Translation I – mRNA Stability & tRNA Charging (Comprehensive Notes)
Page 1 – Introductory Metadata
• Lecture: “Translation I – mRNA Stability and tRNA Charging”
• Instructor: James Bert Flanegan, Ph.D.
• Department: Biochemistry & Molecular Biology, College of Medicine, University of Florida
• Course: BCH 5413
• Contact: Flanegan@ufl.edu
• Framing: First of a two-part series on translation; this installment focuses on how the cell (i) controls the half-life of mRNA molecules and (ii) accurately “charges” tRNAs so that the ribosome can interpret the genetic code.
Page 2 – Overview of Gene-Expression Pathway in Eukaryotes
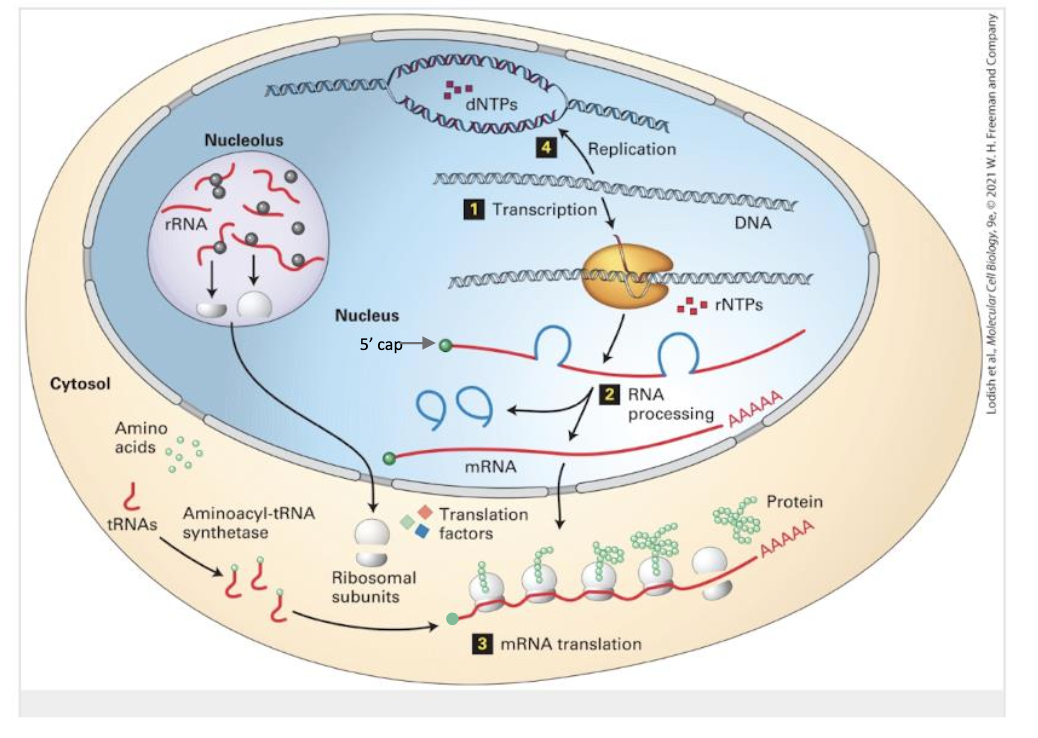
Transcription – DNA-dependent RNA polymerase synthesizes pre-mRNAs in the nucleus.
RNA Processing – Capping, poly-adenylation, and spliceosome-mediated removal of introns convert pre-mRNA → mature mRNA.
Translation – Cytoplasmic ribosomes read the processed mRNA 5′→3′ and polymerize amino acids into protein.
DNA Replication – Mentioned for completeness; covered later.
Key structural feature: the 5′ cap added during processing protects transcripts from 5′ → 3′ exonucleases and aids initiation-factor binding.
Page 3 – Why mRNA Stability Matters
• Cells fine-tune protein levels by modulating how long each mRNA persists.
• Categories:
– Stable mRNAs encode high-demand proteins (e.g., ribosomal proteins); .
– Unstable mRNAs encode transient or signaling proteins (e.g., cytokines); .
• Steady-state concentration obeys
• Thus degradation is as important as transcription for regulating gene expression.
Page 4 – Deadenylation-Dependent Decay: The Default Pathway
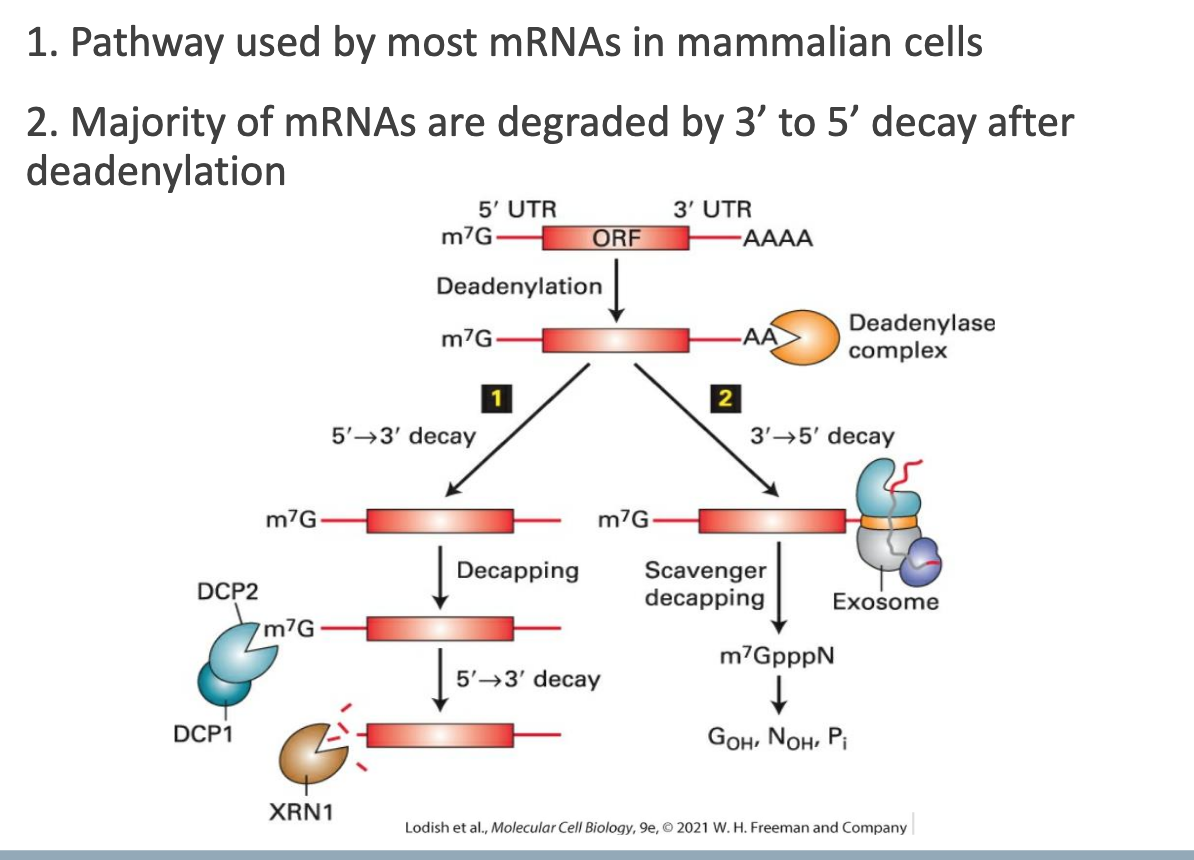
• Used by most mammalian mRNAs.
• Sequence:
Enzymatic shortening of the 3′ poly(A) tail (deadenylation).
Subsequent 3′ → 5′ decay by the exosome complex (primary route in mammals).
• A minority of transcripts may instead be decapped and degraded 5′ → 3′ (yeast bias).
Page 5 – Consequences of Poly(A) Shortening
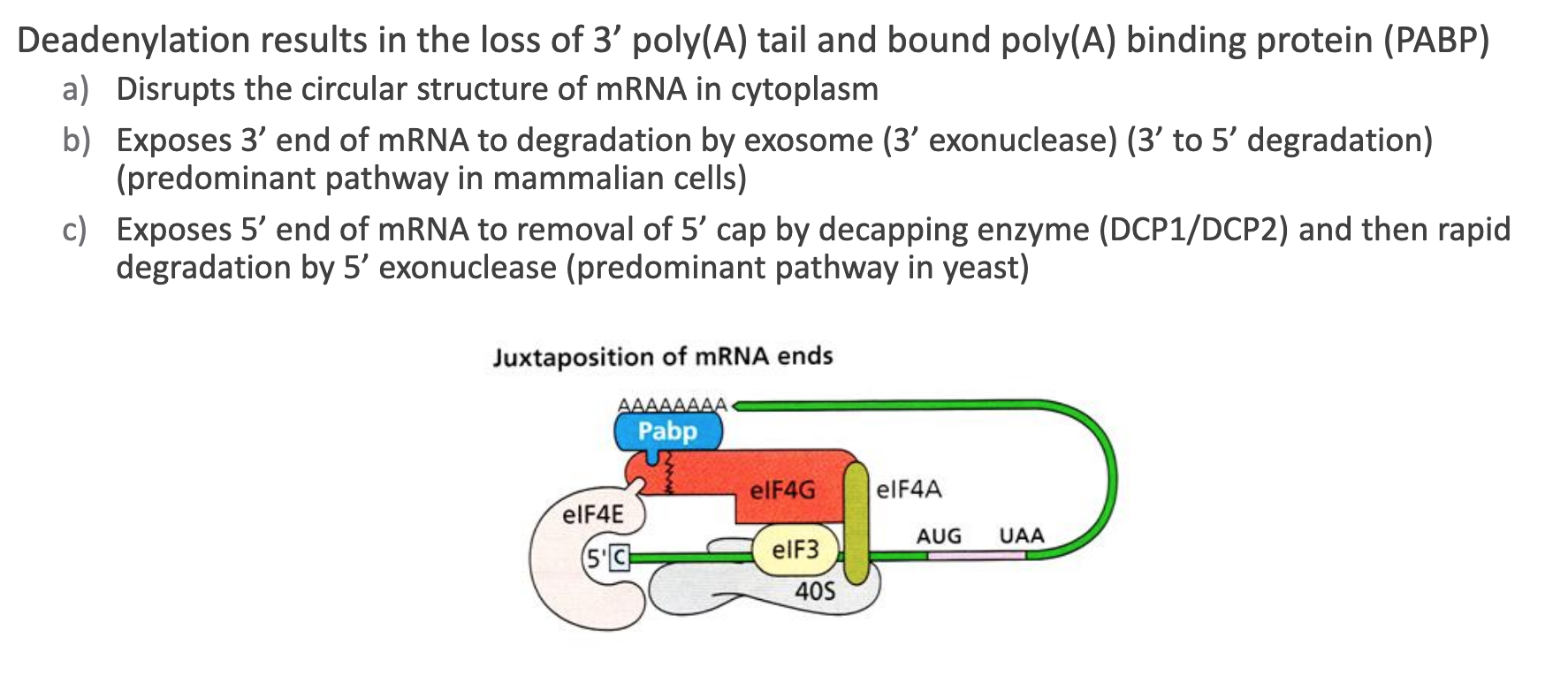
Loss of PABP (Poly(A)-Binding Protein) dissolves the closed-loop mRNP formed by 5′-cap/EIF4E ↔ PABP interactions.
Once unlooped, the exposed 3′ end recruits the exosome → rapid 3′ → 5′ nucleolysis.
Alternatively, the unshielded 5′ cap is removed by DCP1/DCP2 decapping enzyme, then the Xrn1 5′ → 3′ exonuclease digests the body (dominant in budding yeast).
Overall: deadenylation is the “commitment step” for bulk mRNA turnover.
Page 6 – Sequence Determinants of Instability
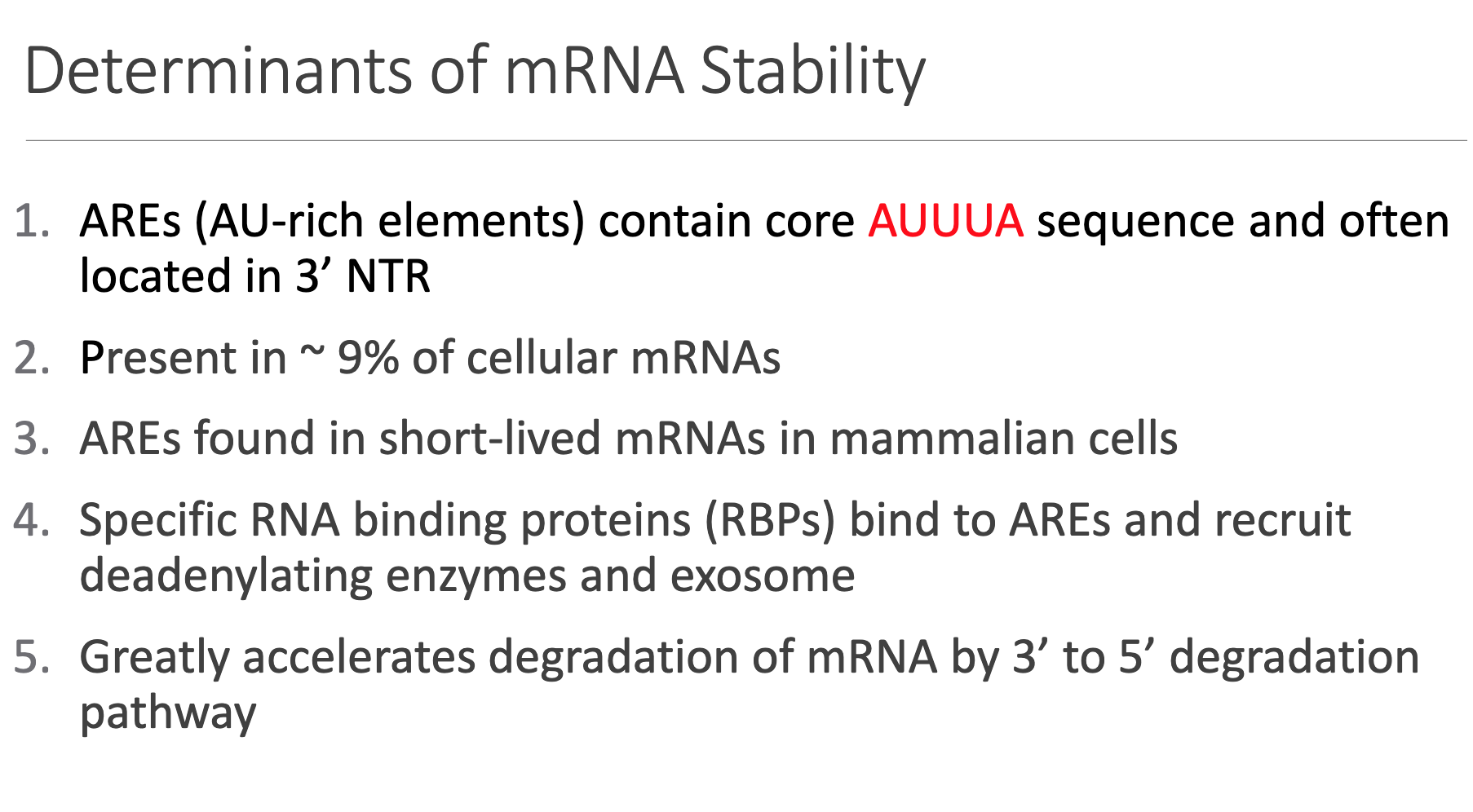

• AREs (AU-Rich Elements)
– Canonical core motif: AUUUA; typically reside in the 3′ untranslated region.
– Present in of human mRNAs; enriched in rapidly-induced transcripts (growth factors, proto-oncogenes, cytokines).
• RNA-Binding Proteins (RBPs) such as TTP, AUF1, and HuR dock onto AREs and recruit the deadenylation machinery/exosome.
• Effect: Orders-of-magnitude acceleration of the 3′ → 5′ degradation pathway, overriding baseline stability.
Page 7 – Endonucleolytic Pathway (ARE-Independent)
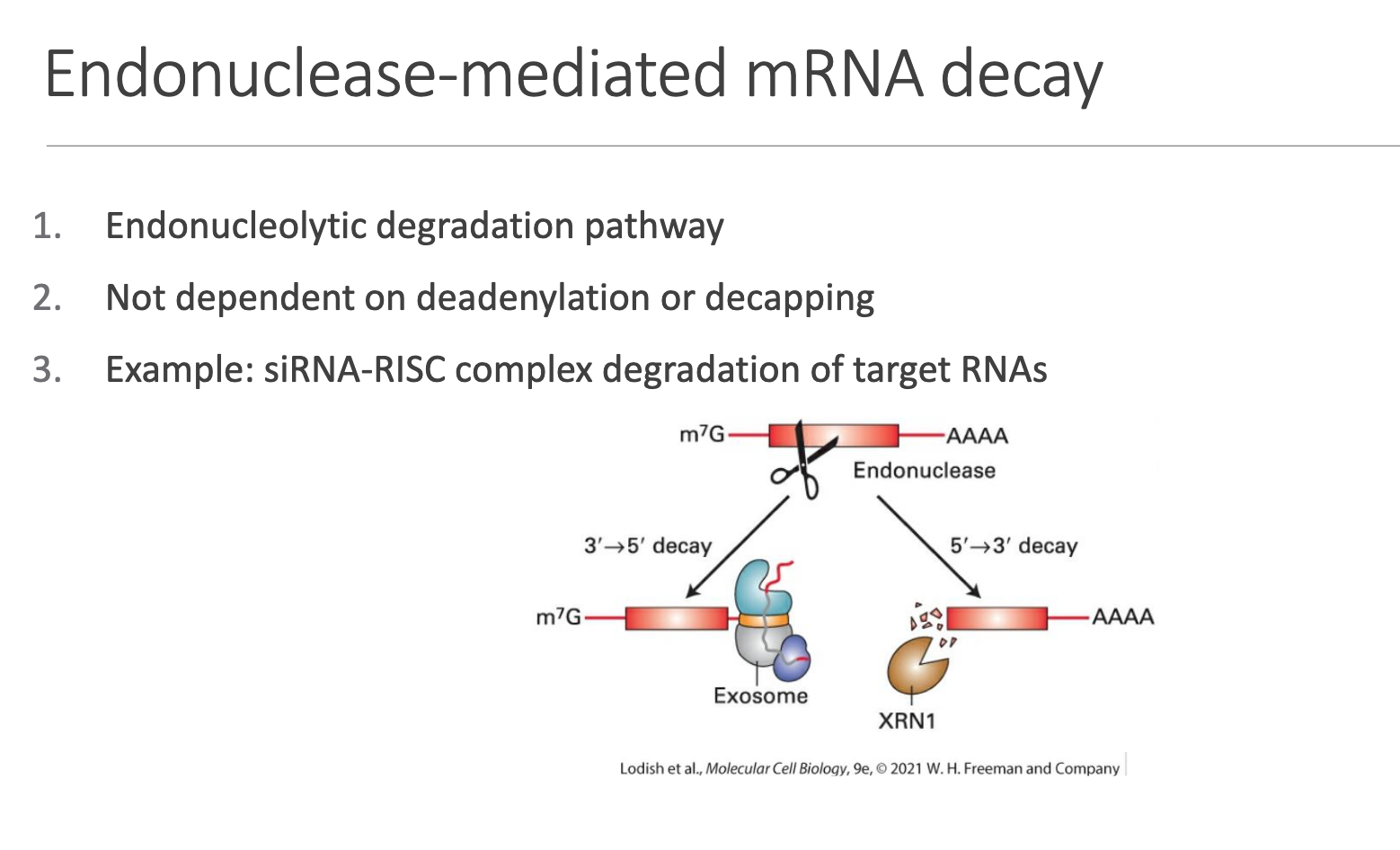
• Mechanism: Site-specific internal cleavage by an endonuclease → fragment ends are then cleared by exonucleases.
• Not contingent on prior deadenylation or decapping.
• Key example: RNA-interference siRNA-loaded RISC mediates AGO2-dependent slicing of perfectly complementary targets.
Page 8 – Translation Essentials
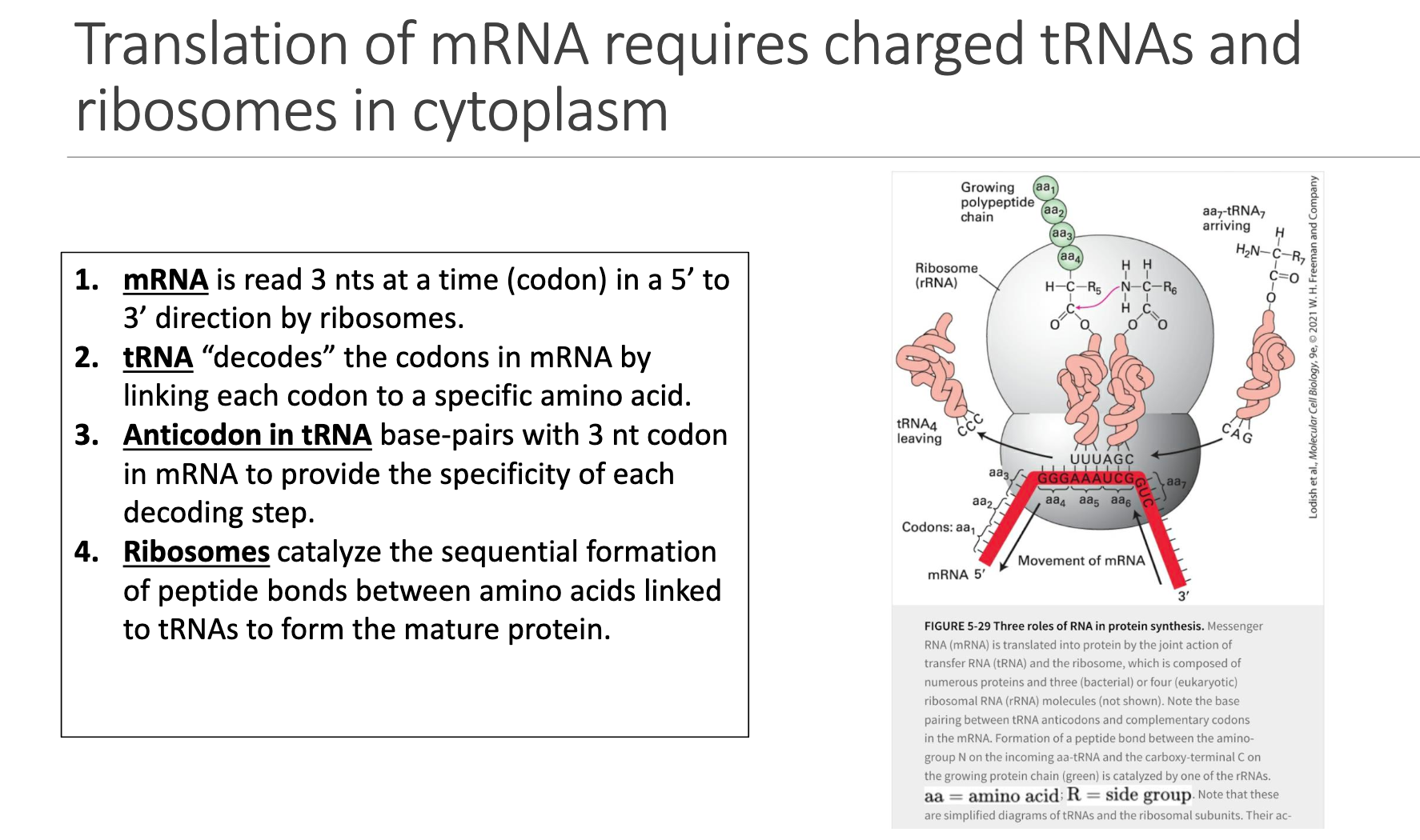
Reading Frame: Ribosome moves 5′→3′, interrogating the mRNA three nucleotides at a time (a codon).
tRNA Decoding: Each tRNA brings an amino acid whose identity is specified by the codon:anticodon interaction.
Anticodon Loop: Three bases in tRNA base-pair antiparallel with the codon (positions 1↔3, 2↔2, 3↔1).
Peptide-Bond Formation: The ribosome’s large subunit peptidyl-transferase center forms peptide bonds, elongating the nascent chain.
Page 9 – Quantitative Anatomy of the Genetic Code
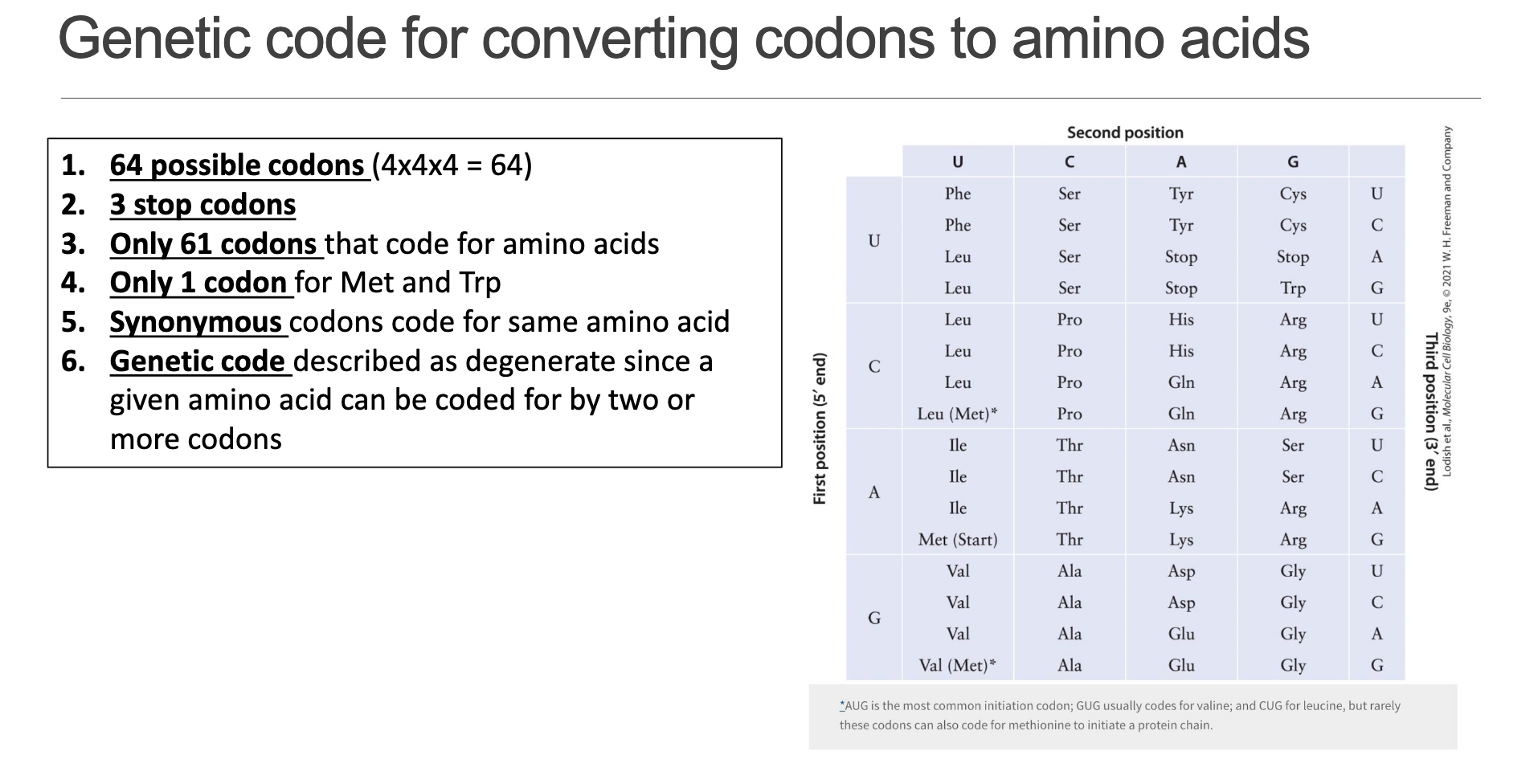

• Total possible triplets: .
• Stop codons: (UAA, UAG, UGA).
• Sense codons: (encode amino acids).
• Only one codon each for Met (AUG) and Trp (UGG); all other amino acids are represented by 2–6 “synonymous” codons.
• Because multiple codons specify the same residue, the code is termed degenerate.
Page 10 – Making the Arithmetic Work

Hierarchy in typical eukaryotes:
20\;\text{amino acids} < \approx50\;\text{different tRNAs} < 61\;\text{sense codons}
Implications:
A single amino acid is often attached to multiple different tRNAs (iso-acceptors).
A given tRNA can recognize multiple codons—possible through wobble base-pairing at the third codon position.
Page 11 – tRNA Structure and Modifications
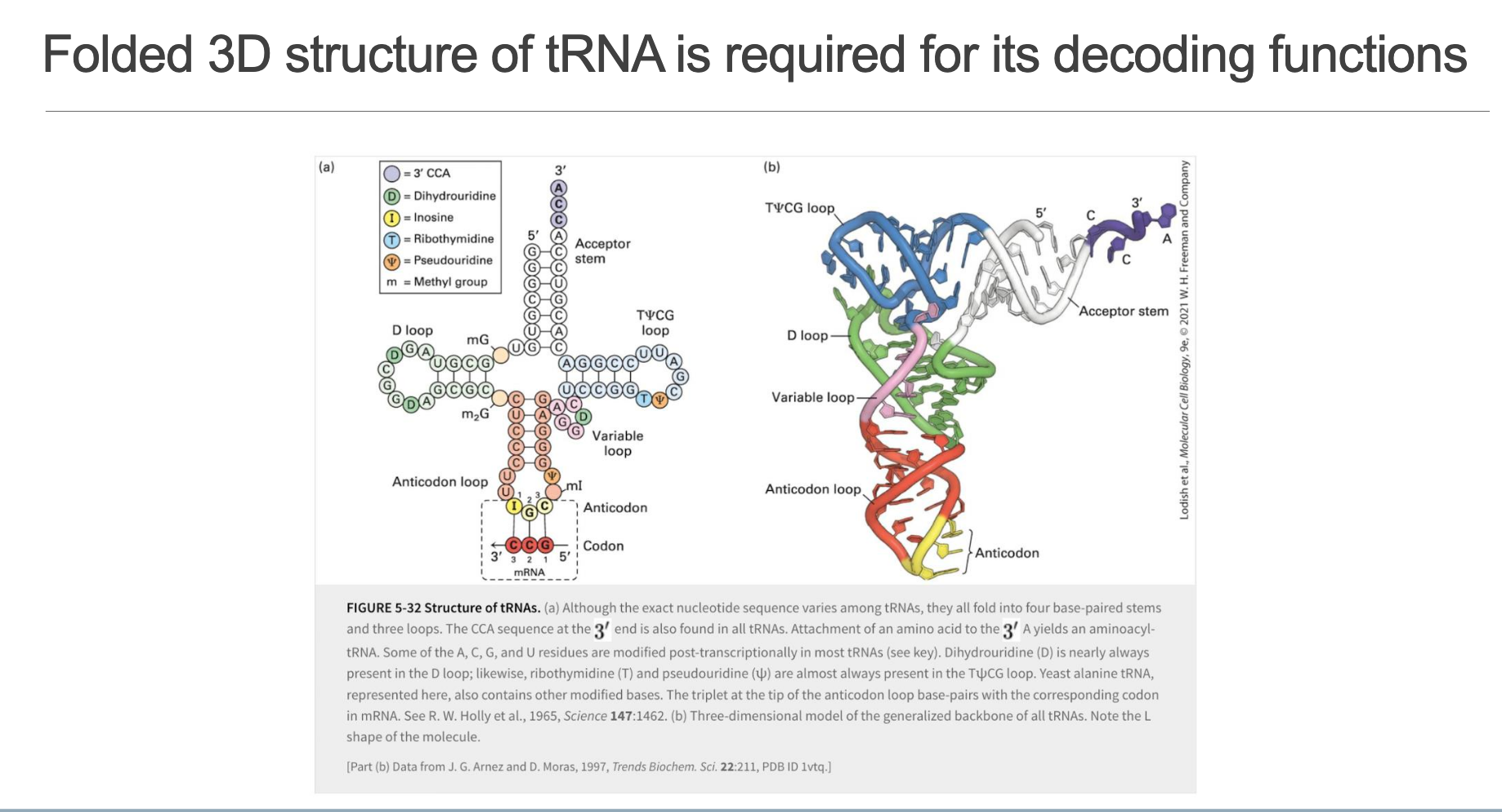
• Secondary (cloverleaf) features:
Acceptor Stem (7 bp) – 3′ terminus ends with the universal CCA sequence; amino acid attaches to the 3′-A via an ester linkage.
D Loop – Contains dihydrouridine (D).
Anticodon Loop – Triplet that pairs with mRNA codon; often possesses base modifications (e.g., inosine (I), m²G, queuosine).
Variable Loop – Size varies (hence name).
TΨC Loop – Almost invariant sequence T Ψ C (ribothymidine–pseudouridine–cytidine).
• Tertiary fold: L-shaped molecule stabilized by coaxial stacking; places the acceptor stem and anticodon loop at opposite ends—ideal for bridging the decoding center and PTC of the ribosome.
Page 12 – Wobble Pairing Rules
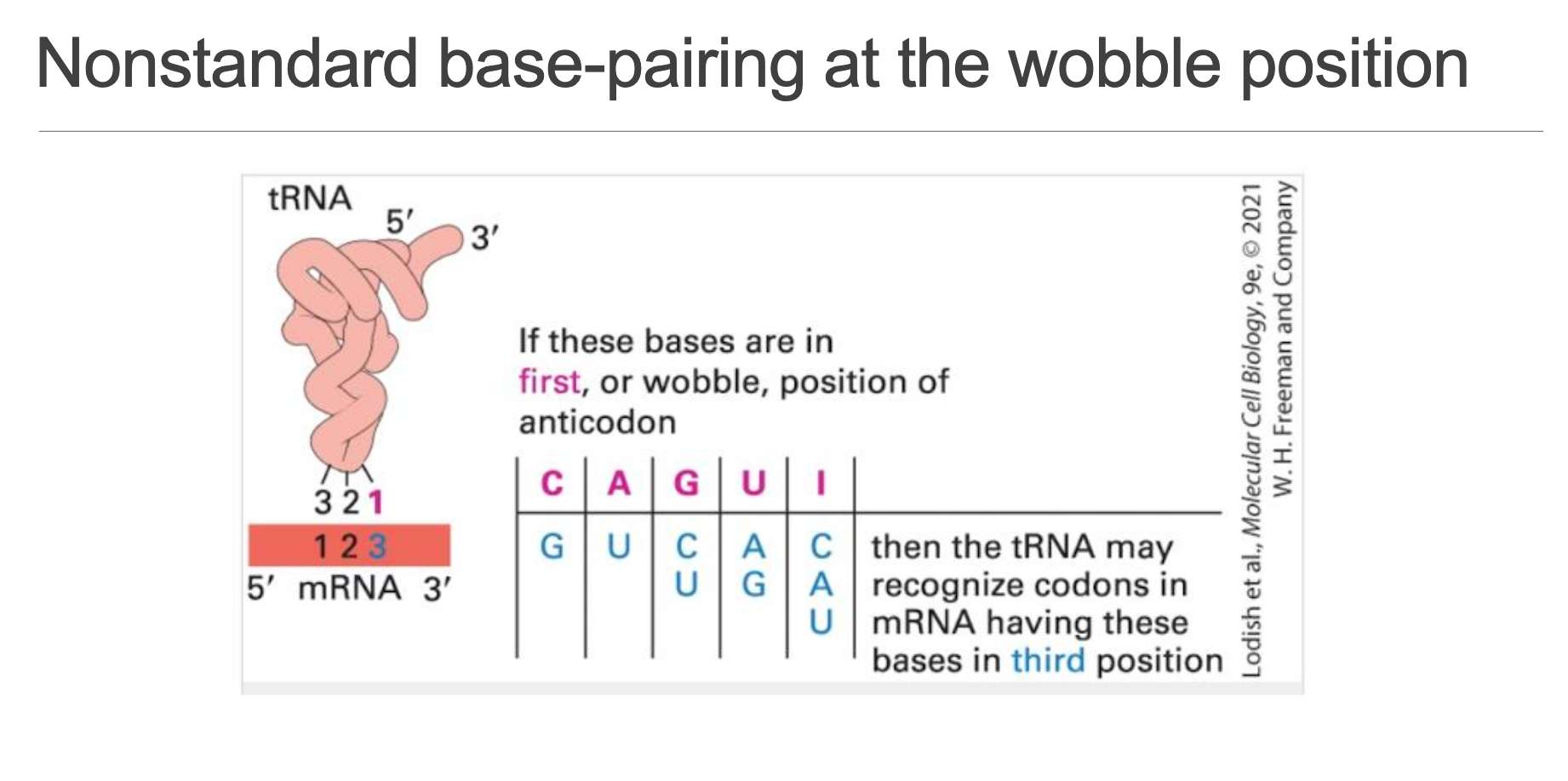
If the first (5′) nucleotide of the tRNA anticodon is … then it can read the following third (3′) codon bases:
• only
• only
•
•
•
Result: fewer than 61 different tRNAs suffice to decode all sense codons without ambiguity.
Page 13 – Aminoacyl-tRNA Synthetases (aaRS): The Accuracy Gatekeepers
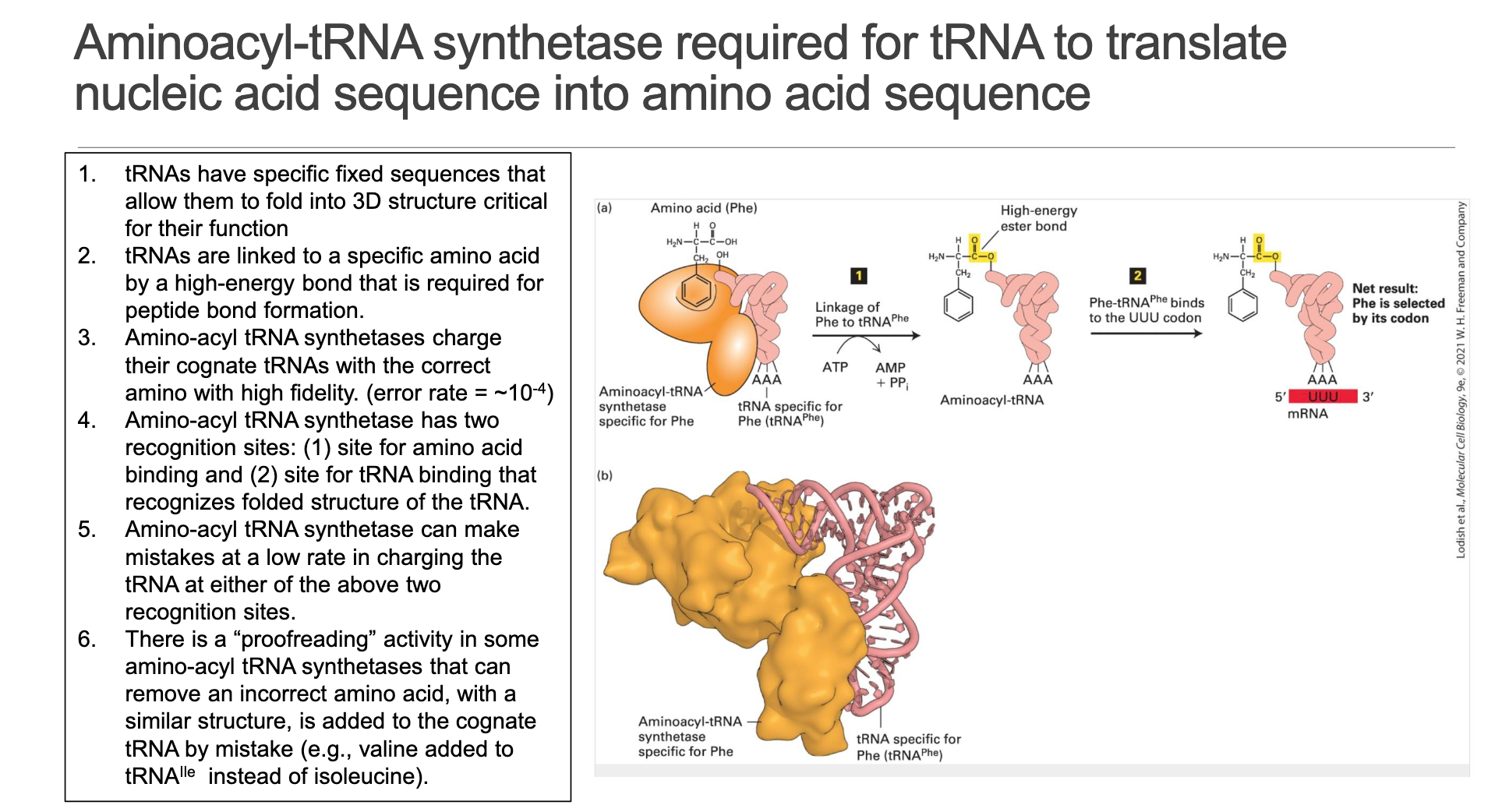
Dual Recognition – Each aaRS possesses an amino-acid binding pocket and a tRNA identity pocket that “reads” shape + key nucleotides (identity elements).
High-Energy Ester Bond – Charging (" aminoacylation ") consumes ATP, forming aminoacyl-AMP then transferring the amino acid to tRNA 3′-A. The resulting aminoacyl-tRNA carries the activation energy for peptide-bond formation.
Error Rate: per charging event—orders of magnitude lower than the ribosome’s decoding error (~).
Proofreading/Editing Sites – Many aaRSs (e.g., IleRS) possess a hydrolytic editing pocket that removes mis-activated amino acids (e.g., valine when isoleucine is correct) after or before attachment.
Functional Impact: Fidelity at this step is critical; a mis-charged tRNA bypasses codon:anticodon quality control and directly installs the wrong residue into every protein copy.
Connections & Context
• mRNA decay pathways intersect with translation: deadenylation uncouples PABP–EIF4G, simultaneously silencing translation and earmarking the transcript for destruction.
• Wobble allows evolutionary flexibility—codon bias can optimize translational speed/accuracy without expanding the tRNA gene set.
• Errors at the aaRS step can cause disease (e.g., Charcot-Marie-Tooth neuropathies from GlyRS mutations) underscoring the importance of enzymatic proofreading.