Enzymes
Introduction to Biological Catalysis:
Enzymes are biological molecules that function as catalysts, significantly increasing the rate of chemical reactions without being consumed in the process.
Molecular Composition: While the vast majority of enzymes are proteins, certain catalytic RNA molecules exist, known as ribozymes.
Fundamental Role: The primary objective of an enzyme is to facilitate reactions that would otherwise occur too slowly to support life. They achieve this by providing an alternative reaction pathway with a lower energy requirement.
Chemical Kinetics and Reaction Rates:
To understand enzyme action, one must first understand the simplest chemical reaction where a reactant (A) is converted into a product (B).
Rate Equations: The reaction rate is defined by the change in concentration of reactants or products over time. In a first-order reaction, the rate is dependent on the concentration of the reactants: Reaction rate = k[A].
The Rate Constant ($k$): This is a constant of proportionality that describes the proportion of reactant that will react per unit of time, typically measured in s-1.
Exponential Decay: As the reaction proceeds, the decrease in substrate concentration follows an exponential decay model, expressed as:
, where [A]0 is the initial concentration and t is time.
Thermodynamics vs. Reaction Kinetics:
A critical distinction must be made between whether a reaction can happen (thermodynamics) and how fast it happens (kinetics).
Thermodynamic Favourability:
Reactions are depicted using free energy diagrams where the progress is shown along a reaction coordinate.
Exothermic Reactions: When the change in free energy (△GR-P) is less than zero (△G < 0), the reaction is thermodynamically favourable and spontaneous.
The Spontaneity Paradox: Spontaneous does not mean fast. For example, the hydrolysis of ATP to ADP and Pi has a highly favourable △G0 = -32kJmol-1, yet ATP remains stable in solution for days at room temperature.
Rate Enhancement by Enzymes:
Without enzymes, biological reactions are incredibly sluggish. For instance, the half-life for glycine decarboxylation is 1.1 billion years. Enzymes provide massive rate enhancements, typically ranging from 105 to 1017 fold.
Chorismate mutase: Increases rate by 106.
Triosephosphate isomerase: Increases rate by 109.
Carboxypeptidase: Increases rate by 1013.
Transition State Theory:
Transition State Theory provides the framework for understanding how enzymes accelerate reactions by focusing on the high-energy barrier between reactants and products.
The Transition State (TS):
The transition state is a high-energy, very unstable chemical species that exists briefly 10-13 to 10-14s during the reaction. It is a point where original bonds are partially broken and new bonds are partially formed.
Activation Energy (△G‡):
The difference in free energy between the reactants in their ground state and the transition state is the activation energy (△G‡).
Rate Relationship: The rate constant (k) is inversely and exponentially related to the activation energy. This is described by the Eyring Equation:
Where κ is the Boltzmann constant, h is Planck’s constant, R is the gas constant, and T is temperature.
Enzyme Action: Enzymes speed up reactions by reducing △G‡ through the stabilisation of the transition state. Crucially, the overall △G of the reaction remains unchanged.
Catalytic Mechanisms:
Enzymes reduce the activation energy by addressing the enthalpic and entropic components of the reaction.
Enthalpic Stabilisation (△H‡):
This involves the direct stabilisation of charges and the facilitation of bond making/breaking.
General Acid-Base Catalysis: This mechanism prevents the development of unfavourable charges.
Acid Catalysis: Stabilises developing negative charges by transferring a proton (H+) from an acidic group on the enzyme.
Base Catalysis: Stabilises developing positive charges by accepting a proton.
Electrostatic Catalysis: The enzyme active site uses charged side chains or metal ions (like Zn2+ in carbonic anhydrase) to create an electric field that stabilises transition state charges.
Entropic Stabilisation (△S‡):
Entropy is a measure of disorder; more disorder is generally energetically favourable.
Proximity and Orientation: Enzymes bring substrates together in the correct alignment for a reaction to occur.
This effectively converts a bimolecular reaction (high entropy loss) into a unimolecular-like reaction (low entropy loss), significantly increasing the effective concentration of reactants.
Covalent Catalysis: In some cases, the enzyme forms a temporary covalent bond with the substrate, providing a highly reactive intermediate that further lowers the energy barrier.
Chymotrypsin is a classic example of an enzyme utilising this mechanism.
Introduction to Enzyme Kinetics:
Enzyme kinetics involves the characterisation of biological phenomena with respect to time, specifically measuring the change in concentration of chemical species as a reaction proceeds.
Role of Enzymes: It is fundamental to note that enzymes affect only the kinetics (rate) of a reaction and do not alter the thermodynamic equilibrium.
Experimental Approach: To analyse kinetics, data is typically collected at a constant enzyme concentration ([E]) using several different substrate concentrations ($[S]$). The initial rate or velocity (V0) is then calculated for each.
Reaction Velocity: A plot of V0 against [S] typically yields a hyperbolic curve.
Region I (Low [S]): The velocity is approximately linear with respect to the substrate concentration.
Region II (High [S]): The velocity becomes approximately independent of the substrate concentration as the enzyme reaches saturation.
The Michaelis-Menten Model:
To quantify kinetic data, the Michaelis-Menten equation is employed: .
Kinetic Parameters:
Vmax (Maximum Velocity): This represents the maximum reaction rate catalysed by a given concentration of enzyme when it is fully saturated with substrate.
Km (Michaelis Constant): This is defined as the substrate concentration at which the reaction rate is exactly half of the maximal velocity (Vmax / 2).
Mathematical Derivation and Assumptions:
Initial Velocity Assumption: By measuring only the initial rate (V0), the back reaction of product (P) to substrate (S) (governed by k-2) is assumed to be insignificant and can be ignored.
Steady-State Approximation: When [E] [S], it is assumed that the concentration of the enzyme-substrate complex ([ES]) remains constant over time. Therefore, the rate of ES formation equals the rate of ES breakdown:
The Michaelis Constant: Km is mathematically derived from these rate constants as
Biological Significance of Km:
Km provides vital information regarding the affinity of an enzyme for its substrate.
Affinity Levels: A low Km indicates strong substrate binding (often in the micromolar range), whereas a high Km suggests relatively weak binding (often in the millimolar range).
Case Study: Alcohol Metabolism: The importance of Km is illustrated by aldehyde dehydrogenase (ALDH).
Most individuals possess a low Km (high affinity) form of ALDH that efficiently clears toxic acetaldehyde.
Some individuals carry a point mutation (E487K) resulting in a high Km (low affinity) enzyme. This leads to acetaldehyde build-up, causing symptoms such as facial flushing and tachycardia
Turnover Number and Catalytic Efficiency:
Beyond Vmax and Km, other parameters are used to assess how "good" an enzyme is.
Turnover Number (kcat): Also known as the molecular activity, kcat represents the number of substrate molecules converted to product per unit of time (usually s-1) per active site when the enzyme is saturated. It is calculated as: .
The Specificity Constant (kcat/Km): This parameter allows for the comparison of different enzymes or the same enzyme acting on different substrates. It is considered a better indicator of overall catalytic efficiency than kcat alone.
Upper Limits of Catalysis: The value of kcat/Km has a physical upper limit determined by the rate of diffusion of the substrate to the enzyme's active site.
Diffusion-Limited Reactions: In these highly efficient enzymes (e.g. carbonic anhydrase), every collision between enzyme and substrate results in catalysis.
The limit for such reactions is approximately 108 to 109M-1s-1.
Analysing Kinetic Data:
While modern techniques use non-linear fitting methods to determine Vmax and Km with confidence intervals, historical methods relied on linear transformations.
Lineweaver-Burk plots 1/V against 1/[S].
The linear transformation of the Michaelis-Menten takes the reciprocal of both sides of the equation:
Linear transformations distort the error on the data.
Reversible Inhibition of Enzymes:
Do not react covalently with the enzyme.
Show rapid equilibrium binding to the enzyme.
Give instantaneous inhibition.
Effect can be reversed by dilution.
The four classes of reversible inhibitors are:
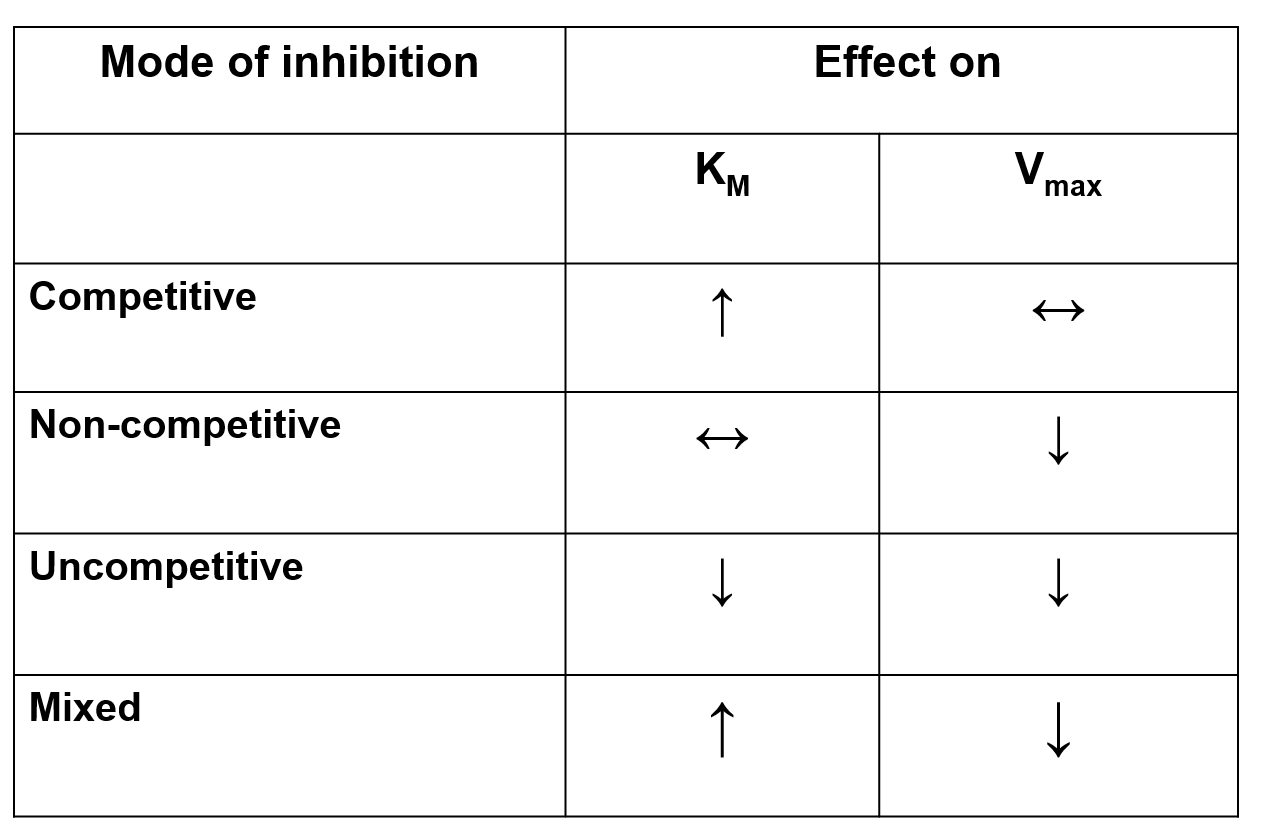
— — — — — — — — — — — —
Competitive Inhibition:
The inhibitor binds to the active site.
Enzyme can function at maximum velocity if the [S] is high enough.
I.e. the substrate can out-compete the effect of the inhibitor.
Vmax is unchanged.
— — — — — — — — — — — —
Non-Competitive Inhibition:
The inhibitor binds at a site other than the active site.
Binding changes the conformation of the enzyme which affects its catalytic activity.
Inhibition cannot be overcome by a high [S].
Km is unchanged.
The Active Site:
The active site is the region of an enzyme where substrates bind and catalysis occurs.
Favourable Binding Energy can offset Energetically Unfavourable Steps:
Binding energy - free energy released upon the interaction of a complementary enzyme and substrate.
It can be significant (e.g. substrate binding with chymotrypsin).
It must stabilise TS to increase the rate of reaction. If only the ES complex is stabilised, activation energy is increased and the rate constant decreases.
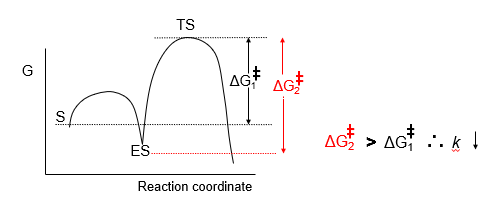
Transition State Binding is Optimised:
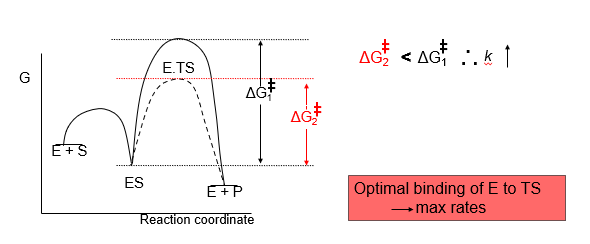
Utilisation of Binding Energy:
Strain - upon binding the substrate is distorted. TS makes better contacts than the substrate.
Induced Fit - structure of the enzyme is complementary to the TS only after binding. Active site closes around substrate to form a new environment for catalysis.
Serine Proteases:
They are a family of proteolytic enzymes that catalyse the hydrolysis of the peptide bond.
They facilitate nucleophilic attack at a normally unreactive carbonyl.
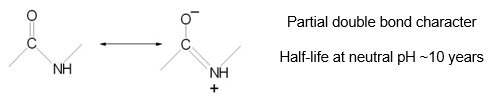
When treated with organophosphates, the serine protease chymotrypsin irreversibly loses all activity.
The mechanism involves a highly reactive serine in the active site.
The reactive serine is part of a ‘catalytic triad’ of residues.
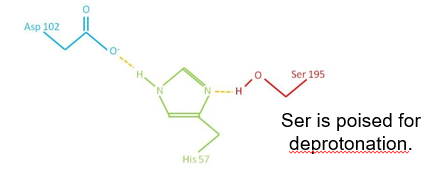
When a substrate is present, His- accepts a proton from Ser-.
Ser- attacks carbonyl carbon of peptide bond.
Forms an unstable tetrahedral intermediate (oxyanion) stabilised by residues from elsewhere in the active site.
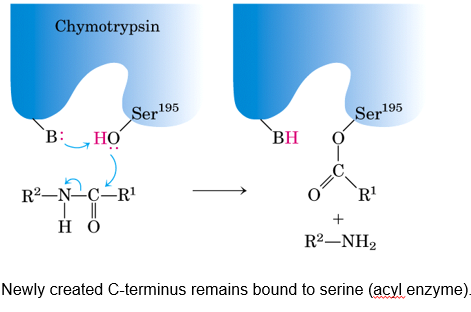
— — — — — — — — — — — —
Serine Proteases have different Specificities:
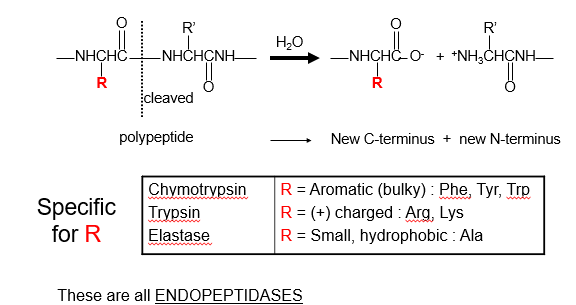
3 Regions of Active Site:
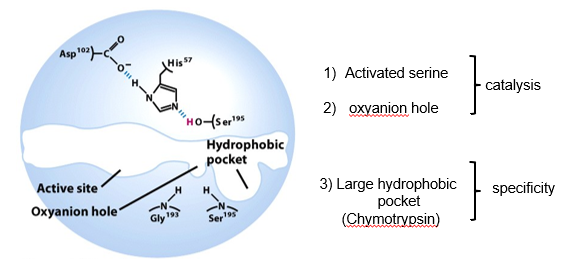
Chymotrypsin is activated by specific cleavage of a single peptide bond:
The inactive form ‘chymotrypsinogen’ is produced by the pancreas.
It is activated when the peptide bond between Arg-15 and Ile-16 is cleaved by trypsin.
Resulting π-chymotrypsin cleaves other π-chymotrypsin molecules to form α-chymotrypsin.
The 3 peptide chains produced are held together by two interchain disulfide bonds.
Type II Restriction Enzymes:
Restriction enzymes hydrolyse the phosphodiester bond in a 1-step mechanism,
Requires Mg2+.
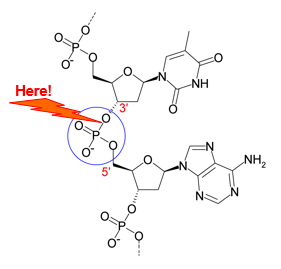
The requirement for Mg2+ stabilises the developing negative charge during hydrolysis.
Restriction enzymes recognise sequences with 2-fold symmetry (i.e. dimers).
In the absence of Mg2+, restriction enzymes have similar affinity for cognate and noncognate sequences.
They can scan along lengths of DNA to find cognate sequences.
❗ Specificity is achieved by distortion of cognate DNA.
Regulatory Strategies:
Proteolytic Activation
Allosteric Control
Covalent Modification (Phosphorylation)
Proteolytic Activation:
The enzyme is inactivate until activated by cleavage of one or more specific peptide bond.
The inactive form is a zymogen or proenzyme.
Activation is not reversible.
❗ Example: Trypsin, Chymotrypsin, Carboxypeptidase, Pepsin.
❗ Example: Blood Clotting Factors
Allosteric Control:
Activity is controlled by the binding of small molecules at regulatory sites distant to the active site.
They do not follow Michaelis-Menten kinetics, often forming a feedback loop.
They often perform feedback inhibition whereby a product produced late in a reaction pathway inhibits an enzyme that acts earlier in the pathway.
❗ Example: Aspartate Transcarbamoylase
It catalyses the 1st step of pyrimidine biosynthesis, and is inhibited by CTP (allosteric inhibitor).
Covalent Modification:
Regulatory groups are attached to the enzymes via a covalent bond.
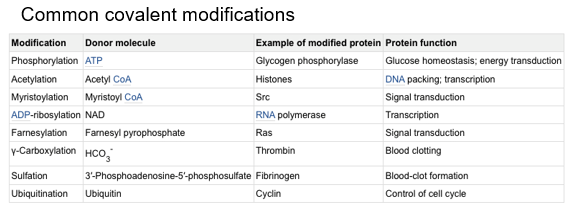
— — — — — — — — — — — —
Phosphorylation:
Protein kinases catalyse the transfer of phosphate groups to proteins.
ATP can act as a donor molecule, while a phosphate group is added to Ser/Thr or Tyr.
Phosphoryl groups add 2 negative charges and potential for 3 H-bonds.
Addition or removal of a phosphate group can cause conformational change within a protein or the formation of a new binding site.
Enzymes involved in metabolism are often regulated by phosphorylation and dephosphorylation.
❗ Example: Glycogen phosphorylase and Acetyl-CoA carboxylase
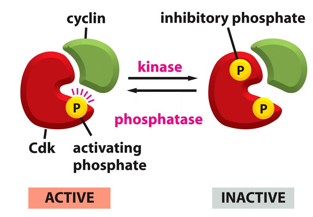
Cyclin Dependent Kinases (CdKs) Control Cell Cycle Timings:
CdK integrates 3 different inputs:
1. Kinase
2. Phosphatase
3. Cyclin
CdK relays a signal only when all three inputs are met.