AP Bio Notes
Unit 1: Chemical Foundation of Life
Chemistry Review:
Chemical bonding: Atoms share electrons by bonding with each other.
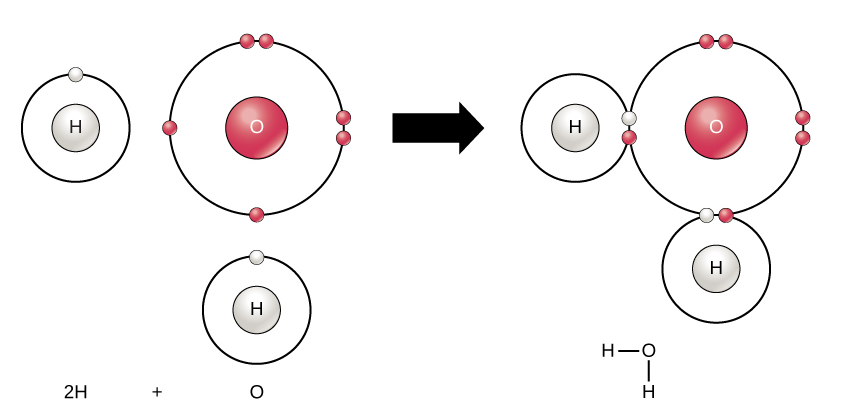
Ion: An atom with a charge, any atom with an uneven amount of electrons and protons.
Cation: Ion with a positive charge by losing electrons
Anion: Negatively charged ion by gaining ions
Electronegativity: (Determined by how many electrons are in the valence shell) Measurement of how much they will pull electrons toward themselves and the closer an electron is to eight electrons the more electronegative that element is.
Most Electronegative element is Fluorine
2nd most electronegative element is Oxygen
3rd most electronegative element is Nitrogen
Electropositivity: Ability of elements to donate electrons and form positive ions
Polar Molecules: Occur when there is an unequal sharing of electrons across a covalent bond meaning that electropositive elements who donate and electronegative elements who take are bonded and that electron is shared more with the electronegative element which is selfish.
Types of Bonds:
Ionic Bond: Formed between ions with opposite charges
Covalent Bonds: Stronger bonds than ionic bonds made with sharing electrons between atoms.
Non-Polar Covalent Bonds: Bonds with an electronegativity difference of less than 0.4
Polar Covalent Bonds: Bonds with an electronegativity difference of more than 0.4
Hydrogen Bond: individually weak bonds created by the attraction of an electronegative element in a molecule with a electropositive element in another molecule
Properties of Water:
Water & Hydrogen Bonding: Water is polar, therefore has uneven amount of charges, and this allows the electropositive end to attract to the electronegative end of water molecules forming hydrogen bonds. Hydrogen bonds are the reason for the properties of water.
Properties:
Cohesion: Water molecules attraction towards each other
Results in Surface Tension by which the cohesion of water makes the surface resistant to rupture by light objects
Adhesion: Water molecules attraction towards other polar molecules
Results in Capillary Action, by which water sticks to side of tubes, or plant roots, and goes up the sides.
High Heat Capacity: Waters resistance to temperature change.
Used by organisms to maintain homeostasis in the body
Takes a lot of energy to break the many hydrogen bonds in water
Density as a Solid: Water is less dense as a solid than liquid.
Creates ice sheets as a insulating barrier to protect life under the sheets from freezing
Solvency: What is a excellent solvent; dissolves other chemicals.
Very good at pulling ionic compunds like salt apart into ions
Biomolecules:
Biomolecule: Large or small molecule used by living organisms; an umbrella term
Macromolecule: Large molecules used by living organisms such as:
Carbohydrate
Lipid
Protein
Nucleic Acid
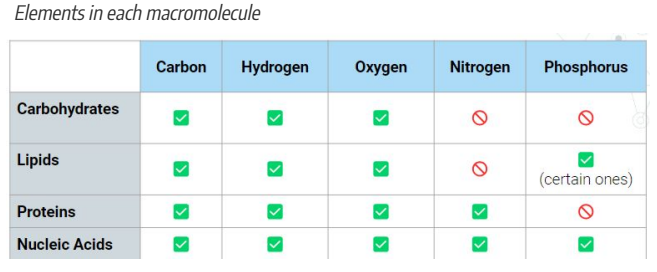
Elements of Macromolecules:
Carbon
Hydrogen
Oxygen
Nitrogen
Phosphorous
Sulfur
Biomolecule Metabolism
Metabolism: The combination of chemical reactions that synthesize and hydrolyze biomolecules for energy storage and release in an organism.
Catabolism: Creates energy
Breaks down biological polymers into monomers to help generate ATP
Anabolism: Synthesizer for storage of energy
Build up monomers into polymers
Dehydration Synthesis: monomers are covalently bonded together into polymers.
Anabolic process
Removes water to build larger molecule
Water is formed as a byproduct outside
Requires the assistance of enzymes to occur
Hydrolysis: Polymers are broken down into monnomers.
Catabolic
Add water to break down molecule
Requires the assistance of enzymes to occur
4 Types of Biomolecules
Carbohydrates: known as sugars
Monomers are monosaccharides
Such as Glucose
Polymers are polysaccharides
Such as Starch
Function:
Short term energy source
Energy storage
Provide structure in organisms
Such as cellulose in plants
Structure is hexamer rings
Always in a ratio of 1Carbon:2Hydrogen:1Oxygen
Types of Carbs:
Branched Structure used to break multiple monomers at once to allow for faster ATP generation -
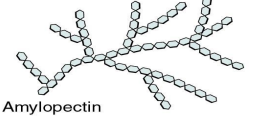
Stacked structure giving organisms structural support -
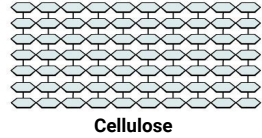
Lipids: known as fats or oils
Monomers are fatty acids
Polymers are lipids
They are hydrophobic, water avoidant
Function:
Long term energy storage
Insulation and protection of organs
Structure is long hydrocarbon chains -
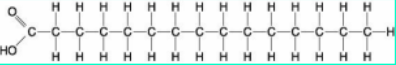
Ratio of 1Carbon:2Hydrogen: Low oxygen
Types of Lipids:
Saturated fats with no double bonds in their structure like the example above
These fats are linear and can stack to creat dolids at room temp which leads to build up in blood vessels
Unsaturated fats with double bonds
Stays liquid at room temp
Proteins: found in meats and muscles
Monomers are amino acids
Polymers are polypeptides
Function:
Wounds and tissue repair
Enzymes
Cell Signaling
Structure is very complex with four levels
Elemental Composition:
Carbon, Hydrogen, Oxygen, Nitrogen and Sulfur
Known as CHONS for short
There are many different types of proteins form proteins embeed into cell membrane to enzymes thtat help catalyze chemical reactions
Nucleic Acid: known as genetic material
Monomers are nucleotides with 3 parts
5-carbon sugar (ribose or deoxyribose
Phosphate
Nitrogenous Base with pairings of A T G C or U
All linked by covalent bonds
Polymers are nucleic acids
Function:
Storage of genetic material
Elemental Composition:
Carbon, Hydrogen, Oxygen, Nitrogen and Phosphorous
CHONP for short
Types of Nucleic Acids:
DNA which is deoxyribonucleic acid
Double helix
Stores genetic code
4 nitrogen bases: ATGC
More stable than RNA
Found in Nucleus
RNA which is ribonucleic acid
Single Stranded
3 types
Messanger mRNA
Transfer tRNA
Riobosomal rRNA
Used for protein synthesis
Instead of Thymine as a nitrogen base it is Uracil
Less stable than DNA
Made in nucleus and transported to cytoplasm
Protein Structure
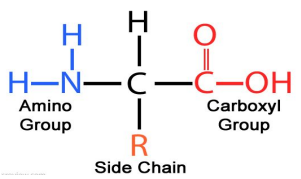
Amino Acid Structure: Bonded to one carbon
Amine group
Indicated by bonded elements of NH2 or NH3
Carboxyl Group
Indicated by bonded elemnts of COOH or COO-
R Side Chain
Varies between the 20 amino acids
Determines if amino acid is hydrophobic, hydrophilic, acidic, or basic
One hydrogen bond to complete 4 covalent bonds of carbon
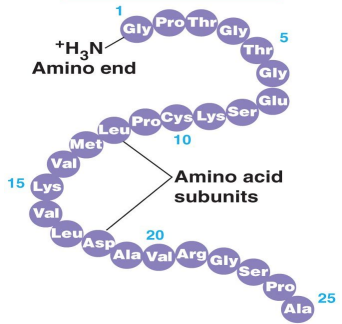
Primary Protein Structure
Peptide Bond: Bond created between two amino acids
Uses dehyration synthesis
Connects the amine group of one amino acid and the carboxyl group of another amino acid
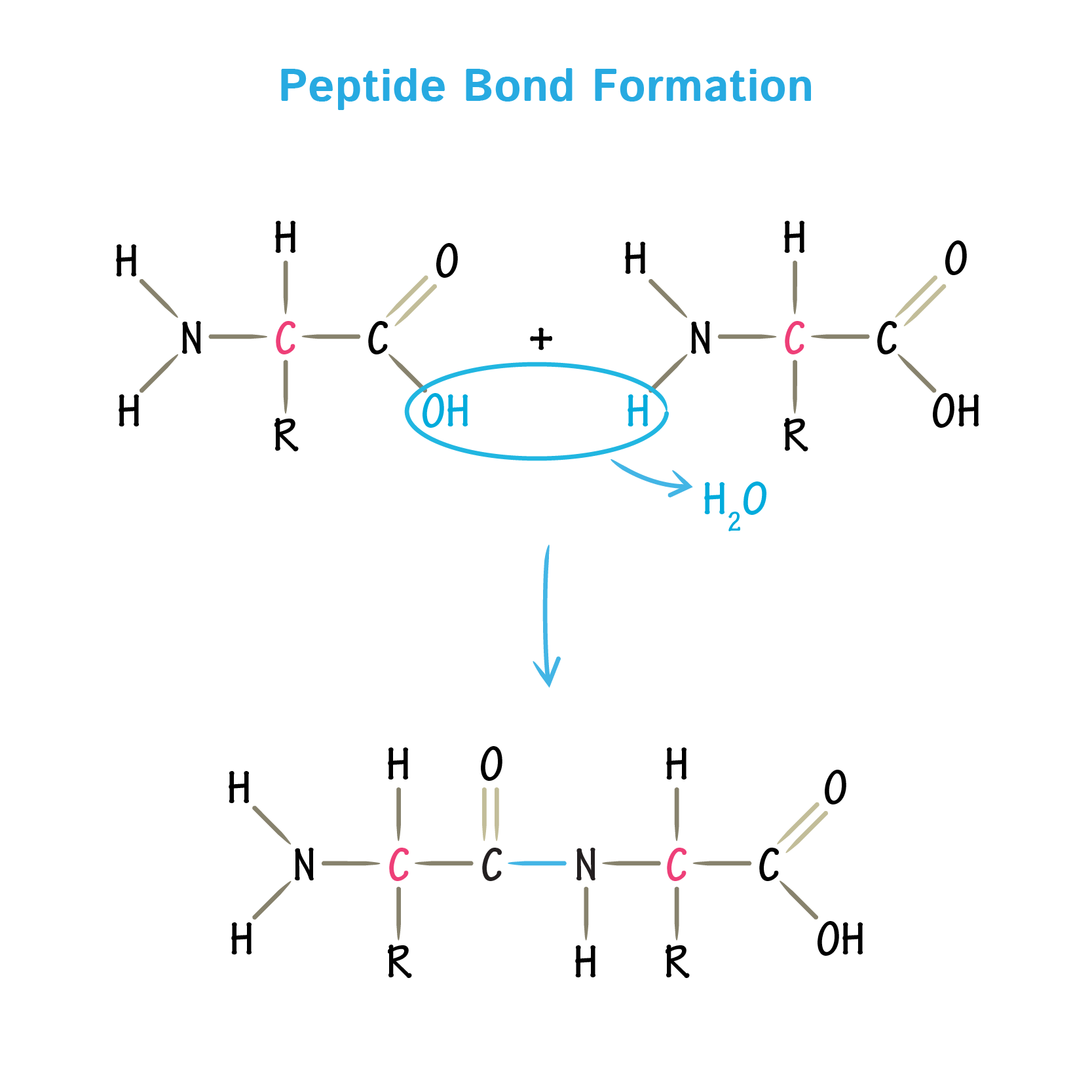
A sequence of connected amino acids creates the primary structue of proteins:
Secondary Protein Structure
Hydrogen bonds between amino acids create two types of secondary structures
Will be broken if protein is outside of normal pH or temp
Alpha Helice:
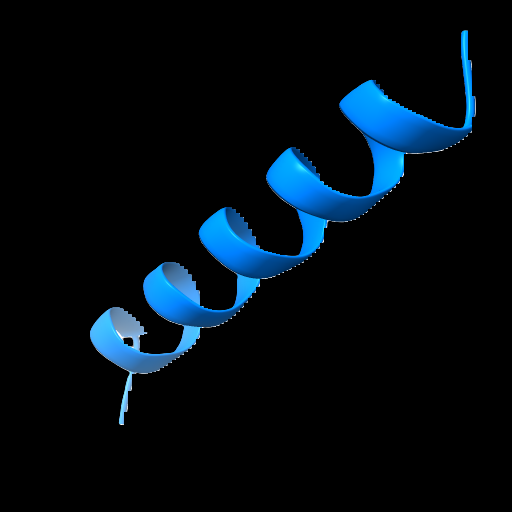
Beta Pleated Sheets:
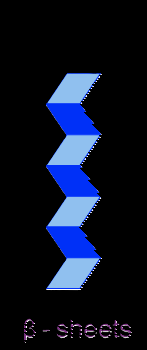
These shapes form because of the hydrogen bond attract certain sections to fold the protein into these forms.
Tertiary Structure
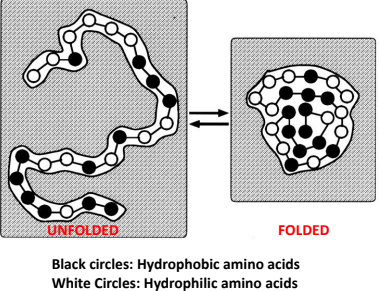
Hydrophobic Collapse: Occurs as the hydrophobic Amino Acids collapse away from the water and into the interior of hte tertiary structure of the protein. Driving factor in the formation of tertiary structure.
R side chains are either hydrophobic or hydrophilic
Hydrophobic side chained amino acids go inside and away from water
Hydrophilic side chained amino acids go outside and face the water
This is called the folding of amino acids
Protein is function at this point
Charge attraction: Occurs between acidic and base charges in R groups of amino acids
Will be broken if protein is outside of normal pH or temp
Disulfied Bridges: Covalent bonds between the sulfur atoms in side chains of two cysteine amino acids
Very strong and will not break outside of normal pH or temp
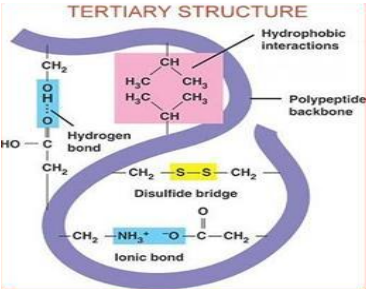
Quarternary Structure
A mix of tertiary protein molecules together
Not all proteins are inovlved in quarternary structures
Unit 2: The Cell
Eukaryotic vs Prokaryotic:
Eukaryotic:
Nucleus
Single or Multi Celled
Membrane bound organelles
Evolved from prokaryotic cells
Cytoplasm region
Prokaryotic:
No nucleus
DNA is an unbound nucleoid region
No membrane-bound organelles
Cytoplasm bound by the plasma membrane
Single celled only
3 Domains of Life:
Archaea and Bacteria - prokaryotic
Eukarya - eukaryotic
All Cells Possess:
Plasma Membrane
Cytosol
Chromosomes
Ribosomes
Organelles & Functions:
Ribosomes:
Make of protein and rRNA
Synthesizes proteins form mRNA
Free-floating or attached
Cytoplasm:
Liquid of water, salt and other nutrients
Helps maintain shape
Site of metabolic reactions
Cytoskeleton:
Helps maintain the shape of animal cells
Help vesicles get transported around the cell
3 parts
Actin filaments, microtubules, intermediate filaments
Centriole:
Creates spindle fibers during mitosis and meiosis
Spindle fibers help pull chromosomes apart
Flagella and Cilia:
Help move cells around
Flagella are longer
Cilia are shorter
Nucleus:
Surrounded by double nuclear membrane which protects DNA from denaturation
Place where transcription (DNA to mRNA) happens
Nucleolus:
Site of ribosome synthesis
Makes rRNA
Smooth ER:
Synthesis lipid/steroid hormone/detoxification
Lipids and hormones get sent to the golgi apparatus
Rough ER:
Highly folded organelle
Contiguous with the nucleus
Has ribosomes attached
Packs and sends proteins to golgi
Golgi:
Fold And modify proteins
Packages proteins/lipids into vesicles
Sends vesicles to destinations
Peroxisome:
Lipid hydrolysis to break down lipids
Fatty acid from break down go to mitochondria for ATP
Uses catalase (enzyme) to break down hydrogen peroxide (toxic)
Lysosomes:
Apoptosis - Programmed cell death
Contains a lipid bubble of hydrolytic enzymes that break down cell waste and denatured proteins into their monomers
Vacuole:
Storing and releasing fluids/biomolecules
Stores waste products until they can be broken down
Mitochondria:
Two membranes which allows for compartmentalization of different chemical reactions
Folding of inner membrane increases surface area to allow for more ATP production
Chloroplast:
Arranged in stacks of thylakoid membranes called grana
Multiple thylakoids increases surface area for more reactions
Also has two membranes allowing for compartmentalization of different reactions
Plasma Membrane:
Phospho-Lipids:
Phosphate head that is polar and hydrophilic
Lipid tail is hydrophobic and nonpolar
Cholesterol:
Controls fluidity of membrane
More or less fluid depending on temperature
Glycolipids:
Facilitated cell to cell adhesion and recognition
Glycoproteins:
Allows for cross linking of cells
Membrane Proteins:
Peripheral Proteins
Found on the inner membrane surface
Integral Proteins
Partially or completely (transmembrane) in the membrane
Amphipathic: Have both hydrophobic and hydrophilic regions
Endosymbiotic Theory:
Compartmentalization:
Mitochondria increases surface area through the inner membranes being folded
Chloroplasts have grana which increase surface area
Endosymbiosis: Ancestral eukaryotic cells engulfed an ancestral mitochondrion establishing a mutualistic relationship
Serves as the explanation for the origins of mitochondria and chloroplasts
Chloroplasts got engulfed after mitochondria
Mitochondria and Chloroplast Common Ancestry Evidence:
Contains circular DNA
Possess ribosomes
Have a double membrane
Are self-replicating
Cell Transport:
Concentration Gradient: Concentration of a particle (element or molecule) is higher in one area than in another.
Particle moving down the concentration gradient is moving from high to low (no resistance)
Particle moving up the concentration gradient is moving from low to high (resistance)
Passive Transport
Passive Transport: Movement of substances across the cell membrane without the input of energy
No energy required
Net movement of substance down their concentration gradient
Diffusion: Movement of substance down a concentration gradient
Dynamic Equilibrium: No concentration gradient as substances move in equal rates. There is no high and low concentration, it’s just constant. Substances STILL move back and forth across a membrane just in equal amounts.
Osmosis: Diffusion of water across a selectively permeable membrane
Moves from higher to lower water concentration across membrane
Moves from lower to higher SOLUTE concentration across membrane
More solute on a side means less water on that side as the solute takes up space for water
Aquaporins: Cells use aquaporins, which are transmembrane proteins, that allow the water to diffuse easily inside the cell.
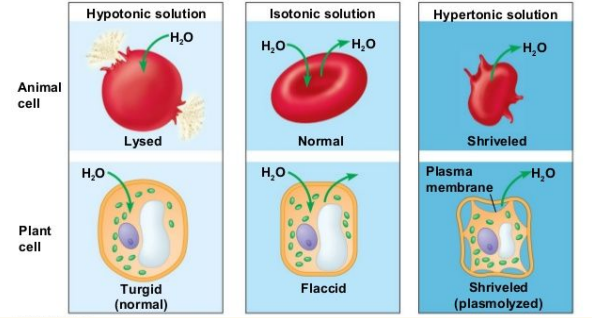
Tonacity: measuring the solute levels inside and outside the cell with three types:
Hypotonic Solutions: Lower amount of solute outside the cell (lower inside)
Causes water to move INSIDE the cell as more water is outside
Hypertonic Solutions: Higher amount of solute outside the cell (higher inside)
Causes water to move OUTSIDE the cell as more water is inside
Isotonic Solutions: Equal amount of solute inside and outside
No NET movement of water but water still keeps moving in equal rates
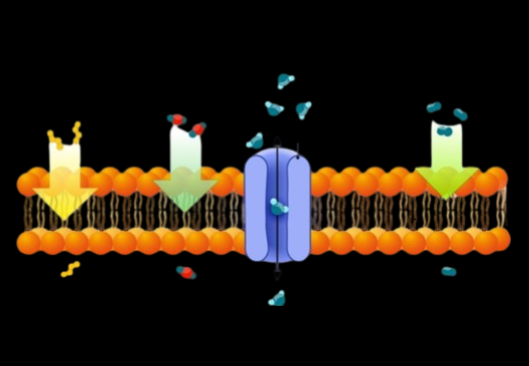
Facilitated Transport
Facilitated Transport: uses membrane proteins to help substances pass the membrane. No energy is required still.
Channel Proteins: Allow specific substances to quickly go through hydrophilic passageways.
Aquaporins
Ion Channels
Carrier Proteins: solutes bind to protein which changes protein shape to let the protein out the other side of the membrane
Glucose carrier proteins
Amino acid carrier proteins
Active Transport
Active Transport: movement of substances against their concentration gradient, requiring an input of energy. (Movement from low to high)
Sodium-Potassium Pump: A protein that uses ATP to transport 3 NA+ ions out of the cell and 2 K+ into the cell.
Electrochemical gradient: A difference in chemical and charge concentrations across a membrane.
With a Sodium-Potassium Pump there is a higher concentration of potassium ions inside and higher concentration of sodium ions (Na+) outside
3 positive charges pumped 2, 2 positive charges pumped in.
Leads to uneven amount of charges and a net negative charge inside from more positive charges being pumped out
Cotransport: protein actively pumps out a substance so it can diffuse back inside through passive transport through a cotransport protein and the substance that is having difficulty diffusing through the membrane takes a ride with the passively transported protein using the cotransport protein.
Technically the cotransport protein in itself is not using active transport but the pump protein that pumps out the substance for the cotransport protein to use DOES use active transport so the cotransport is using active transport INDIRECTLY.
Bulk Transport: Large molecules being transported across the membrane through vesicles.
Exocytosis: Intracellular vesicle often made by the golgi goes and fuses with the plasma membrane and the contents of the vesicle is excreted out.
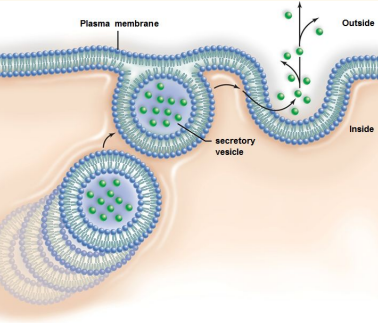
Endocytosis: Vesicles form around big molecules entering the cell and a piece of the plasma membrane separates to form a vesicle inside the cell.
Phagocytosis (cell eating): Large, solid material is taken in by endocytosis.
Water Potential & Osmolarity
Osmoregulation: maintains water balance and solute concentrations to keep the cell in homeostasis
Water Potential
Water Potential: Measurement of the amount of free energy found in a mole of water. How likely water molecules are to move from their current position to somewhere else.
Mole: SI (International system of units) measure of amount of substance
Represented by greek letter psi ψ
Measured in bars
Free Energy: Movement
Low ψ means low movement and water molecules stay still
High ψ means high movement and water molecules move out when possible
Often for plant cells
Calculating Water Potential:
Formula: ψ = ψp + ψs
ψp = Pressure potential
Measures Turgidity of plant cells which is the pressure exerted by the cell wall on the water inside
ψs = Solute Potential
Formula: ψs = -iCRT
i = Ionization constant (how many elements result from dissolving one molecule)
Sodium Chloride: NaCl → 1Na+ + 1Cl- (i = 2)
Glucose: C6H12O6 → C6H12O6 (i = 1)
Calcium Chloride: CaCl2 0< 1Ca2+ + 2Cl- (i = 3)
C = Molar Concentration (M) (moles/liter)
R = Pressure constant (R = 0.0831 L-bar/Mol-K)
T = Temperature in Kelvin (K = C +273)
Pure water with an open container has a water potential of 0
High movement of water
Water potential goes down as solute is added to negatives
Low movement of water