RNA
Nucleic Acids and Proteins
RNA Overview
Ribonucleic Acid (RNA): Polymer of nucleotides similar to DNA with key differences.
Sugar Component: Contains ribose (not deoxyribose as in DNA).
Nitrogen Base: Contains uracil (U) instead of thymine (T) found in DNA.
Strand Structure: Synthesized as a single strand, unlike the double helix structure of DNA.
Structural Features: Can fold upon itself, creating secondary structures important for function.
Pairing: RNA can pair with complementary DNA or other RNA strands to form double helices.
Types of RNAs
Various RNA Types and Functions:
Ribosomal RNA (rRNA): Major component of ribosomes.
Composed of a large and small subunit, which are both made of several proteins
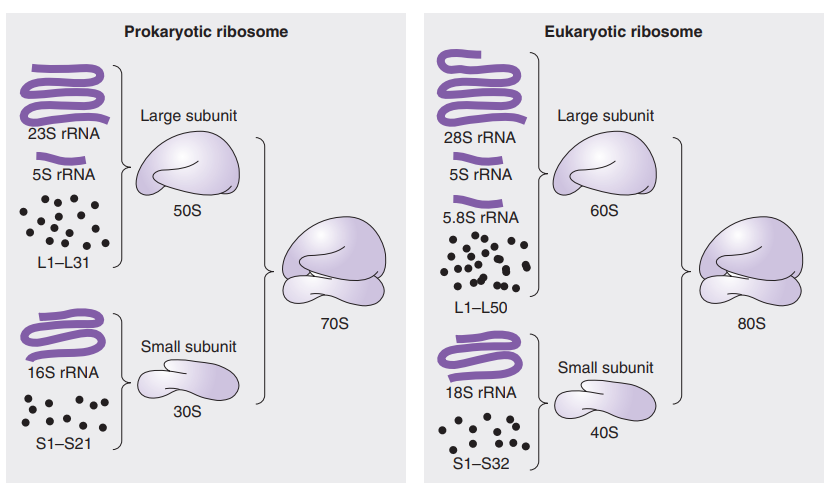
Messenger RNA (mRNA): Carries genetic information from DNA to ribosomes for protein synthesis.
Transfer RNA (tRNA): Brings amino acids to ribosomes during translation.
Small Nuclear RNAs (snRNA): Involved in RNA processing.
Function: Primarily involved in RNA processing, including splicing of pre-mRNA, where they play a crucial role in the removal of introns and the joining of exons to form mature mRNA.
Transcription
Process of Transcription: The copying of a specific DNA strand into RNA.
Enzyme Involved: RNA polymerase, which varies by organism.
Prokaryotes: One type of RNA polymerase.
Eukaryotes:
RNA Polymerase I (Pol I): Synthesizes ribosomal RNA (rRNA), which is a key component of ribosomes, essential for protein synthesis.
RNA Polymerase II (Pol II): Responsible for synthesizing messenger RNA (mRNA), which carries genetic information from DNA to ribosomes for protein production, and small nuclear RNAs (snRNAs) involved in RNA processing.
RNA Polymerase III (Pol III): Synthesizes transfer RNA (tRNA), which translates mRNA codons into amino acids during protein synthesis, along with some small nuclear RNAs.
Transcription Sites: Occurs at discrete stations within the nucleus, specifically at the promoter region of the gene.
Initiation of Transcription:
RNA polymerase and accessory proteins bind at promoter.
The promoter typically contains specific short sequences, such as the TATA box, which play a crucial role in transcription initiation.
The binding of RNA polymerase at the promoter is facilitated by transcription factors, which assist in the recruitment of the polymerase and help to unwind the DNA strands, exposing the template for RNA synthesis.
Eukaryotic transcription requires additional factors compared to prokaryotes.
Types of Transcription:
Constitutive Transcription: The continuous and unregulated transcription of certain genes that produce essential proteins required for basic cell functions
Inducible (Regulatory) Transcription:
Produced proteins only when needed. Activates transcription only during certain times in the cell cycle or particular conditions.
Transcription Elongation and Termination
Elongation: RNA polymerases synthesize RNA strand complementary to the DNA template.
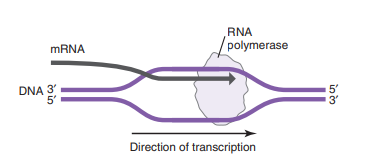
RNA polymerase moves along the DNA template, synthesizing the RNA strand by adding complementary ribonucleotides to the growing mRNA molecule.
This process involves the unwinding of the DNA double helix
RNA polymerase reads DNA in the 3' to 5' direction and synthesizes RNA in the 5' to 3' direction
Termination: Involves different mechanisms:
Prokaryotic: Can be rho-dependent or -independent, based on sequences in DNA.
Rho-Dependent: a protein involved in the termination of RNA synthesis in prokaryotic organisms
When Rho reaches RNA polymerase, it causes the complex to disassemble, resulting in the release of the newly synthesized RNA strand from the DNA template.
This process is crucial for preventing unnecessary transcription and ensuring that RNA synthesis stops at the correct location in the DNA sequence.
Rho-Independent:
Formation of Hairpin: While the RNA is being made, a section folds over to create a loop (like a hairpin).
U-Rich Sequence: Right after this loop, there is a long stretch of uracil (U) bases.
Weak Bonds Break: The bonds between the U's in the RNA and the A’s in the DNA are weak. When the hairpin forms, these weak bonds help the RNA separate from the DNA.Basically, the hairpin tells the RNA to stop, causing it to break away, and that’s how transcription ends!
Eukaryotic: involves cutting the RNA at the polyadenylation signal and adding a poly(A) tail, ensuring proper processing and stability of the mRNA.
In eukaryotic cells, termination of RNA synthesis occurs at a specific point called the polyadenylation signal.
The polyadenylation signal usually consists of the nucleotide sequence AAUAAA.
When RNA polymerase transcribes this signal, it indicates where to cut the RNA chain. An endonuclease will execute the cut
After the cut, a poly(A) tail, which is a long string of adenine nucleotides, is added to the RNA's 3' end.
This process does not rely on a specific termination sequence.
The cleavage event helps ensure proper processing of the RNA molecule.
Any extra RNA that is synthesized after the cut is quickly removed, ensuring only the correct RNA is used for protein production.
RNA Processing in Eukaryotes
Pre-mRNA Processing: Modifications that are done to RNA (mRNA) before it is considered functional and stable enough to be translated into proteins
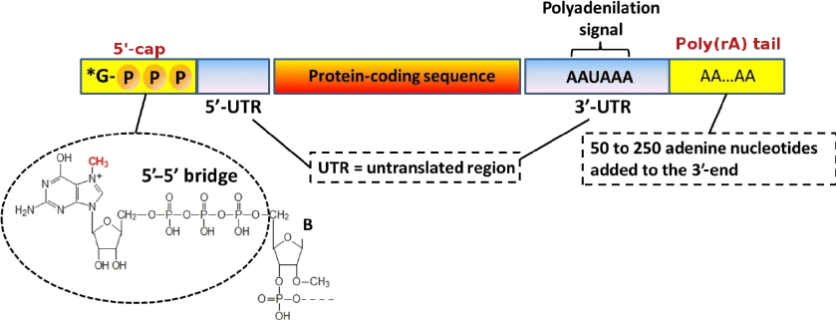
Capping: Addition of a methylated guanosine at the 5' end, providing stability and recognition for translation.
Polyadenylation: Addition of a polyA tail at the 3' end, critical for stability and translation efficiency.
Once RNA polymerase transcribes the polyadenylation signal, typically AAUAAA, the strand will be cleaved, and a poly(A) tail will be added to the 3’ end of the cleaved mRNA
Stability: The poly(A) tail protects the mRNA from degradation by exonucleases
Translation Efficiency: It also aids in the recruitment of the ribosome for translation, ensuring that the mRNA is efficiently translated into protein.
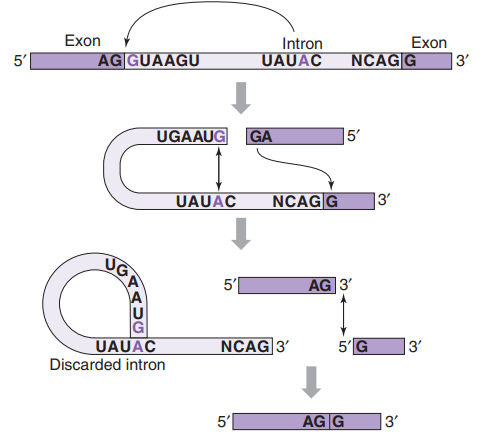
Splicing: Introns (non-coding regions) are removed and exons (coding regions) spliced together to form mature mRNA
Heteronuclear RNA (hnRNA) is the product of RNA synthesis. This is the form of RNA before the introns are removed.
Intron Removal: The spliceosome catalyzes two main reactions:
First, the 5' end of the intron is cleaved, and the intron is attached to the branch point (forming a lariat structure).
Second, the 3' splice site is cut, allowing the exons on either side of the intron to join together and release the intron in the form of a lariat structure, which is subsequently degraded.
Alternative Splicing:
In some cases, not all exons are included in the final mRNA, allowing for the production of multiple protein variants (isoforms) from a single gene. This process is known as alternative splicing and contributes to protein diversity and regulation in cells.
Functions of RNA
Ribosomal RNA (rRNA): Key structural and functional component of ribosomes, synthesized from DNA and critically involved in protein synthesis.
80-90% of RNA is rRNA
Messenger RNA (mRNA): Functions as a carrier of genetic instructions from DNA to the ribosomes, provides the information necessary for translation.
Monocistronic in eukaryotes - produce only one protein per mRNA strand
Although only one protein can be produced, by starting the RNA synthesis in different places or by processing the mRNA differently, diverse protein outcomes are possible (different type of protein)
Polycistronic in prokaryotes. - produce multiple proteins per mRNA strand
Transfer RNA (tRNA): Acts as an adaptor molecule, translating mRNA codons into amino acids for protein assembly.
tRNA is responsible for recruitment of amino acids. The anticodon on tRNA pairs with the codon of mRNA
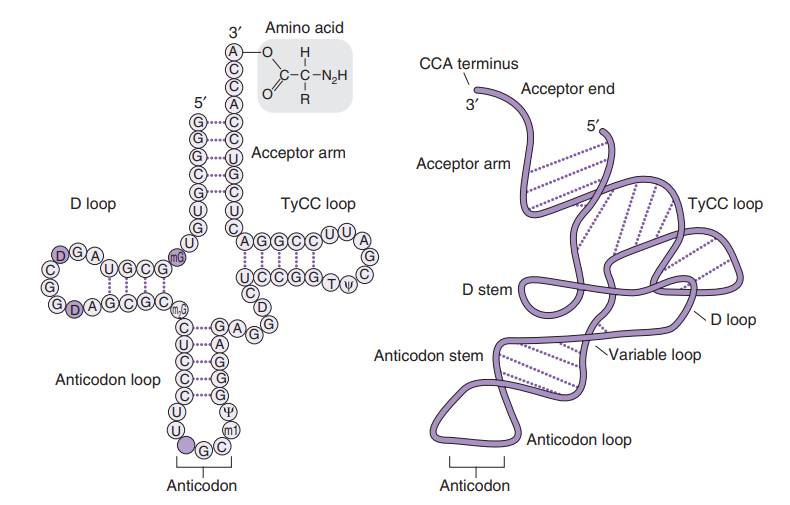
L-shape: tRNA molecules are typically described as having an L-shaped three-dimensional structure due to instrastrand hydrogen bonding
Anticodon: At one end of the tRNA is a region called the anticodon, which consists of three nucleotides that are complementary to a specific mRNA codon. This complementary pairing ensures the correct amino acid is delivered during translation.
Amino Acid Attachment Site: At the opposite end of the tRNA, there is an attachment site for a specific amino acid. Each tRNA is specific to one amino acid, allowing the genetic code to be translated accurately into proteins.
Protein Synthesis
Translation:
Starting Molecule: Initiated with methionine (AUG) in eukaryotes or N-formylmethionine in bacteria.
Ribosomes: Composed of rRNA and proteins; facilitate mRNA translation into proteins.
Codon-Anticodon Pairing: Essential for accurate translation, ensuring correct amino acid incorporation.
Termination: Signal by stop codons (UAA, UAG, UGA) leads to polypeptide release.
Additional Concepts
RNA Polymerases: Catalyze RNA synthesis; distinct for prokaryotic and eukaryotic systems.
DNA-dependent RNA polymerases - Require a DNA template
RNA-dependent RNA polymerases - Require a RNA template

Note that there are various versions of RNA polymerase in eukaryotes
2 alpha (α) subunits: Involved in the formation of the core enzyme and binding to regulatory proteins.
1 beta (β) subunit: Plays a crucial role in substrate binding and catalysis.
1 beta prime (β') subunit: Also involved in DNA binding and catalysis.
1 omega (ω) subunit: Helps in the assembly of the core enzyme.
1 sigma (σ) factor: Necessary for promoter recognition and the initiation of transcription. The sigma factor helps the holoenzyme bind to the promoter region of DNA, facilitating the start of RNA synthesis.
tRNA Charging: Activation of amino acids via tRNA by aminoacyl-tRNA synthetases, crucial for translation fidelity.
Chaperones: Assist in the proper folding of newly synthesized proteins, preventing aggregation.
Evolutionary Perspective: RNA is theorized to be the original genetic material from which DNA evolved.