Lab I
Lecture Video 1: Mitochondria DNA
Questions:
Do you have your well number & sample access code?
Yes
What size (bp) do we expect the PCR product to be?
16520 - 15972 + 1 = 549
What is a FASTA file?
A file format used for storing nucleotide sequences
What haplogroup do you think you are?
American
Drag the activities in the correct order
manually determine, annotate your mtDNA, create two phylogenies
In order to complete the Lab I activity, I will need the well number and sample access code my TA gave me in Lab E. If I don't have that information:
I need to come to one of the drop-in lab sessions in person to receive my code.
Notes:
Compare you mDNA to find out the haplogroup
Try to work on the worksheet and then go to drop in office hours
how to identify the haplogroup
have to annotate the sequence
cheek cell extraction
we ran the agarose
used the sanger sequencing to get the 4 color data
make a phylogenetic tree
We targeted the hypervariable sequence and amplified it with PCR
The PCR product size will be right primer start- left primer start + 1
Fasta File: file format used in bioinformatics
FASTA means fast
in the indicator of the header in line 1 we should add
haplogroup, ethinicity, region, country, sample ID
You can do a multiple sequence alignment
grouping based on similar sequences
Making a haplogroup tree
549 base pairs
Look at sequence (manually determine your haplogroup)
align it with the reference sequence to see where the mutations are
try to pinpoint the haplogroup
Use blast to find the haplogroup (annotate)
compare manual vs computed haplogroup
explore haplogroup information
make comparison between section haplogroups
Make two phylogenetic trees
align your mtDNA to make a tree
align your mtDNA with all of the other mtDNA sequences to make a class section tree
Lecture Videos Part 2
Questions
Which enzyme is commonly used in Sanger sequencing to perform DNA synthesis?
DNA polymerase
In Sanger sequencing: What is the purpose of incorporating dideoxynucleotides (ddNTPs), into the sequencing reaction?
To terminate DNA synthesis
Which oxygen atom(s) is/are missing in dideoxynucleotides (ddNTPs) compared to deoxynucleotides (dNTPs)?
Oxygen at the 3’ position only
What are the ingredients necessary for the Sanger Method of DNA sequencing?
dNTPs, DNA template, DNA polymerase, ddNTPs, primer
Why are we still using Sanger Sequencing in LS7L? (Select all that apply.)
Method is easy to explain
Only want to amplify small region
To protect anonymity
Notes:
Sanger Method
DNA sequencing method
In ddATP
we will take away the oxygen from the 3’ position which will stop elongation from happening
dNTPs
will cause the elongation to stop
will cause the
the sanger method is a slow progress
we have fast processes of DNA sequencing
will use the sanger sequence since we’re amplifying a small region of DNA
Lecture 3:
part 1:
questions
What is the password to access the MtDNA database?
science
What is 4-color data?
Automated DNA Sequencers generate a four-color chromatogram, which is the raw data of a sequencing run
Why is the beginning of a sequencing gel often occluded?
Primer dimers and solvent, which eluate first occulde the smaller DNA fragments.
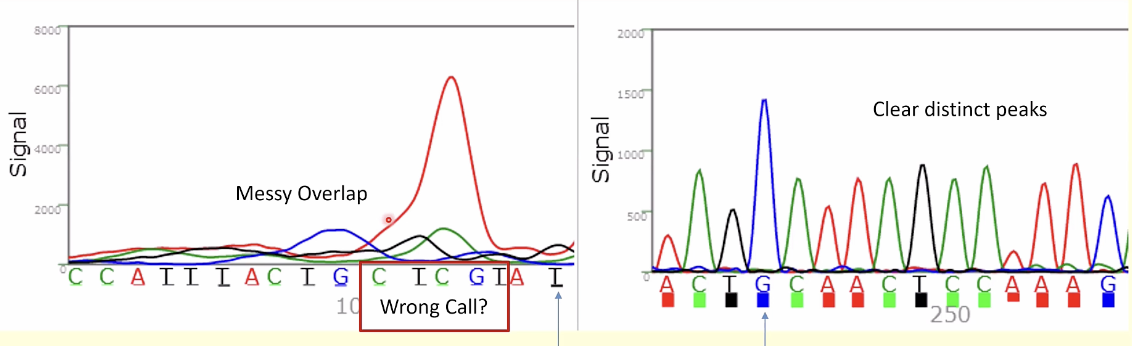
In the shown sequence the most likely reason for the change in data quality is:
c track in the target sequence, all cs will confuse the sequence since the sequence might slip
get a spare sequence if the BLAST fails
Based on what you learned in the lab manual chapter for Lab I and Appendix A, which of the following is an indication that you might need to use a spare sequence?
There are so many mutations that it is impossible to determine your haplogroup.
notes
ls7l, password : science
fill in the room info for the sample code in the database to access the four color data
you need to make a visual assessment of the sequence
there will be an indicator under the base → base calls
base calling: assigning nucleotides depending in the sequence based on the graph lapping
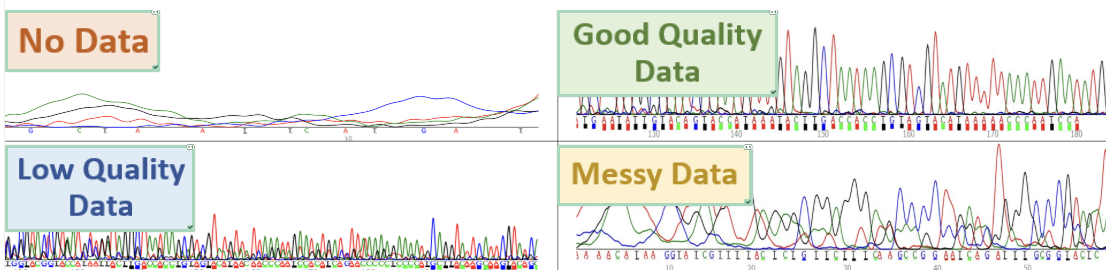
there are distinct and messy overlaps
at 20 bp there are primer dimers, these sequences are washed out by primers and solvent
at the end of the sequences that gaps are broader
ghost peaks, when its longer than 549, use blast
poor quality, use blast
slipage, use blast
part 2:
questions
What is the most likely purpose for aligning your mtDNA Sequence with the RSRS?
To identify SNPs in the mtDNA sequence to determine the haplogroup.
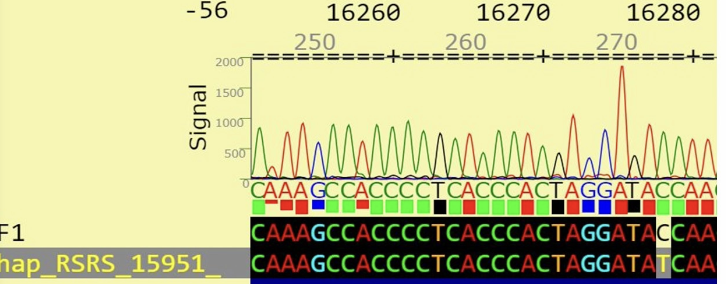
What is the correct mutation notation for the mutation pictured below?
T16278C
notes
Q4 on the worksheet
alignment helps idnetify the differences between personal mtDNA and the reference
help identify real mutations
once you do this you will identify all mutations in the different areas, top,middle, bottom
look at the phylogeny
part 3:
questions:
When annotating your mtDNA, what type of information should you include?
Your haplogroup, country of origin, and the length of your sequence
My mtDNA matches the mtDNA from an individual in the USA. That means?
My egg-parental lineage matches somebody living in the USA
If the multiple Sequence Alignment doesn’t work:
Check your Annotation Blast information field (there should be no DNA sequence)
try to delete shortest sequence
Try to restart with different set of sequences.
Try different browser (check marks misaligned)
notes:
questions 5,6,7,8
you will also need to include the reading at 260 and 280 on the annotation
by blasting you will retrieve sequences that are similar to your own
after determining the haplogroup you will need to find information from wikipedia
when making the tree deselect the outliers, such as the short sequences
Lab Manual:
We will be doing a comparison of our DNA to known sample to determine which egg parental lineage our DNA comes from
DNA synthesis is terminated through the addition of ddNTPs since it has a hydrogen in the 3’ position instead of a hydroxyl group
Being able to determine our mitochondrial lineage by comparing our sequences to the RSRS reference sequence and determining the mutations to see what our haplogroup is'
Assignment Overview:
Experimental Materials
Manually determine the haplogroup through alignment
Annotating mtDNA sequence
2 Trees