Antibodies
4: Antibodies are glycoproteins only able to attach to a specific antigen. Produced by B cells and recognise proteins, carbs, nucleic acids, glycolipids and small inorganic molecules. Can mediate many responses by interacting with complement for cell lysis or attaching of innate cells, interacting with phagocytes via Fc region and have neutralising effects.
5: Discovery of antibodies took time. In the late 1800s von Behring and Kitasato realised serum from animals immunised against diphtheria/tetanus toxins could transfer immunity to others. In the early 1900s it was demonstrated that something in serum could neutralise and precipitate toxins, and lyse and agglutinate bacteria. It took until the 1930s to realise this was down to one molecule and in the 1950s the antibody structure was shown by Porter and Edelman.
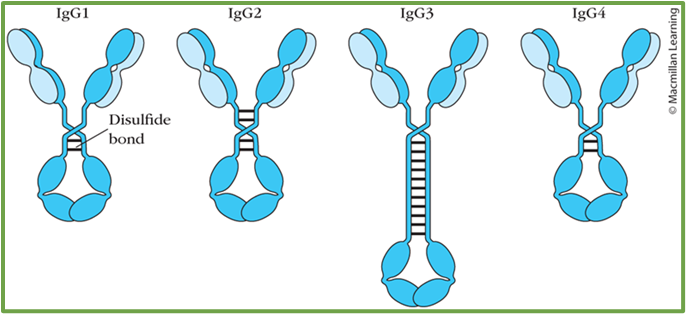
6: 5 antibody types distinguished by structure and function. IgG is broken down into a further 4 subtypes. LOOK AT 2ND YEAR SLIDES
IgG = most abundant in circulation, activates classical complement pathway, mediating ADCC by NK cells and opsonisation to enhance phagocytosis
IgD = role uncertain
IgE = works against parasite but also involved in type I hypersensitivity reactions (allergy)
IgA = dimer, activates lectin complement pathway, microbial adherance to mucosal surfaces and involved in inflammation
IgM = pentamer, first antibody to appear, undergoes class switching to other types, aids in removal of apoptotic cells
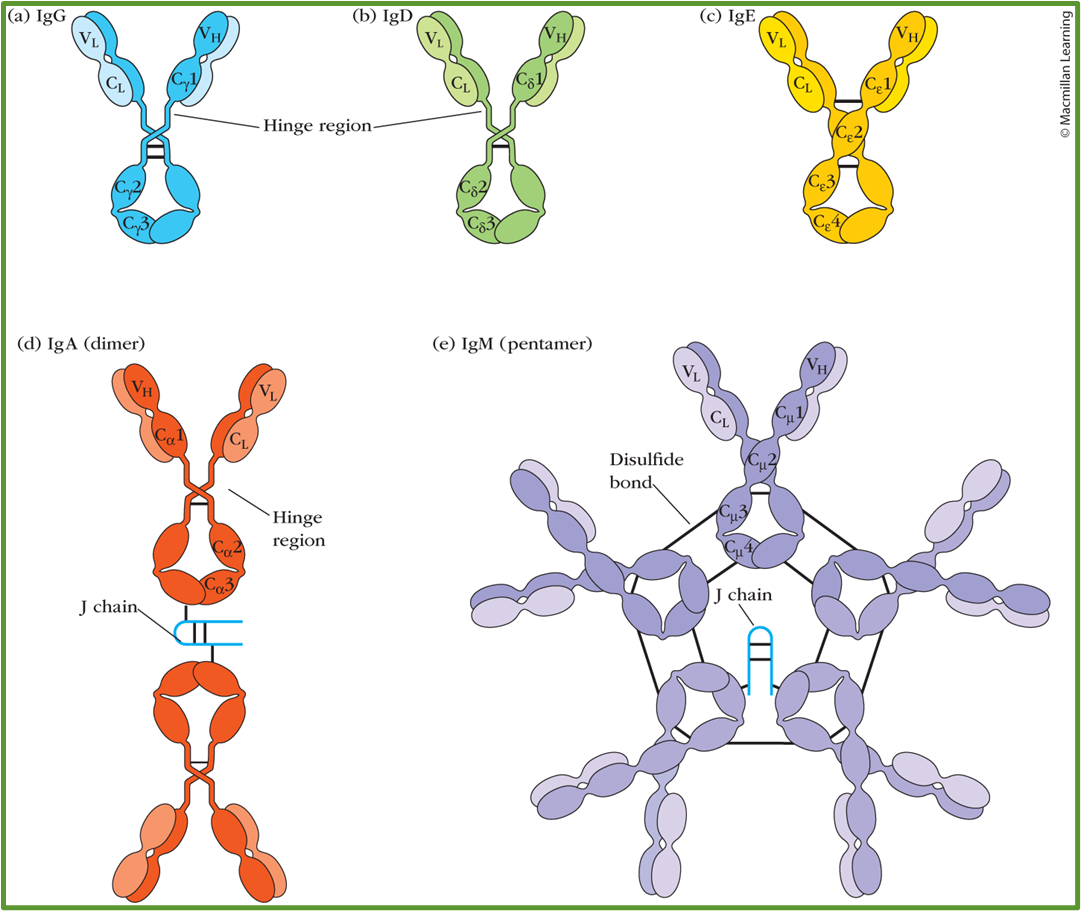
7: Antibodies consist of two identical light chains (25 kDa) and two identical heavy chains (53 kDa). The heavy chains determine which isotype an antibody is. The binding/variable region (Fab) is made up of both light and heavy chains, with two antigen sites per antibody. The Fc portion (heavy chain constant region) binds cell surface receptors. Variation found in types of light and heavy chain.
8:
9:
10: IgG molecules exhibit flexibility due to a hinge region between Fc and Fab regions which allows some independent movement of the Fab arms. This is done by Fab arms waving or rotating, the Fc tail ‘wagging’ or the Fab elbow bending. This is important for allowing IgG to simultaneously bind pathogens and immune system effector molecules/receptors.
Antigens made up of two homologous molecules which can crosslink two antigen-binding sites is used to create antigen-antibody complexes (bind multiple pathogens).
11: complementarity-determining regions (CDRs) = hypervariable (HV) regions
framework regions (FRs) = regions between CDRs
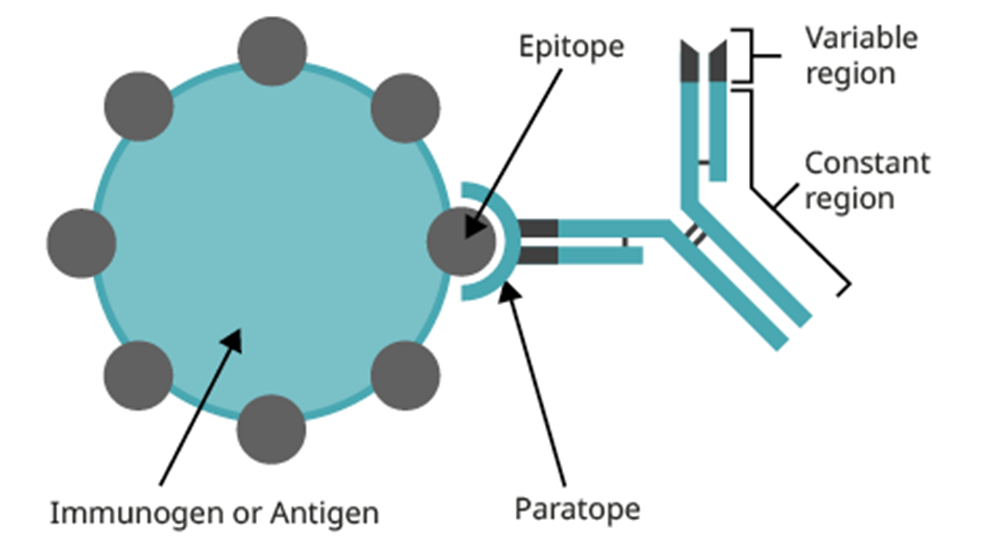
paratope = CDRs which make up antibody combining site
epitope = antigen combining site which the paratope is complementary to
12: Each antibody has a different V region, however sequence variability isn’t evenly distributed throughout the V region. There are three especially variable segments in the VH and VL domains - HV1, HV2 and HV3. HV = hypervariable. The most variable part is in HV3. Need 6 hypervariable regions to be different enough to bind individual specific pathogens.
The rest of the V domain (non-hypervariable regions) are less variable and known as framework regions, which form the beta sheets providing structure to the Ig.
13: Antibody domains are barrel-shaped and consist of polypeptide beta strands running antiparallel to each other, packing together to form two beta sheets held together by a disulphide bond.
In the folded light chain V domain, HV regions come together to form antigen-binding regions.
When the VH and VL domains pair in the antibody, the three HVs from each domain are brought together to form a single hypervariable site at the tip of each arm (antigen binding site) = 6 hypervariable loops/CDRs
14: Non covalent forces (electrostatic, hydrogen bonds, Van der Waals, hydrophobic interactions and cation-pi interactions) hold the antigen-antibody complex together.
Hydrophobic and van der Waals only work over very short distances and their purpose is to pull two complementary surfaces together. For some antigens, most of the binding energy is accounted for by hydrophobic interactions. In most antibody-antigen interactions many van der Waals forces form.
Electrostatic interactions occur between charged side chains, with at least one being found in most antibody-antigen interactions. Hydrogen bonds occur between hydrogen and oxygen/nitrogen. These allow more specific interactions and strengthen the overall interaction.
Partial charges shown in dipoles for H bonding and Van der Waals are shown as delta + or -.
15: There were two original theories for antibody diversity: germline and somatic variation. The germline theory suggested the genome contains a very large number of genes encoding Igs. Somatic variation theory suggested the genome contained a small number of Ig genes specificity is generated by alteration eg mutation or recombination.
Cloning antibody genes showed elements of both theories to be right. The DNA sequence encoding the variable region is generated by rearranging a relatively small number of inherited gene segments. Diversity is enhanced by somatic hypermutation in mature activated B cells.
In 1965, Dreyer and Bennett suggested two separate genes encoded a single heavy or light chain and in 1976, Tonegawa and Hozumi showed that separate genes encode the V and C regions of antibodies, and also showed the genes are rearranged during B cell development.
Antibody repertoire = total antibody specificities available to an individual. It’s at least 10^11 in humans and probably much greater than the size of our genome.
16: Different gene segments encode different parts of the antibody. There is many different possible gene segments for each segment of each chain, especially for the variable region.
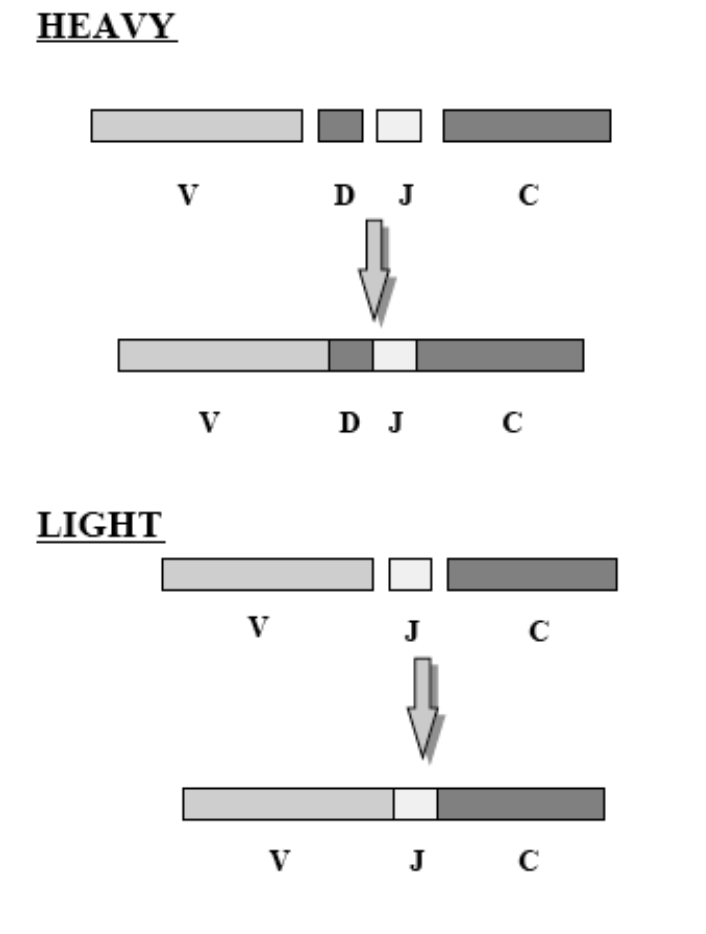
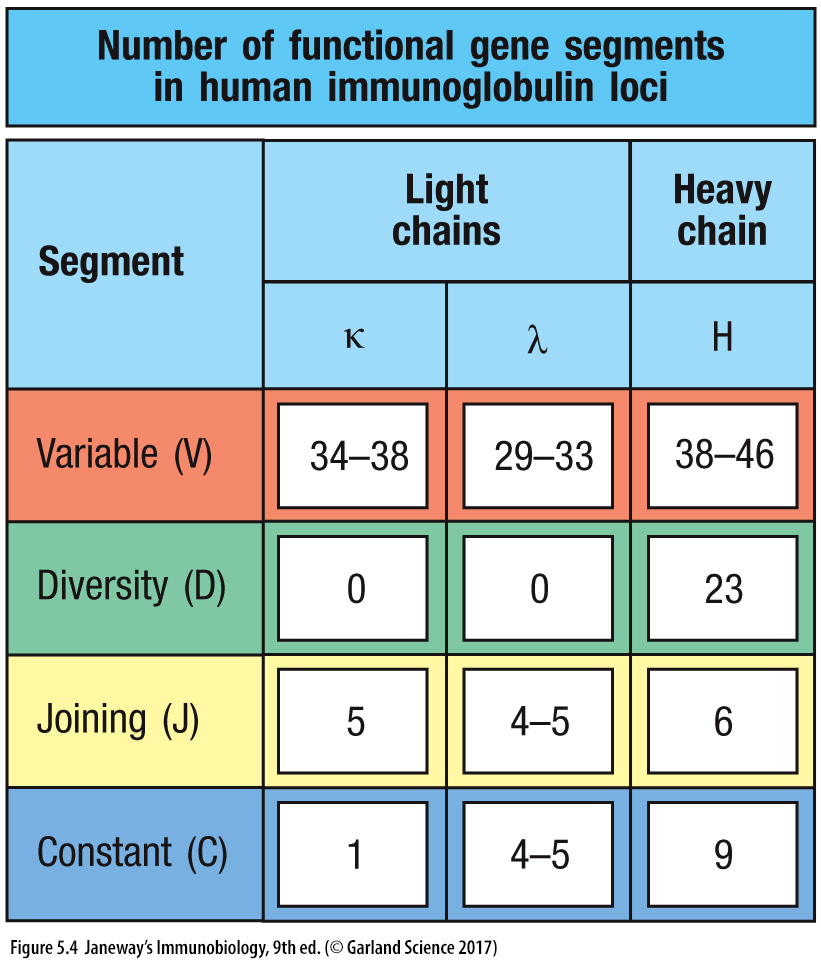
17: 30 different gene loci for variability region. Gene segments which encode the heavy and light chains are organised into three genetic loci - kappa, lambda and heavy chain loci. Each of these can assemble a complete V region sequence. Each locus is found on a different chromosome and organised slightly differently. Instead of the heavy chain locus having 1 C region, it contains multiple consecutive C regions, each corresponding to a different isotype. The total length of the heavy chain locus is more than 2 megabases (1 megabase = 1 million bases)
18: diagram, showing germline organisation of gene segments maybe - joining together of different regions and recombination - encoding variability
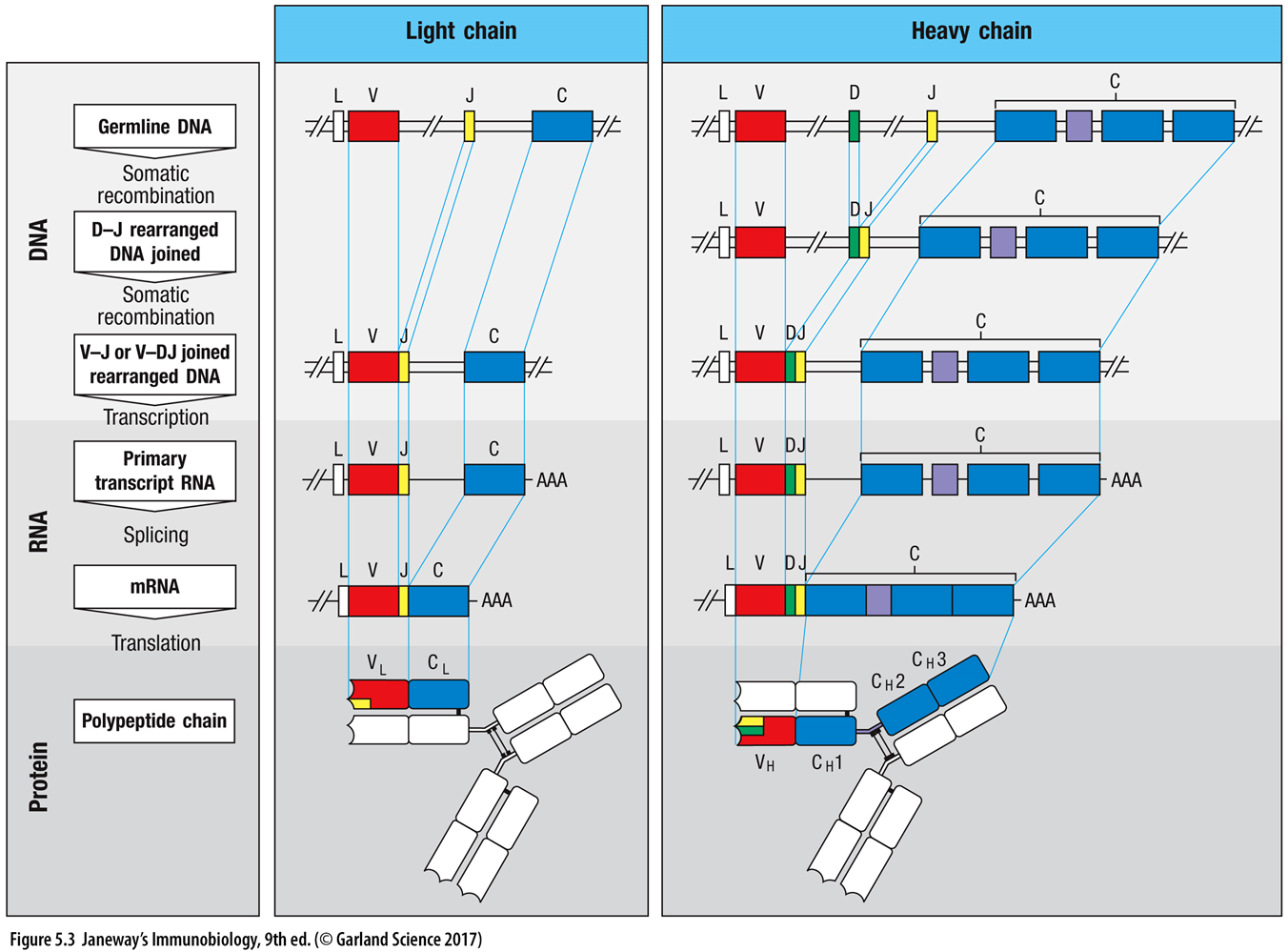
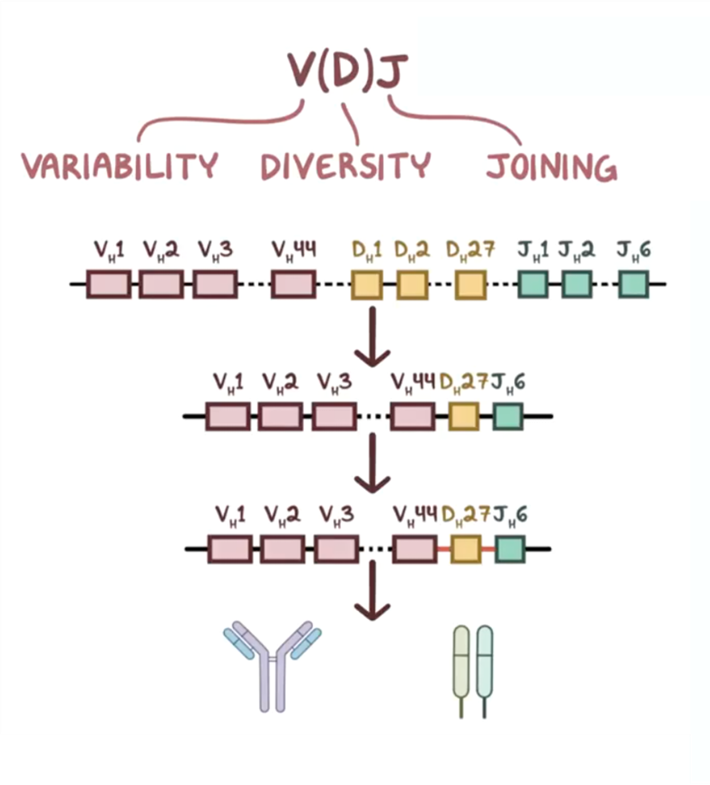
19: Antibody diversity is key to recognising a vast range of pathogens, and several mechanisms come together to create this diversity: V(D)J recombination, junctional diversity, somatic hypermutation, class switch recombination and combination of heavy and light chains.
V(D) J recombination is where the variable (V), diversity (D) and joining (J) segments recombine in developing B cells. Each person inherits multiple of these segments and they randomly recombine - heavy chains undergo V, D and J recombination whereas light (kappa and lambda) chains only recombine V and J segments. This shuffles the segments to create different combinations with purposeful recombinational inaccuracy to create additional variation and a wide range of antibodies. This process is mediated by RAG 1 /2 (recombination activating genes) enzymes which cut and join different segments.
Don’t have RAG = severe immune deficiency
20: Junctional diversity occurs during V(D)J recombination where RAG 1/ 2 and TdT (terminal deoxynucleotidyl transferase) enzymes add/remove nucleotides between V, D and J segments, altering recombination site sequences and creating unique AA sequences in antigen binding regions. Estimated that V(D)J recombination + junctional diversity could create at least 1011 different receptors expressed by naive B cells. Sometimes this produces an antibody which can’t bind - this antibody is destroyed.
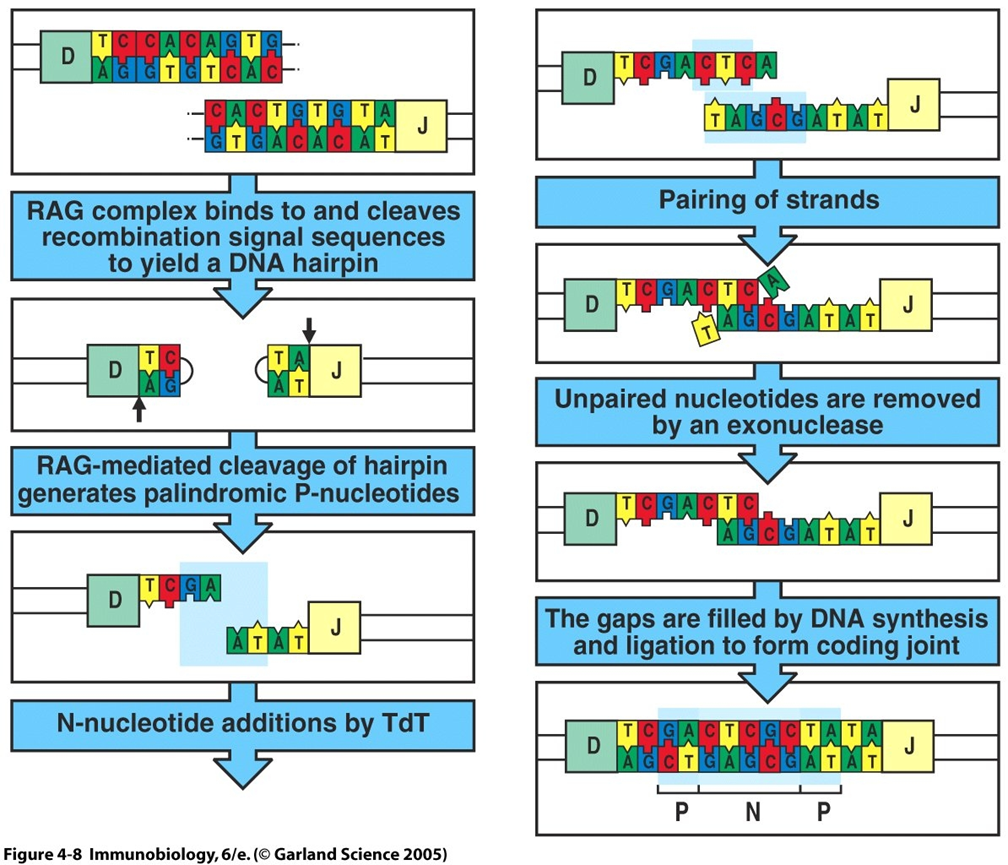
Above diagram shows junctional diversity.
RAG complex cleaves recombination signal sequences → DNA hairpin
hairpin cleavage by RAG → palindromic P-nucleotides
TdT adds N-nucleotides
strands pair
unpaired nucleotides removed by exonuclease
gaps filled by DNA synthesis and ligation → coding joint
What is a coding joint?
21: Somatic hypermutation occurs after a B cell encounters an antigen (in response to infection), rapidly introducing point mutations especially in antibody CDRs. The B cells producing antibodies with higher affinity to the antigen are selected, increasing specificity and diversity allowing enhanced antigen binding.
Overview:
antigen encounter
point mutations in CDRs
specific B cell selected for
point mutation = single base inserted, deleted or substituted
22: B cells can change constant region of the heavy chain - class switch recombination, as originally IgM is produced in response to every infection. This means the same antigen specificity (variable region) can be linked with different functions.
Random heavy and light chains may also be paired, increasing diversity as each combination recognises different antigens. This can be done in a lab to study therapeutics.
23: Antibody diversity is important for pathogen recognition, immune adaptability, targeting precision and vaccination.
Pathogen recognition: diversity allows millions of pathogens and toxins to be identified and neutralised
Immune adaptability: quick adaptation to new/evolving infections due to diverse antibodies
Targeting precision: specific, targeted immune responses with minimal damage to healthy tissues
Vaccination: antibody diversity allows vaccines to be effective through generation of specific antibodies with high affinity
Extra reading REQUIRED:
antigen binding site structure
mechanisms of antibody diversity