Polymerase Chain Reaction (PCR)
PCR Overview
The Polymerase Chain Reaction (PCR) is a revolutionary technique developed by Kary Mullis in 1987. It allows for the artificial amplification of specific DNA sequences, enabling the production of billions of copies of target DNA within a few hours, thus facilitating various applications in molecular biology, diagnostics, forensics, and evolutionary studies.
Basic Principles and Tools of PCR
Key Concepts
Target DNA: The specific sequence of interest, typically a fragment of a gene.
Primer: A short segment of DNA that binds to the template strand of the target sequence. Primers are essential for allowing DNA to be amplified by Taq polymerase.
Taq polymerase: A thermostable DNA polymerase derived from the bacterium Thermus aquaticus, which enables DNA synthesis at high temperatures.
Thermal Cycler: An automated instrument that rapidly cycles through the necessary temperature changes for denaturation, annealing, and extension during PCR.
dNTPs: Deoxynucleotide triphosphates, the building blocks for new DNA strand synthesis.
Magnesium ions (Mg²⁺) and buffers are also requisite components for the reactions.
PCR Process Overview
The PCR process can be visualized in three main stages:
Denaturation: DNA strands are separated by heating the sample to approximately 95°C, breaking the hydrogen bonds between complementary bases, allowing the two strands to become single-stranded DNA.
Annealing: The temperature is lowered to around 55°C, allowing the primers to bind to their complementary sequences on the target DNA. Optimal binding occurs at a temperature typically set at 3°C below the melting point of the primers, which can be calculated using the formula:
\text{Melt Point} = 4(G+C) + 2(A+T)Because G and C have stronger hydrogen bonds than A and T
Extension: The temperature is raised to 72°C, the optimal temperature for Taq polymerase activity, allowing for the synthesis of the new DNA strand starting from the bound primers.
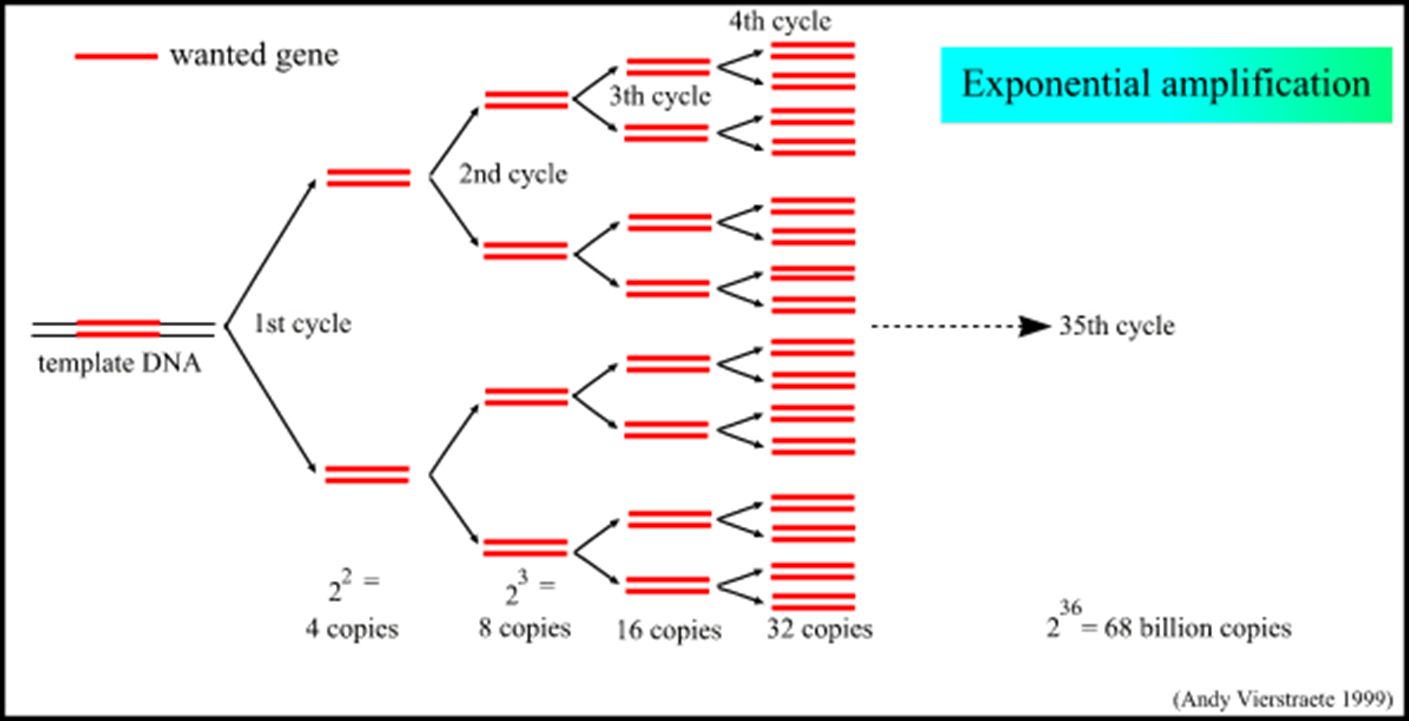
Repeated Cycles

Each complete cycle doubles the amount of target DNA, leading to exponential amplification.
For example, upon completing 32 cycles, the number of double-stranded target molecules can reach approximately 2^{32} = 1,073,741,824 copies.
Designing Primers
Designing effective primers is crucial for successful PCR amplification. The following guidelines are recommended:
Avoid excessive G/C or A/T content; aim for approximately 50% of each to ensure balanced binding.
Primers should be 16-24 nucleotides long for specificity while maintaining the target's fidelity.
The melting point of primers should exceed 55°C to ensure successful binding.
Limitations of Taq Polymerase
While Taq polymerase is favoured for its rapid DNA synthesis and high processivity, it possesses some limitations:
Taq polymerase is error prone; rate of misincorporation of bases = 285/106 (cf with E. coli DNA polymerase I = 9/106).
Single errors will multiply at each point in subsequent cycles.
Can be reduced by use of VentTM DNA polymerase that has proofreading activity.
Also recombinant Taq that is a genetically modified version with a proofreading enzyme fused to it... high processivity,
Types of PCR Reactions
Single-locus PCR
Requires knowledge of the target sequence, including nested PCR and inverse PCR.
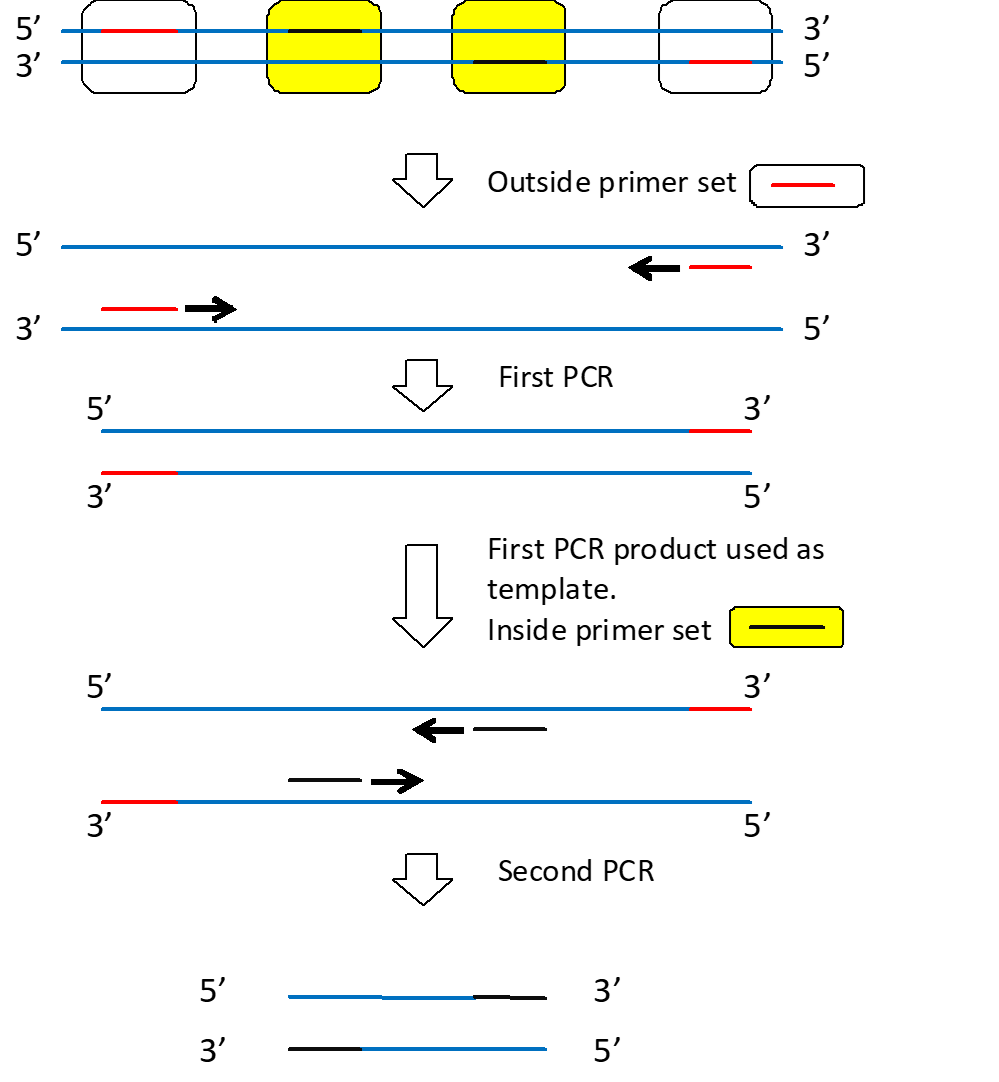
Nested PCR
Useful for enhancing target amplification at very low levels or where the first amplifying primer misprimes. This involves performing an initial PCR, then using the PCR products with a second set of internal primers to reamplify for greater specificity and copy number.
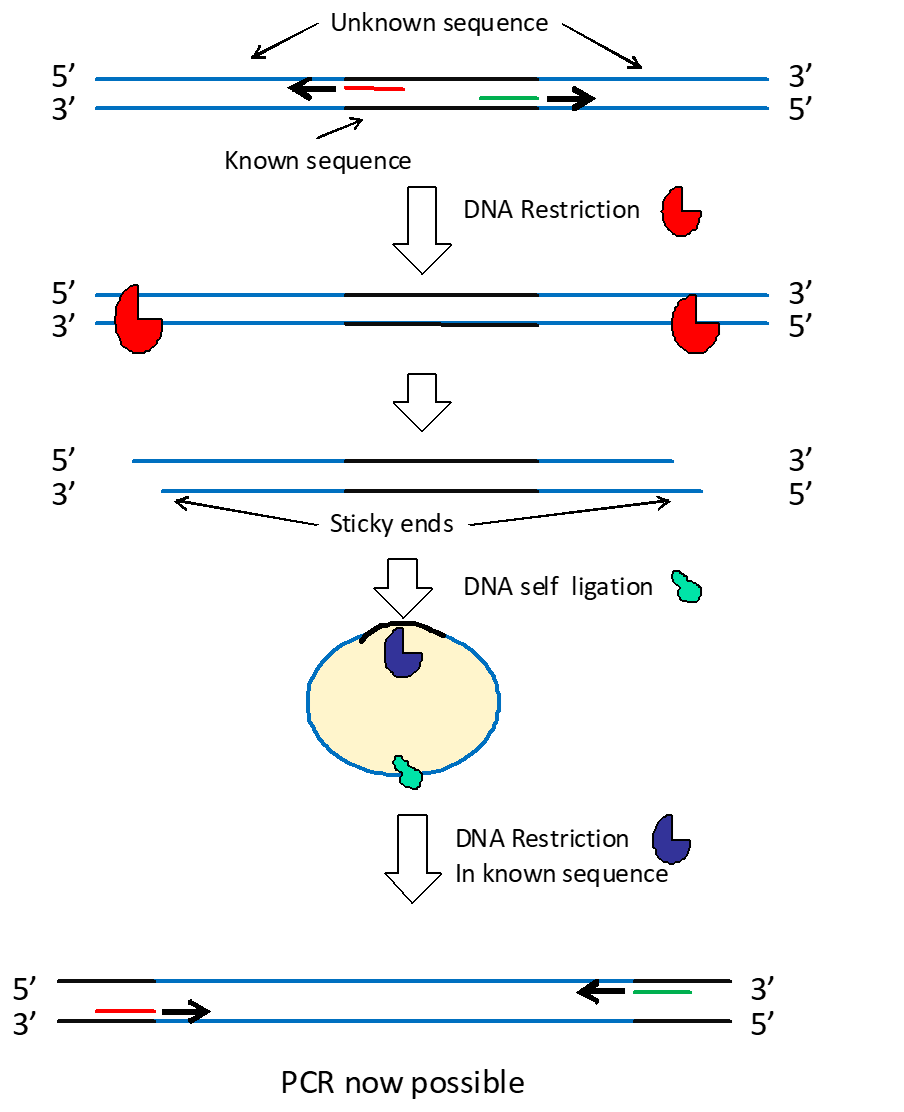
Inverse PCR
Applies when part of the DNA sequence is known, but the surrounding DNA is not. The DNA is fragmented with restriction enzymes, circularized, and PCR is conducted with primers amplifying outwards from the known sequence.
Multi-locus PCR
Does not require prior knowledge of the DNA sequence, including methods like RAPDs (Random Amplified Polymorphic DNA), ISSRs (Inter Simple Sequence Repeats), and AFLPs (Amplification Fragment Length Polymorphisms).
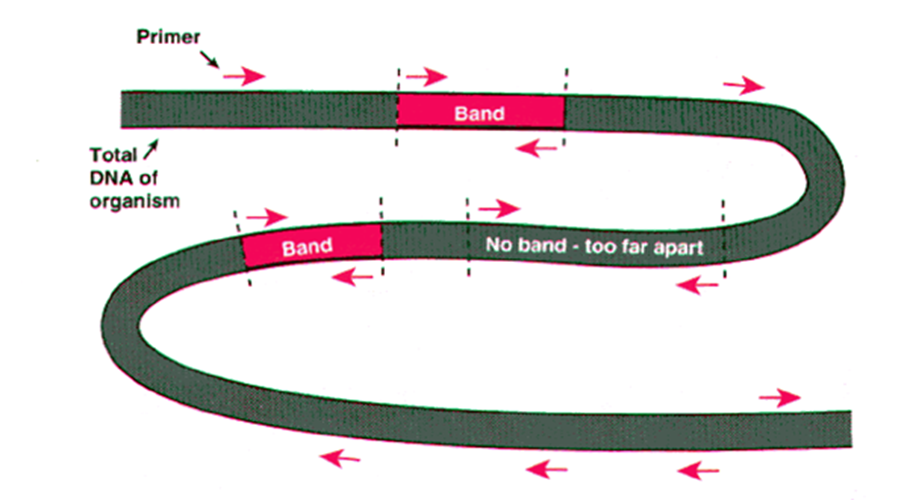
RAPDs
Employ random sequences of short primers that can amplify polymorphic regions of DNA, with a statistical notion that the chance of a random sequence matching an existing sequence is governed by the equation 4^{nt} , where nt = length of the primer.
ISSRs
Are a multi-locus PCR technique that amplifies regions between simple sequence repeats without prior DNA sequence knowledge.
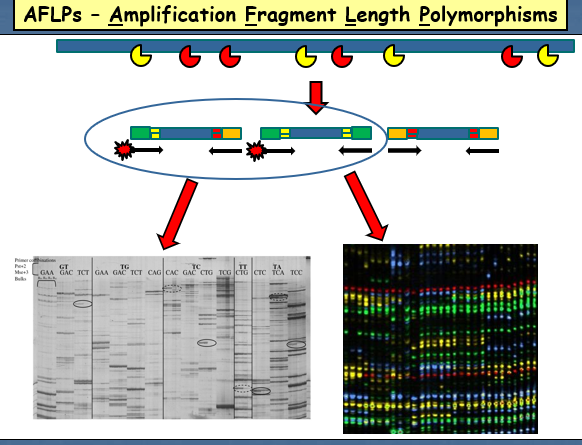
AFLPs
Are also a multi-locus PCR method that does not require prior knowledge of the DNA sequence, used for amplifying polymorphic regions.
Applications of PCR
Forensics and Paternity Testing: PCR is extensively employed to analyze specific DNA regions to discern individual identities and establish family relationships.
Clinical Diagnostics: Used for detecting latent infections, e.g., HIV and tuberculosis. PCR aids in rapid diagnostics by amplifying specific sequences related to these pathogens, facilitating early intervention.
Population Genetics and Gene Mapping: Involves isolation and amplification of genes, contributing to genetic diversity studies and gene location in organisms.
Archaeological Studies: PCR's sensitivity allows for the examination of ancient DNA, aiding in understanding evolutionary biology.
Sex Determination in Embryos: PCR can also be used for determining the sex of embryos by targeting sequences unique to the Y chromosome.
Conclusion
PCR stands as a cornerstone technique in molecular biology, significantly amplifying DNA with high precision and sensitivity. Its applications span across various fields, proving indispensable for research, clinical diagnostics, and forensic investigations. Understanding its mechanisms, limitations, and optimal use of reagents is essential for harnessing the full potential of PCR.