosomal recessive inheritance→ inherited in 2 genes aka 2 mutant alleles at the locus
one comes from each parent
keep in mind if parents are related is huge for autosomal genes suually 2nd cousins or 1st cousins and since genomes are related with 1/8th -1/32 high chance of having same allele
affects both sexes equally and two carriers have a 25% chance of having an affected child
for disorders with unknown inheritance, parental consanguinity is a strong indicator of autosomal recessive inheritance
consanguinity produces affected people with identical mutant alleles due to common ancestry
if recessive disorder is frequent in population there are a lot of carriers
two unrelated parents can carry different mutant alleles
compound heterozygous 2 different mutation present in the gene
typically, compound heterozygotes are phenotypically similar unless they are at different severities
Example of autosomal recessive disorder→ cystic fibrosis → mutations in CFTR gene encodes chloride channel and loss of CFTR function leads to abnormal fluid and electrolyte transport which results in common symptoms below
Failure to reabsorb chloride in the sweat
Depletion of airway surface liquid in the lung
Defective secretion of pancreatic enzyme
reduced fertility
x linked recessive disorders→ mostly males are affected born to unaffected parents
mother of affected male is carrier and has affected male relatives
IS NOT TRANSMITTED FROM DAD TO SON
dad with mutation will transfer the allele to all daughters who will be carriers
can
if mom is carrier and dad is unaddected → son has it; if next child is son there is 50% chance the son has it or is unaffected and if the next child is a female zero chance of having it and 50% chance of being a carrier
complications with inbreeding cause it makes it harder to tell if its Y linked or X-linked until a female gets the disorder
X- linked dominant disorders → one parent is affected and affected individuals can be male or female
higher chance of being an affected female than males
affected females typically have milder disease then males
if the father is affected all daughteres will have the disorder and none of the sons (depending on the mother) will be affected
if the mother is affected the transmission is equal in the sexes
in this case A causes disease
females can be heterozygous and unaffected cause of scilencing of the affected X
she has a genetic mosaic
males are constitutionally hemizygous (1 allele) and females are functionally hemizygous have 2 alleles but only use the one
Klinefelter’s syndrome will have 2+ silenced genes
consequence of genetic mosaic females that are affected will have partial functionally compared to males who are affected with 0/ defective products
25-20% of silenced genes escaped
skewed x inactivation→ the favoring of silencing 1 gene over another
Ex. if quite gene makes functional protein may be more favored for it not to be silenced
can occur cause inactivation of mutant X provides survival advantage to cells, inactivation of normal X provides advantage, and random chance
if x linked recessive disease and the mutant X is activated then bad luck if x linked dominant and the dominant allele is silenced then good luck
male lethality→ being homo/ hemizygous is so severe that the male dies but the x inactivation in males allows them to live
lethality in males → is X linked dominance disease
pseudoautosomal regions → 2 gene containing regions in common present on X and Y or X and X
don’t produce X and Y linked patterns
pair during male meiosis and undergo recombination, including obligate crossover in major PAR
contain ~29 genes
mutation will be autosomal
pseudo autosomal inheritance is like autosomal
gene pairs in pseudoautosomal regions are effectively alleles
copies are present on both X and Y and can move between chromosomes by recombination so don’t act X- linked or Y- linked
males can pass it on to either female/ male offspring
resembles autosomal inheritance ie dominant inheritance of a mutation in the PAR
one person can be affect but another won’t be cause of crossing over
Y linked inheritance→ male specific region of the Y chromosome makes 31 different proteins
proteins are involved in the normal development of testes, germ cells, and fertility
no known trait gives stereotypical Y-linked pedigree
Y chromosome infertility typically caused by new mutations
IVF fertility can’t get passed on for male infertility
pedigree features → only males are affected; exclusive father to son transmission
females have 2 allele for all genes in the pseudo autosomal regions both alleles are active; in males both are active and have an allele on each chromosome
mtDNA is more prone to mutation than nuclear DNA
disorders primarily affect tissues with high energy requirements think muscle, brain
not packed with histones and stuff it’s circular, prone to mutation from ROS
inheritance is matrilineal → affected individuals can be either sex but affected males don’t transmit the condition
most people with mtDNA disorder have a mix of normal and mutated mtDNA
aka they are heteroplasmy
homoplasy→ all the DNA is mutated (rare)
disease pheontype is only observed if the proportion of mutant exceeds threshold
typically 60-807 mutations
the ratio of normal to mutant mtDNA can change over a person’s lifetime due to relaxed replication of mtDNA
if most of the mtDNA is mutant means it sloppy replication
mtDNA disorders can have a highly variable phenotype within families due to variable heteroplasmy
oocytes contain >100.000 mtDNA molecules but during development go through a bottleneck stage with little mtDNA so the expansion of these few can create different levels of mutant mtDNA in the oocytes
Oocytes are immature egg cells found in the ovaries. They are the female germ cells that can develop into mature eggs through the process of oogenesis.
will see skipping of generation in the pedigree
if mom doesn’t have the disorder probably cause the amount of DNA passed on through the bottleneck was large
example of mtDNA disorder leigh syndrome→ severe neurological disorder that manifests in 1st year of life cause patches of damaged tissue develop in the brain leading to progressive loss of movement
dead by the age of 3
caused by mutation in more then 75 different genes that function in ATP production
~20% of people carry a mutation in mtDNA
most common mtDNA mutation affects mt-ATP6 which is apart of ATP synthase
can take mitochondria DNA from healthy eggs and is allowed in the UK and Australia but banned in Canada cause of ethical concerns
Genetic Variation and Disease causing abnormalities
Pathogenic DNA variation can cause disease in two broad ways:
Change in sequence of gene product
Total loss of function or reduced normal function
Altered or new function that is harmful (= gain of function)
Change in the amount of gene product through:
Altered copy number
large >50 bp; small change <50
most common is small scale changes.
more affective is when it’s a large change or a lot of small changes
Change in gene regulation
Premature termination codon
Loss of function mutations:→ final gene product is not produced/ produced at low levels or is made but doesn’t work properly
Results in recessive disease when both alleles are pathogenic variants
May sometimes result in dominant diseases with one pathogenic allele
Effects include haploinsufficiency and dominant-negative effects
Example: Cystic fibrosis
Caused by loss-of-function mutations in CFTR both of which enode for a chloride channel
normal stimulates water movement and produces thin, freely flowing mucus
Disease occurs when both alleles have a loss-of-function mutation bringing function close to zero
Haploinsufficiency: genes that are very sensitive to dosage and 50% is not sufficient for normal function
Reduction from 100% to 50% of gene product results in disease
Results in dominant disease observed in heterozygotes with 1 mutant allele
Example: Brachydactyly mental retardation syndrome→ associated with large deletions at 2q37
one copy remaining of the deletion for all genes means it’s down 50%
Caused by haploinsufficiency of HDAC4
causes intellectual and behavioral issues, short fingers and short toe
Dominant-Negative effects: NOT A DELETION
Mutation produces a non-functional mutant protein that inhibits the function of the normal protein
occurs because protein forms a multimer that becomes non-functional when mutant is incorporated
Disease results because there is <50% functional product
Example: Osteogenesis imperfecta→ brittle bones collagen is gone
depends on severity mild means there is 50% collagen present while severe will have an abnormal 1A1→3/4 polypeptide sequence baby will die in utero or 1-2 day after birth
Shows the genetic mechanism involving COL1A1 and COL1A2 mutations
Gain of function mutations:
Results in gene products with new or harmful functions
protein might not properly respond to regulatory signals
protein that exhibits inappropriate expression
eg: over expression ro expressed at wrong time in wrong tissue or in response to wrong signal
enzyme with alter target specify
protein that forms toxic aggregates
Leads to dominant disease even with one copy of the pathogenic mutation or other allele is producing the correct product
Example: Achondroplasia
Caused by mutations in FGFR3 leading to ligand-independent activation of signaling
Unstable expansion of repeats:
Some tandem repeat sequences can expand and result in disease aka dynamic mutations
disease results due to production of toxic protein or RNA that is harmful
is gain of function dominant disease
Anticipation:
Individuals with high repeat numbers are more severely affected
transmission of a mutant allele in a family can lead to increase repeat number and disease severity with each generation =Anticipation
Disease severity increases with each generation due to repeat expansion
can exhibit a premutation stage → repeat array has expanded to a size that is unstable
won’t result in disease but may readily increase in length causing disease in subsequent generations
Example: Huntington disease→ motor abnormalities, personality changes, gradual loss of cognition and death
CAG repeats 6-26 is normal
CAG repeats>40 is huntington disease
36-39 repeats is incomplete penetrance (ie may or may not have it)
27-35 repeats is risk of expansion
sperm is more tolerant to these changes
Symptoms and age of onset correlate with increased number of repeats
higher expansion when transmitted by father
pedigree with square and : inside to show symptoms cause they have more than 40 repeats
with dominate diseases can expect there to still be normal in kids
recessive diseases →always a loss of function
duplication think gain of function
difference between dominant negative→ interacting with another protein that’s good causing it to lose function→ key it functions in a dimer
DNA forensics
use DNA to find criminals, heritage and parents
OG way to do this DNA fingerprinting → different cut sites and banding leads to a lot of DNA
0.5 to a ml of sample is needed and is not well automated
needed digestion of DNA, gel electrophoresis and Southern blotting of fenomic DNA
probes hybridized to hypervariable minisatellites spread across the genome
found to prouce individual specific fingerprints
is super time consuming
PCR became gold standard with the use of microsatellites (STRs) cause little DNA is needed
STRs have variable amount of repeats in each person the more sites you compare the lower the chance that there are multiple people that have that DNA sequence in population
these repeats come from slippage and inheritance
king tut’s family→ sequenced with pcr to find patents/ others with number of repeats and lineage
PCR is touchy and sensitive cause DNA from researchers can interfere with the results
in this technique your mostly calculating probability (unless mismatch)→ can’t say for sure it’s X’s DNA cause you looking at some markers and not the whole genome
especially when DNA amount is small/ degraded
coduses (dataset and standard of gene markers in population) are base on the population
match probability is 1/10 trillion
new tech for DNA samples→ can PCR different samples at the same time and find results
can use hardy weinberg equation and multiplication to get frequency of the allele in population depending on allele and frequency present use propitiate part of equation then put value to -1 and get occurrence in population
hardy weinberg equation=p²( for AA) +2pq (for net)+q² (for aa)
then multiply all the net frequency’s for the replicates
inverse the value to get repeats in population
Familial DNA searching→ only works for close relatives
then they try to zone in on the killer using other factors or using y chromosome which has conserved DNA from father to son
based on IBDs or identical by descent → more closely related people will have greater IBD than more distantly related people
rec. DNA databases which is not in codus look for +700,000 SNPs using microarrays before SNP of interest
genomic DNA is amplified and fragmented, then hybridized to a microarray contain 100s of 1000s of probes attached to beads
probe sequence ends one base before the SNP of interest
single BP extension adds a single labelled base to the probe complementary to the SNP
DNA is then washed away and the signals are read
most DTC test don’t voluntarily cooperate with law enforcement but thrid party application that provide further analysis like GEDmatch and FAmilyTreeDNA do depending on the case and the person opting in or out of the program
y chromosome markers stay the same since they are transferred from father to son only and are germline cells
Journal article notes
What are DTC tests?
tests that are sold to consumer directly instead of under doctor recommendations
what type of info do they give and what information do they get?
get raw data and then go to databases to analyze them
raw data is the 700,000 SNP/ genetic variants
can give limited amount of analysis
difference between DTC and 3rd party services like GEDmatch?
3rd party services are used for further analysis of the data while DTC will get the raw data and give you an analysis
IBD segments
large sections of DNA that are retained with in a family as time passes these sections become smaller
the larger the sections are the more related the people are
segments become smaller during meiosis cause of gene cross over and recombination
long range familial searches
used by law enforcement to find suspects
public data bases are used to find long distance relatives of the suspect and narrowing it down to the criminal
relative must be in data base and this is what is referred to as a match
DTC databases look at more matches then those in the forensics database
was used to find the golden state killer
DTC and forensic databases ethical biases
forensic databases are mostly people of color and immigrants cause they are more likely to be arrested as suspects in crimes
DTC are more white and wealthy people cause these people want to do the tests and can afford it
if you are white or wealthy you are more likely to have a match in the DTC then people of colour
Figure 1a: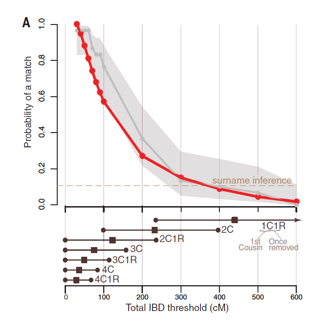
red line made using all public DTC databases and grey is only a portion so that they could get a confidence interval
x-axis shows relatedness to target
chance of finding a match increases when the relatedness decreases
allow for more variation that allows for more ancestors
Figure 1b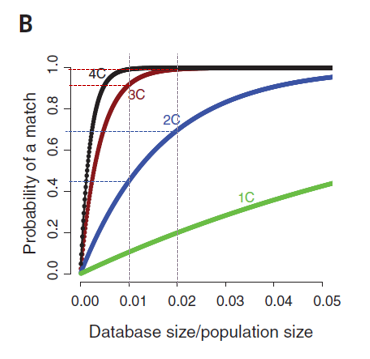
the larger the database is the easier it is to find a more closely related match
optimal fast plateau with small database size
Figure 2A: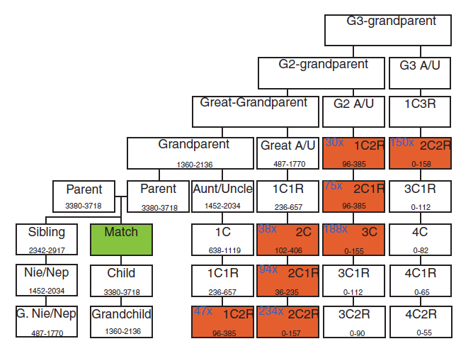
match is the relative in the databases
people in red are the potential suspect with range of cM at the bottom of box
the average number of relative at the positions add in blue
add them together to get total number of suspects
Figure 2B: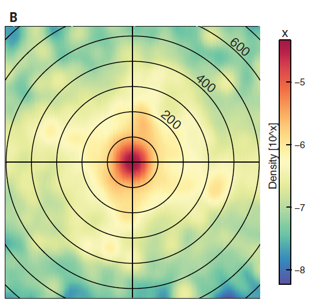
how close or far way they are from each other to narrow down location and limit this radius according to the match and the scene of the crime
Figure 2C and D: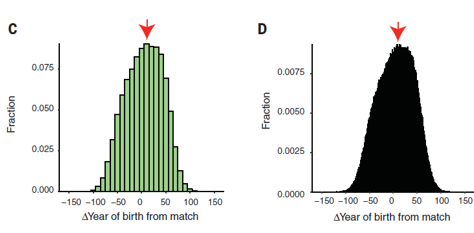
showed amount of matches and their age relative to the match →used to narrow down suspects
closer the resolution of year the less suspects present
Pipeline of these searches:
entire population→ gene match→ geography→ sex (gained from DNA sample)→ age (10 year resolution 16-17 people; 1 year resolution 1-2 people )
Main barrier of technique
it is not finding a match it’s the time and resources it takes to find a target
also requires info about the family tree that is public
Privacy concerns exist:
wanted to see how hard it would be to re-find the people in studies and their ancestry couple
found couple with in an hour and target with-in the day
anyone could do this and locate people used in studies
gene data isn’t classified as identifiable and should be; DTC companies should encrypt txt files to ensure that 3rd party is authorized to have access
Figure 3: 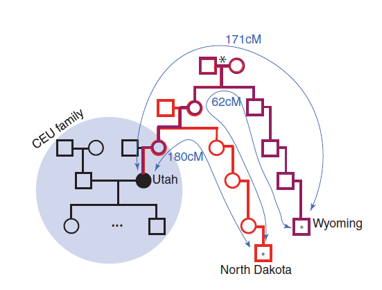
black circle is the target person; dots are distant causes that are matched made by going back to find most recent common ancestor