Protein Structure and Folding
Introduction to Protein Folding
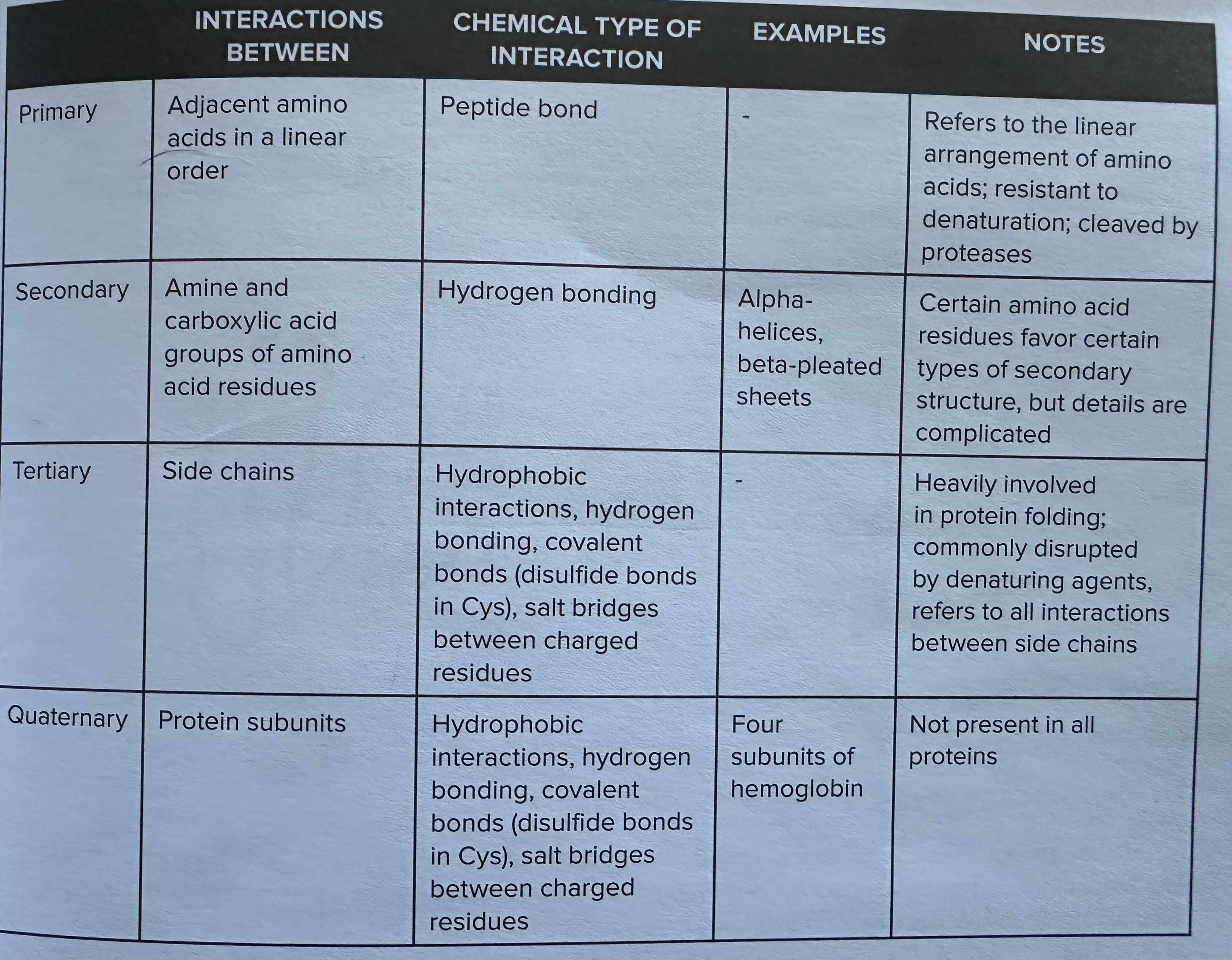
Three main types of protein folding
Primary Structure
linear chain of amino acids resides that make up the protein
covalent bonding = peptide bonding between residues
NOT disrupted by denaturation
proteases can disrupt primary structure
Secondary Structure
local structures that are formed by hydrogen bonding between the amino and carboxyl groups of amino acid residues
EX: beta pleated sheets and alpha helices are key examples of secondary structures that contribute to the overall stability and functionality of proteins.
IMPT TO NOTE: proline is particularly well know for for disrupting secondary structures = proline kinks due to its unique cyclic structure, which limits the flexibility of the polypeptide chain and can prevent the formation of regular alpha helices and beta sheets.
Tertiary
three dimensional structure that result from interactions among the side chains of the amino acids , including hydrogen bonds, ionic bonds, hydrophobic interactions, and van der Waals forces, ultimately determining the protein's shape and function.
charge driven interactions - salt bridges are formed between positively and negatively charged side chains, playing a crucial role in stabilizing the tertiary structure of proteins. EX; disulfide bondsare covalent bonds that form between the sulfur atoms of cysteine residues, providing additional stability to the protein's tertiary structure and often influencing its functional properties.
formations is through oxidation reaction
only form in TERIARY structures not secondary ones
Additionally, the hydrophobic effect contributes significantly to the tertiary structure, as nonpolar side chains tend to cluster together in the interior of the protein, away from the aqueous environment, further stabilizing the overall conformation.
Bonus: Quarternary Structure
Quaternary Structure: This level of protein structure involves the assembly of multiple polypeptide chains into a single functional unit, which can include various combinations of different subunits that interact through non-covalent bonds, contributing to the protein's functionality and stability.
Uses tertiary structures to do this - noncovlant interactions and disulfide bonds
Ex: hemoglobin
protein with 2 subunits = dimer, 2 = trimer, 4 = tetramer
A protein with more than four subunits is referred to as an oligomer, which can exhibit complex interactions and functionalities depending on the arrangement and nature of its constituent subunits. These oligomeric proteins can play critical roles in various biological processes, including enzyme activity and cellular signaling.
Types of Proteins
Globular Proteins:
Rounded and spherical in shape.
Typically serve functional roles (e.g., enzymes, hormones, regulatory molecules).
Generally water-soluble.
Fibrous Proteins:
Elongated and hydrophobic.
Primarily serve structural roles (e.g., keratin, collagen).
Membrane Proteins:
Attached to the plasma membrane.
Protein Folding and Thermodynamics
Entropy:
Fundamental concept: increases in entropy are thermodynamically favorable.
In a physical context, entropy correlates with disorder.
Unfolded vs. Folded Proteins:
An unfolded protein is a linear chain with high entropy (more flexibility and disorder).
A folded protein is rigid with low entropy (less flexibility due to interactions).
Why Proteins Fold:
Folding decreases the entropy of the protein but compensates by increasing solvent entropy.
The interaction between non-polar (hydrophobic) and polar (hydrophilic) residues drive this process.
Hydrophobic Interactions
Hydrophobic residues cluster to minimize contact with water, leading to:
Polar residues positioned on the surface to interact with the solvent.
Non-polar residues buried in the interior of the protein structure.
This arrangement reduces the need to form ordered solvation shells around non-polar residues, thus increasing solvent entropy.
Simple Takeaways on Protein Folding
Hydrophobic clustering: Hydrophobic molecules gather away from hydrophilic molecules, shaping protein structure.
Residue placement: Polar residues (e.g., aspartic acid, tyrosine) are usually found on the exterior, while non-polar residues (e.g., valine, tryptophan) are typically located in the interior.
Protein Denaturation
Denaturation refers to the loss of a protein's secondary, tertiary, or quaternary structure while preserving the primary structure.
Key Points about Denaturation:
Primary structure remains intact, as it involves covalent peptide bonds
Non-primary structures are held by weaker interactions (ionic, hydrogen bonds).
Denaturing Agents
Factors that lead to denaturation include:
Temperature extremes: High temperatures disrupt non-primary interactions, rendering proteins non-functional. Low temperatures can reduce activity due to decreased molecular collisions.
pH extremes: High or low pH can alter charge-based interactions (salt bridges) crucial for tertiary structure stability.
Detergents: Amphipathic compounds disrupt hydrophobic interactions that maintain folding, with SDS being a notable example in labs.
Reducing Agents: Break disulfide bonds, reducing interactions between cysteine residues (e.g., 2-mercaptoethanol).
Recovery from Denaturation
Some denaturing conditions are reversible; proteins can regain their structure once the conditions are normalized (e.g., removal of urea).
Irreversible Denaturation: Some methods (like cooking) permanently alter structure without a possibility of refolding.
Proteases and Protein Breakdown
Enzymes called proteases break down proteins by hydrolyzing peptide bonds, leading to amino acid release.
Categorization of Proteases:
Based on catalytic mechanisms or target spchain ecificity (e.g., serine proteases cleave near specific amino acids).
Examples include trypsin, which cleaves at the C-terminal side of lysine and arginine residues, and aminopeptidases/carboxypeptidases, which remove residues from terminal ends.
Important Notes
defining feature of each level of structure is the interaction BETWEEN the various structures
EX: hydrogen bonding
occurs in 2, 3, and 4
Difference is the key:
2nd: occurs between the amine
and Carboxyl residues in the AMINO ACID chain itself
3rd: occurs between varoius side chains of the protein
4th: between the protein subunits