Genetics - lecture 12 - Gene Mutations
Understanding Mutations
Mutation
Defined as a change in the DNA sequence.
Caused by DNA replication errors, spontaneous mutations, and chemicals and irradiation.
Present in every organism; not always manifesting as distinct phenotypes.
New mutations can occur every generation.
Types of mutations include gene mutations, point mutations.
Types of Mutations
Gene Mutation
Occurs within individual genes from changes in nucleotide sequences, affecting protein structure and gene expression.
Categories of Gene Mutation:
Somatic Mutations:
Occur in non-germline cells (body cells).
Non-inheritable.
Germ-Line Mutations:
Occur in gametes (egg and sperm).
Inheritable.
Can lead to cancer.
Point Mutations
Base-Pair Substitution Mutations:
Changes one nucleotide base pair for another.
Silent Mutation:
No change in amino acid due to genetic code redundancy.
Missense Mutation:
Results in an amino acid change; can be:
Conservative: changes to a chemically similar amino acid, less likely to alter function (e.g., alanine to glycine).
Non-conservative: changes to chemically different amino acid, more likely to alter protein function (e.g., glycine to glutamate).
Nonsense Mutation:
Changes an amino acid codon to a stop codon, resulting in truncated, often nonfunctional polypeptides.
Frameshift Mutation:
Caused by insertions or deletions of bases, altering the reading frame and affecting downstream sequences.
Transcriptional Mutations
Mutations affecting gene expression without changing amino acid sequences.
May occur in regions such as promoter sites or regulatory protein binding sites, affecting transcription levels.
Regulatory mutations: Alter the amount of protein product produced by a gene.
Promotor mutations: Alter the binding sites of transcription and regulatoy proteins
Infere with efficient transcription initiation.
Could reduce, abolish, or enhance transcription.
Causes of Mutations
Induced Mutations:
Caused by environmental mutagens, such as chemicals, radiation (e.g., UV light).
Mutagenesis: Process of inducing mutations by mutagens.
Types include:
Nucleotide base analogs (altered pairing properties)
Deaminating agents (remove amino groups).
Alkylating agents (add methyl/ethyl groups).
Hydroxylating agents (add hydroxyl group)
Oxidative agents (addition of oxygen or removal of hydrogen)
Intercalating agents (insert between bases).
Flat, planer molecules intercalate between base pairs distorting the DNA helix which disrupts DNA replication and causes addition/deletion of nucleotides leading to frameshift mutations.
Spontaneous Mutations:
Arise in absence of known mutagens, often from DNA replication errors or tatomeric shifts altering base pairing.
i.e. DNA polymerase can inset the wrong nucleotide which is corrected by proof reading 99% of the time
Trinucleotide repeat disorders: Caused by DNA polymerase slipping during DNA replication and increasing the number of trinucleotide repeats within a gene which results in longe stretches of the same AA within the protein.
Nucleotide base changes: DNA nucleotide bases can occasionally convert to alternative structions (tautomers) with slight differences in bonding and hydrogen places.
amino (A+C) (C-NH2) → imino
keto (T + G) (C=O) → enol
Leads to base-pair mismatches and incorporation of incorrect bases.
Transposable Elements
DNA sequences that move within a genome.
Two types:
DNA transposons: Move position using protein transposase found in both prokaryote and eukayote genomes.
Retrotransposons: Move position via RNA intermediate and reverse transcriptase - found in eukaryotes.
Have the ability to cause mutations on insertion into genes.
DNA transposons - IS elements in E. Coli
Each genome contains several IS elements.
F plasmid contains three IS elements.
They carry a transposase gene bracketed by inverted repeat (IR) sequence.
Transposase: The protein involved in moving the IS element from one DNA location to another
Steps:
Transposase molecule binds to each IR and blunt end cuts on both sides of the IS elements
Then it cuts the new target site in the genome with staggered cuts
If a transposon moves and inserts into a gene, it will create an insertion mutation that will cause premature termination.
DNA Repair Mechanisms
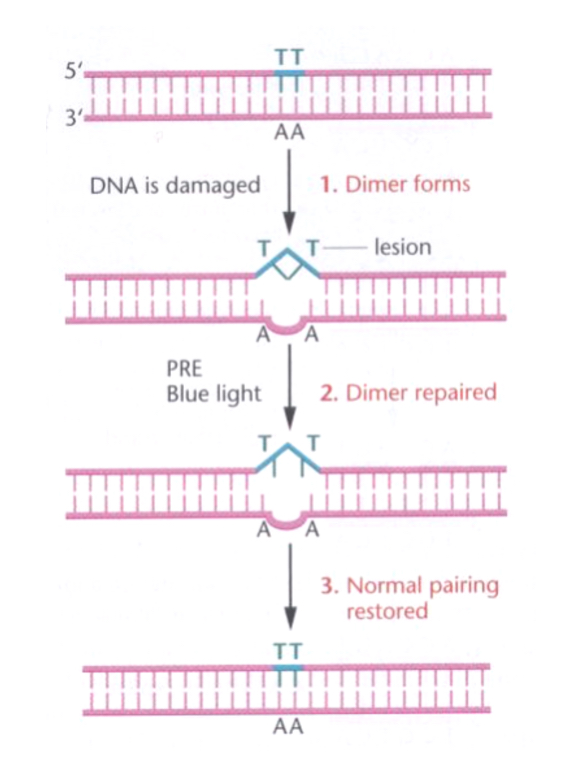
Photoreactivation Repair:
Photoreactivation enzyme (PRE) repairs UV-induced thymine dimers through cleavage (e.g., photolyase in E. coli).
Called photolyase in E. Coli
Not present in humans
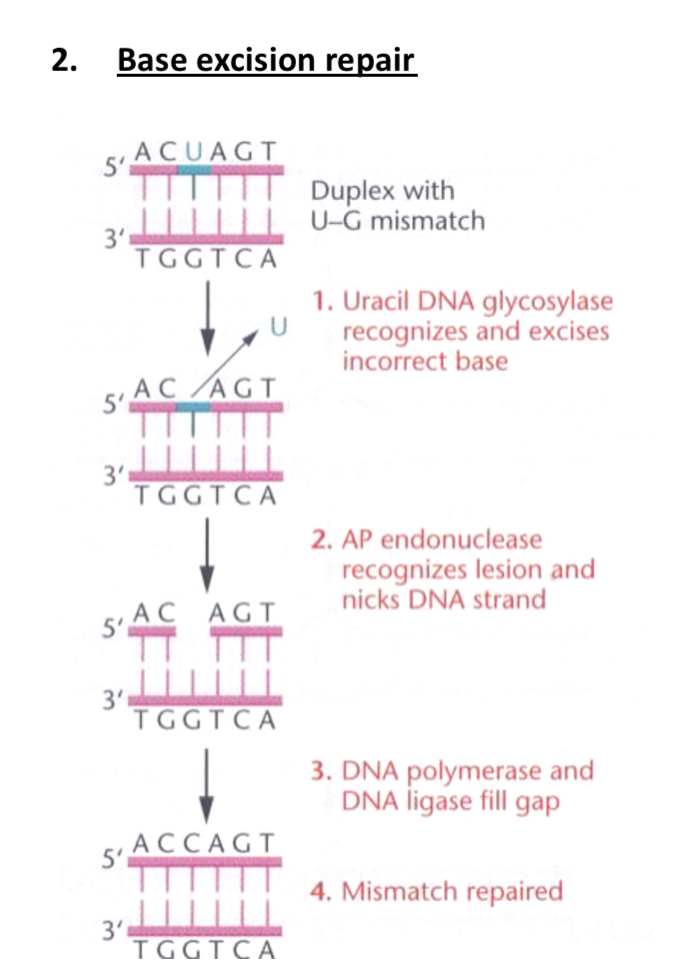
Base Excision Repair:
Performed by a base specific glycosylases, AP ligases and polymerases to correct mismatched bases.
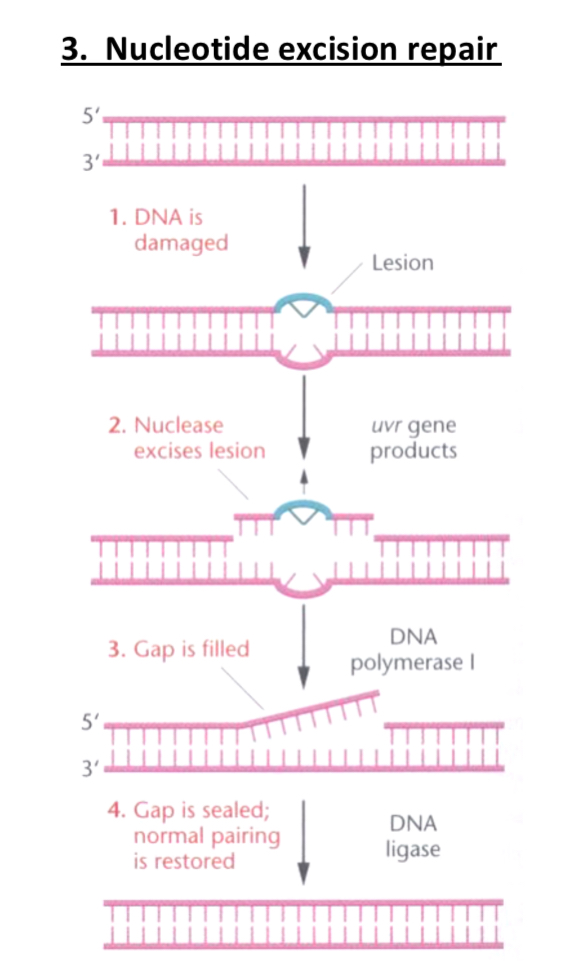
Nucleotide Excision Repair:
Recognizes and repairs bulky DNA lesions, utilizes enzymes (e.g., UvrA, UvrB, UvrC in E. coli) to excise damaged sections.
DNA polymerase I synthesizes the new DNA strand and DNA ligase joins the single standed nick.
Human Diseases Caused by Mutation
Xeroderma Pigmentosum:
Caused by mutations in genes involved in nucleotide excision repair, leading to extreme photosensitivity and skin cancer.
Hereditary Cancers:
Linked to mutations in DNA repair genes, such as BRCA2 for breast cancer, MSH2 for colorectal cancer.