Molecular Biology Techniques
Molecular Biology Techniques Exam 2
- Genomics * Gene Transfection: inserting intact genes * Eg. GFP linked to gene of interest * Editing genes: Adding/removing genes or single bases * CRISPR/Cas 9 (62 pig retroviruses removed) * Controlling gene expression * Knockin and knockout transgenic animals and cells * siRNA, shRNA, antisense RNA * DNA sequencing: determining the exact order of base pairs in a segment of DNA * “$1000 human genome” - Oxford Nanopore Technologies minION system uses bacterial pores (nanopores)
- Nanopore sequencing: drawing individual strands of DNA through protein pore in membrane, identifying unique shape of base pairs to obtain a readout of the DNA
- Transcriptome: the set of all the RNA molecules transcribed in a cell/cells
- Proteomics: the study of the entire genome
- Epigenetics: inheritable non-base changes in DNA due to environment * Potential epigenetic influences: died, disease, microbiome, psychological state
- Cancer cells and the metabolome * If we can understand metastatic cancer cells decision-making process (in how they spread in the body) then attacking them may be easier * Vanderbilt U researchers discovered metastatic cancer cells take the lowest energy route -> targeting metabolome may be important * Prefer large spaces (they are lazy)
- Secretome: secreted proteins, cfDNA, exosomes and vesicles from cells * biomarkers for cancer (cfDNA -> Galleri Test) * Stem cells secrete factors in exosomes that are important to tissue repair and other physiological changes -> clinical trials * Replace stem cells * Calm cytokine storm of covid 19 via inhalation * Advantage: stem cells are forever, exosomes have half life
- Mutations: change in DNA sequence that affects genetic information * Can be caused by UV light, chemicals … * Absolutely critical in molecular genetics
- Mutagen: a chemical or physical agent that interacts with DNA and causes a mutation
- Yeast cell characteristics * Can be diploid or haploid * Temperature sensitive mutants * Easy to grow -> simple basic solution
- Studying mutations using yeast cells process * Yeast in liquid culture -> add mutagen, distribute into smaller aliquots * Incubate at permissive temp (23 C) * Plate out individual aliquots * Grow one in permissive temp (23) and other in non growth temp (36)
- Permissive temperature: 23 C * Yeast will grow * The temperature at which a temp sensitive mutant allele is expressed the same as the wild type allele
- Nonpermissive temperature: 36 C * Temp sensitive mutants will not survive * The temp at which a TD mutant allele does not express itself
- What permissive temperature experiments tell us about involved genes * Temp sensitive genes are essential for cell growth (cell does not grow in a different environment that inhibits activity)
- Complementation analysis * Determines if recessive mutations are in the same gene * Test for determining whether two mutations associated with a specific phenotype represent two forms of the same gene (alleles)
- Complementation analysis procedure * Mate haploids of opposite mating types and carrying different temp sens cdc mutation * Plate and incubate one at permissive temp and replica at nonpermissive temp * Testing for TS cdc phenotype
- Interpretation of complementation analysis * Growth at nonpermissive temp: mutations are on different genes; respective wild-type alleles provide normal function * Absence of growth: mutants are on same gene; both allele nonfunctional
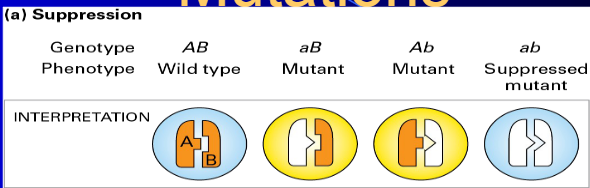
- Suppression - rare
* Suppressed mutant has wildtype
*
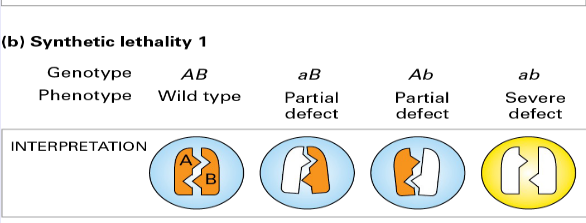
- Synthetic lethality 1
* Severe defect -> cannot bind
*
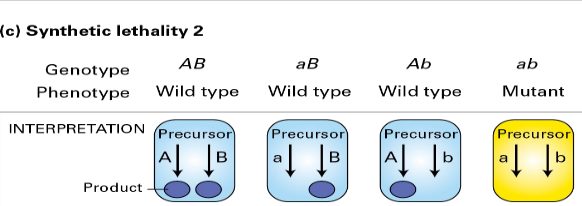
- Synthetic lethality 2
*
- Restriction enzymes/nucleases * Bacterial enzyme that cuts DNA at a specific sequence of nucleotides * Blunt and sticky ends * EcoRI
- Gene splicing: uses restrictive enzymes to insert DNA sequence into vector or plasma
- Nucleic acid hybridization: base pairing between a gene and a complementary sequence on another nucleic acid molecule * FISH, antisense DNA/RNA
- Nucleic acid hybridization procedure * Probe put into cell * Binds to select mRNA * Ribosome does not bind to doubly bonded mRNA
- DNA engineering: technology that involves manipulating DNA to insert in another organism
- Native gel electrophoresis types * Polyacrylamide GE: small sequence of DNA (10 - 200 bases) * Agarose GE: larger DNA (200 - 20 Kb)
- Native Gel electrophoresis applications * Restriction enzymes: band/gel shift assay (DNA binding proteins) * Apoptosis vs necrosis
- Band/gel shift assay * Objective: search for protein that binds to gene of interest
- Apoptosis vs necrosis native gel electrophoresis * Control: will not go anywhere * Apoptosis: ladder (nonrandom DNA cleavage) * Necrosis: smear (random DNA cleavage
- Northern Blot * Analyze RNA expression (gene expression) * Qualitative (molecular weight) and quantitative (relative abundance) * LIMIT: only one mRNA species can be assessed at a time
- Southern Blot * EG technique designed to detect gene sequence (DNA) * VNTR -> parental/forensic analysis * SNP -> helps to predict diseases * Personalized medicine * Bone marrow transplant example
- Southern Blot and bone marrow transplant * Expectation is for donar bands to match patients after surgery - no residual cancer * They now express the donor’s bands, and not their own
- VNTR (Variable Number of Tandem Repeats): distinguish DNa from multiple samples (forensic and parental analysis)
- SNP (Single Nucleotide Polymorphism): one base-pair variation in the genome sequence * Usually occur in junk DNA, bt can occur in coding DNA * Predict symptom of severity of autism spectrum disorder * Helps predict diseases, side effects, doses, construct genetic maps
- Molecular beacon probe * Gene expression -> look for specific DNA/RNA strand * Better than FISH * Molecular Beacon probe’s fluorescence is quenched (not fluorescent) unless bound to target RNA or DNA strand * Fluorescent dye on one end and quencher on other end of oligonucleotide * Only fluoresces when binds to DNA/RNA strand (in situ hybridization) because now quencher and fluorescent are separated
- cDNA microarray: analyze gene expression up to 8600 at once * Alternative to northern blots * Cluster analysis
- cDNA microarray procedure * Two groups: one without serum (control), one with serum (insulin example) * Isolate total mRNA of cells * Reverse transcriptase to cDNA * Label each with its own fluorochrome (red or green) * Mix to hybridize on array * Assess ratio of intensities of red, yellow, and green fluorescence
- Analysis of cDNA microarray * Green: expression of gene decreases in cell after serum is add -> control (no serum) binds * Red: expression of gene increases in cells after serum added -> insulin binds * Yellow: gene expressed in both cells -> both present
- cDNA microarray applications * What genes are expressed in x-type cells in response to Y? * Subtyping two cancer cells to identify differences in genes * Cryosurgery of prostate
- Cluster analysis: determining if groups of genes whose expressions are altered in response to the same conditions
- Single cell RNA sequencing: can identify individual cells at select points in types based on their unique RNA signatures * Improvement over cDNA microarrays
- Application of single-cell RNA sequencing: compare RNA signature of one cell to another (comparing artificial heart cells vs real heart cells)
- DNA cloning * Starting material is all genomic DNA * Adv. Gene of interest is present, * dis. huge library to search
- cDNA cloning * Starting material is mRNA and not DNA -> (smaller library) * Dis advantages * Need to convert mRNA to DNA to clone E.coli using reverse transcriptase * If gene is not being expressed, gene wont be cloned
- Issue with using prokaryotes to prompt DNA/protein synthesis * Very little protein of interest secreted * No post translational modification (glycosylation, phosphorylation, folding) * Required for protein to function
ENBREL: a miracle biologic for Rheumatoid arthritis
- PCR (Polymerase Chain Reaction): methods to amplify amounts of DNA or mRNA using Tac polymerase
- Short tandem repeats * Sections of a chromosome in which DNA sequences are repeated * Can be detected by DNA sequencing and PCR amplifying
The Jackson Laboratory in Bar Harbor Maine has over 6000 transgenic mice strains
- Knock-in genetic modification: Genetic engineering to activate a particular gene
- Knock-out genetic modification: genetic engineering to disable a particular gene
- siRNA Huntington’s disease: * No cure and causes breakdown of nerve cells in the brain * People with HD over over abundance of huntington's toxin (siRNA) which is toxic to nerve cells and cancer cells * People with HD have 80% fewer cases of cancer
- siRNA (short inhibitory RNA) * Adv. * Simple methodologies * Fast and effective transfection * Modifications are available * Disadv. * Not passed on to daughter cells * Non renewable * Only transient knockdown
- shRNA (short hairpin RNA) * Adv. * Renewable resource (passed down) * Transient or stable knockdown * Transfection or viral delivery * Disadv. * Technically challenging
\ \