BIOL 3130 LEC 3
1) Why is gene regulation important if all cells have the same DNA?
Gene regulation allows different cell types to express different genes, enabling specialization, environmental response, and proper cellular function.
2) What level of gene regulation is emphasized in this lecture?
Transcriptional regulation, which controls RNA synthesis by RNA polymerase II.
3) What is the technical nomenclature for RNA polymerase and what is its fundamental function?
RNA polymerase is technically referred to as a DNA-directed RNA polymerase, and its fundamental function is to synthesize RNA using a DNA template during the process of transcription.
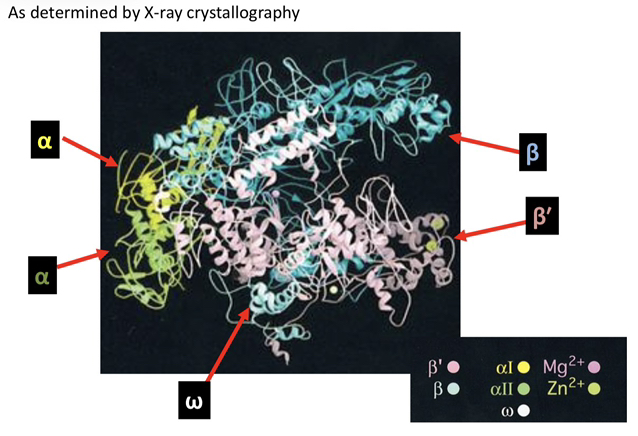
4) List the specific subunits that compose the E. coli RNA polymerase and their respective molecular weights.
The enzyme is composed of two large subunits, β (150 kD) and β′ (160 kD); two α subunits (40 kD each); one σ factor (70 kD); and one ω subunit (10 kD).
5) Describe the role of the ω (omega) subunit within the RNA polymerase complex.
The ω subunit is considered non-essential for the catalytic activity of the enzyme, but it plays a critical role in the proper assembly of the enzyme complex.
6) Distinguish between the "Core enzyme" and the "Holoenzyme" in terms of subunit composition.
The Core enzyme consists of the subunits α2ββ′ω, whereas the Holoenzyme includes all core components plus the σ (sigma) factor.
7) How does the absence of the sigma factor affect the transcriptional capability of the Core enzyme on natural DNA?
Without the sigma factor, the Core enzyme is unable to initiate transcription on intact, natural DNA templates.
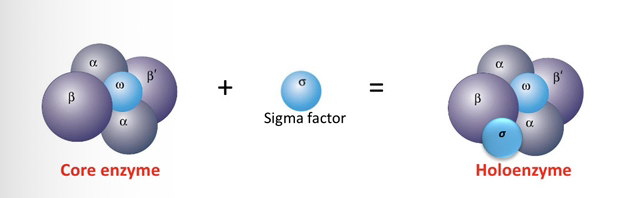

8) Based on the 1969 experiments, how does the Core enzyme's activity differ when using "nicked" DNA versus "intact" DNA?
The Core enzyme is capable of transcribing DNA that contains nicks(single-strand breaks), but it requires the addition of the sigma factor to transcribe intact DNA.
9) Why is the sigma ( σ) subunit specifically referred to as a "Specificity Factor"?
It is called a specificity factor because it is required to direct the RNA polymerase to initiate transcription at the correct locations on intact, natural DNA rather than at random nicks.
10) Define a "nick" in the context of a DNA template and explain its relevance to Core enzyme activity.
A nick is a single-strand break in the sugar-phosphate backbone of the DNA; these nicks allow the Core enzyme to initiate RNA synthesis in vitro even in the absence of a sigma factor.
11) Define a "Promoter" and describe its typical location relative to the transcription process.
A promoter is a specific DNA sequence where RNA polymerase binds to initiate transcription, and it is usually located upstream of, or includes, the Transcriptional Start Site (TSS).
12) Why is the sigma ( σ) subunit essential for transcription on natural, cellular DNA?
The sigma subunit is essential because it permits the RNA polymerase to recognize and bind to authentic promoter sequences, ensuring that transcription is specific and directed rather than random.
13) Explain why the Core RNA polymerase is capable of transcribing nicked DNA in vitro but not intact DNA.
Nicks and gaps in the DNA provide artificial entry points that allow the Core RNA polymerase to bind nonspecifically and carry out transcription; however, because normal DNA lacks these nicks, the enzyme cannot bind without the sigma factor.
14) Contrast the initiation of transcription on nicked DNA versus promoter-driven transcription.
Transcription on nicked DNA is considered "not natural" and occurs nonspecifically in vitro, whereas transcription at promoters is specific, occurs on intact DNA, and is strictly directed by the sigma ( σ) subunit.
15) What was the purpose of the filter binding studies conducted by Hinkle and Chamberlin?
Hinkle and Chamberlin used filter binding studies to compare how tightly the Core RNAP binds to DNA compared to the Holoenzyme, demonstrating that the sigma factor specifically facilitates promoter binding.
16) What does the presence of the sigma subunit change regarding the binding behavior of the RNA polymerase holoenzyme?
The presence of the sigma subunit transforms the enzyme from a molecule that binds DNA nonspecifically at nicks into a specialized complex that recognizes and binds tightly to authentic promoter sites.
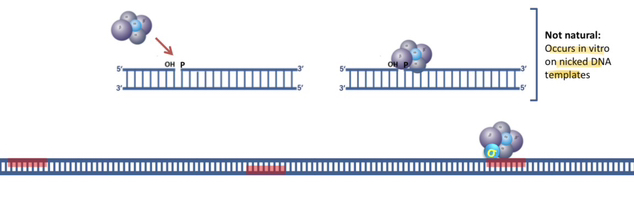
17) What is the fundamental purpose of a filter binding assay in molecular biology?
The filter binding assay is an in vitro technique used to determine if a specific protein of interest possesses the ability to bind to double-stranded DNA (dsDNA).
18) Describe the specific binding properties of nitrocellulose filters regarding proteins and nucleic acids.
Nitrocellulose filters have a natural affinity for proteins and single-stranded nucleic acids, but they do not bind to double-stranded DNA on their own.
19) How is the DNA prepared in a filter binding assay to ensure that interactions can be detected?
The double-stranded DNA used in the experiment must be radioactively labeled so that its presence on the filter can be measured after the washing process.
20) In a filter binding assay, what does the presence of radioactivity on the filter indicate after the final wash?
The presence of radioactivity indicates that the protein bound to the dsDNA, forming a complex that was trapped on the nitrocellulose filter while the unbound DNA passed through.
21) Explain why radioactive dsDNA passes through the nitrocellulose filter if the protein of interest does not bind to it.
Because dsDNA does not inherently stick to nitrocellulose, it will simply flow through the filter and be washed away unless it is physically anchored there by a protein that has successfully bound to it.
22) Outline the basic experimental steps of a filter binding assay from incubation to detection.
The process involves incubating a protein with radioactively labeled dsDNA, passing the mixture through a nitrocellulose filter, washing to remove unbound material, and finally measuring the radioactivity remaining on the filter.
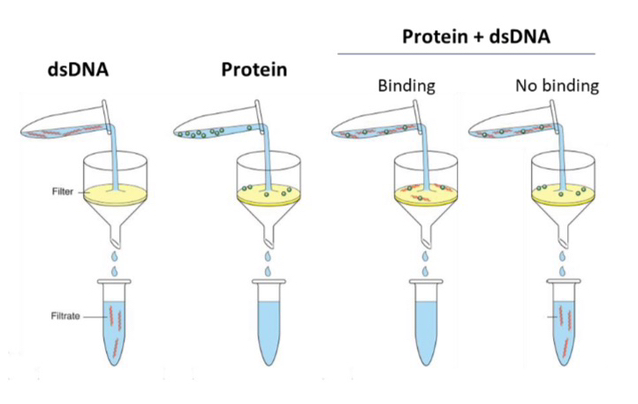
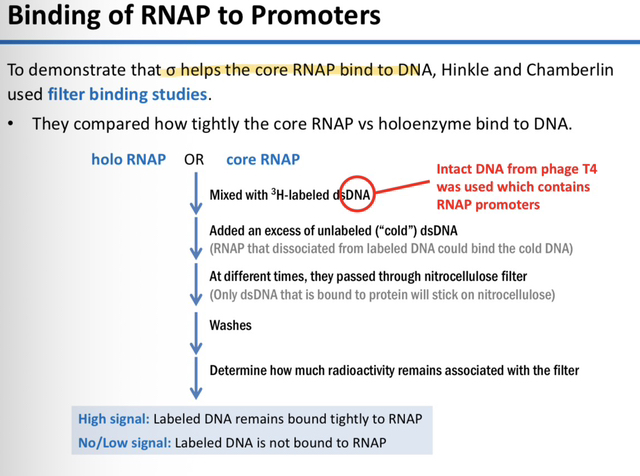
23) What was the primary goal of the filter binding studies conducted by Hinkle and Chamberlin?
The goal was to compare the binding affinity (tightness) of the Core RNAP versus the Holoenzyme to DNA to determine how the sigma (𝜎) factor affects the stability of the RNAP-DNA complex.
24) Why was an excess of unlabeled ("cold") dsDNA added to the mixture during the experiment?
The cold DNA acted as a competitor; if the RNAP dissociated from the radioactive DNA, it would likely bind to the excess cold DNA, allowing researchers to measure how long the original radioactive complex stayed together.
25) In this specific experiment, what does a "High Signal" (high radioactivity) on the filter represent?
A high signal indicates that the labeled DNA remained tightly bound to the RNA polymerase and did not dissociate during the incubation period or the wash.
No/Low signal: Labeled DNA is not bound to RNA
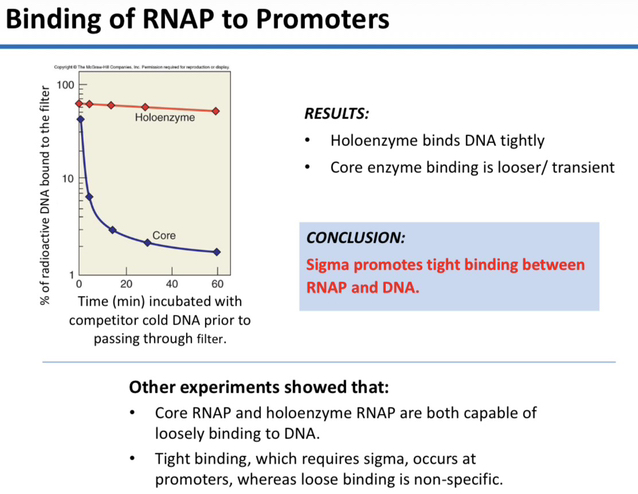
26) Based on the results of the filter binding assay, contrast the binding of the Holoenzyme with that of the Core enzyme.
The results showed that the Holoenzyme binds DNA tightly and stably, whereas the Core enzyme binding is loose and transient, leading to a rapid loss of signal over time.
27) What major conclusion was drawn regarding the role of the sigma factor in DNA binding?
The researchers concluded that the sigma (σ) factor promotes tight binding between RNA polymerase and the DNA template.
28) Compare the locations of "tight binding" versus "loose binding" for RNA polymerase.
Tight binding occurs specifically at promoters and requires the sigma factor, while loose binding is non-specific and can be performed by both the Core and Holoenzyme.
29) In a graph plotting "% of radioactive DNA bound" vs. "Time," what would the curve for the Core enzyme look like compared to the Holoenzyme?
The Core enzyme curve would drop rapidly toward zero, reflecting loose/transient binding, while the Holoenzyme curve would remain relatively high or flat, reflecting stable, tight binding.
30) How does the presence of sigma change the RNAP-DNA interaction from a "natural" perspective?
The sigma factor transforms the interaction from a non-specific, loose association into a highly specific, tight interaction at authentic promoter sequences.
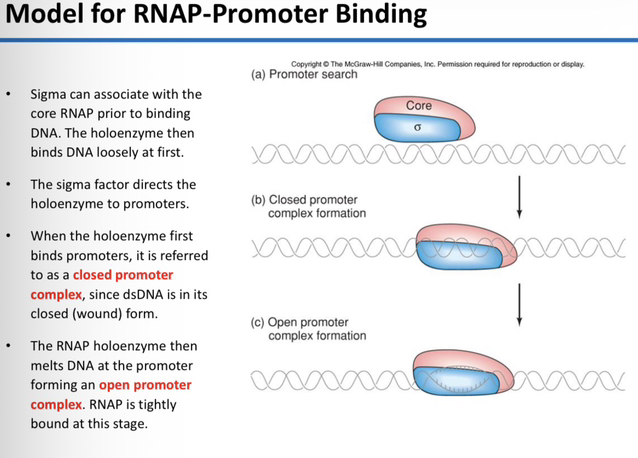
31) Describe the initial association between the sigma factor, Core RNAP, and DNA. Define the "Closed Promoter Complex" and explain the state of the DNA during this stage. What transition must occur for the "Open Promoter Complex" to form, and how does this affect binding affinity?
The sigma factor associates with the Core RNAP to form the holoenzyme prior to DNA binding; once formed, the holoenzyme initially binds to the DNA loosely.
The closed promoter complex occurs when the holoenzyme first binds to a promoter site while the DNA remains in its closed, double-stranded (wound) form.
The holoenzyme must melt (unwind) the DNA at the promoter to form the open promoter complex, at which stage the RNA polymerase becomes tightly bound.
32) What is the specific role of the sigma factor during the transition from non-specific DNA binding to promoter binding?
The sigma factor acts as a guide that directs the holoenzyme to recognize and bind specifically to authentic promoter sequences.
33) What is a "DNA element" in the context of gene regulation?
An element is a specific DNA sequence that is often recognized and bound by a specific protein, such as RNA polymerase or a transcription factor.
34) Identify the promoter element discovered by David Pribnow and describe its typical location.
David Pribnow identified the Pribnow box, a 6-7 bp sequence located approximately 10 bp upstream of the transcriptional start site (TSS).
35) What is the consensus sequence of the -10 element (Pribnow box) in E. coli?
The consensus sequence for the -10 element is TAtAaT (often written as TATAAT).
36) Why is the identification of a "consensus sequence" like the Pribnow box significant for understanding RNAP function?
The consensus sequence represents the common sequence element across different promoters that allows the holoenzyme (via the sigma factor) to identify exactly where to initiate transcription.
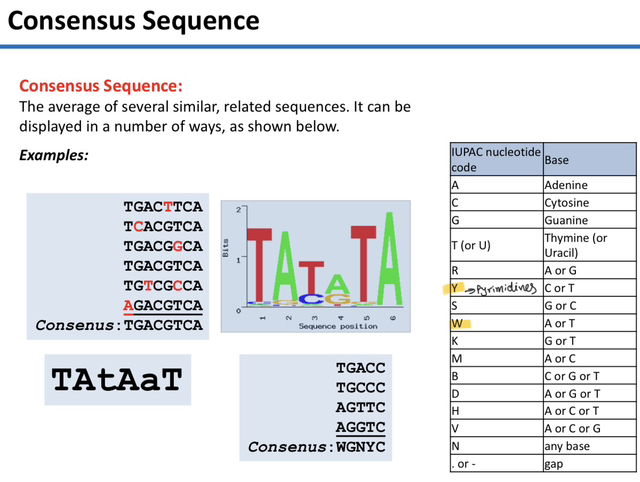
37) What is a "consensus sequence" and how is it determined?
A consensus sequence is the calculated average of several similar, related sequences, representing the most frequent nucleotide found at each position across multiple examples.
38) In the IUPAC nucleotide code, what do the symbols R, Y, and Nrepresent?
In the IUPAC code, R represents a purine (A or G), Y represents a pyrimidine (C or T), and N represents any base.

39) Identify the two primary elements that make up the E. coli core promoter and state their approximate locations.
The core promoter consists of the -10 box (Pribnow box) and the -35 box, named for their approximate positions in nucleotides upstream of the transcriptional start site (TSS).
40) On which DNA strand are promoter consensus sequences typically reported?
Consensus sequences for promoters are traditionally reported as they appear on the coding strand (the non-template strand).
41) Define "Promoter Strength" in the context of bacterial transcription.
Promoter strength refers to how effectively a promoter attracts RNA polymerase to initiate multiple rounds of transcription.
42) According to the "Consensus Model," what determines if a promoter is considered "strong" in E. coli?
A promoter is considered stronger the more closely its sequence matches the consensus sequence of the -10 and -35 boxes.
43) Aside from the actual sequence of the boxes, what other physical factor significantly affects promoter strength?
The spacing between the -10 and -35 elements is a critical factor that influences how well the RNA polymerase can bind and initiate transcription.
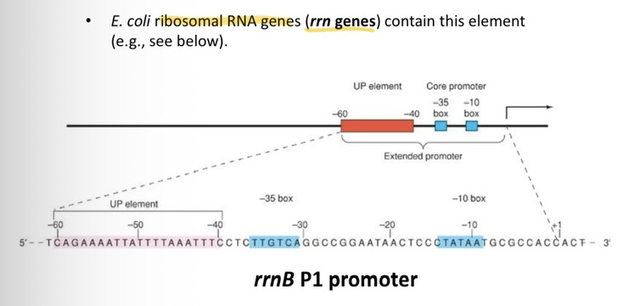
44) What is the "UP element," and what is its specific effect on transcription levels?
The UP element is an additional DNA sequence located upstream of the core promoter that significantly enhances promoter strength, stimulating transcription by a factor of 30 times.
45) Provide an example of a specific gene set in E. coli that utilizes an UP element and explain why this is necessary.
The ribosomal RNA (rrn) genes, such as rrnB P1, contain an UP element to ensure high levels of transcription, as these genes are considered very strong promoters
46) What are the three distinct phases of transcription, and what chemical event marks the transition into the first phase?
The three phases are initiation, elongation, and termination, with initiation officially beginning upon the formation of the first phosphodiester bond in the RNA transcript.
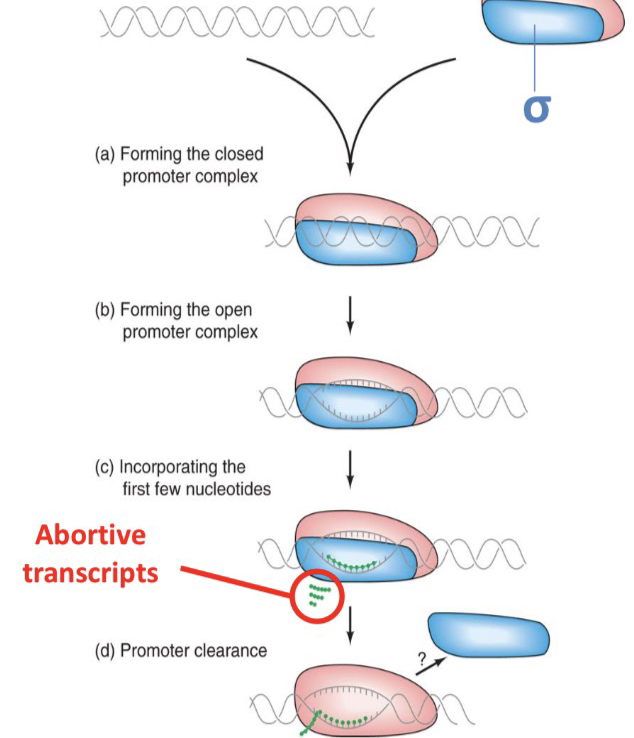
47) Define "abortive transcripts" and describe their characteristics as discovered by Carpousis and Gralla.
Abortive transcripts are very short RNA oligonucleotides (typically 2–6 nucleotides long) that are synthesized and released by RNA polymerase before it leaves the promoter to begin full-length synthesis.
48) In the Carpousis and Gralla experiment, what was the function of adding Heparin, and what did it prove?
Heparin acts as a competitor that prevents RNA polymerase from re-initiating transcription; because multiple abortive transcripts were still found per RNAP, it proved that these short RNAs are generated while the enzyme is still bound to its original promoter
49) Describe the stages of transcription initiation:
RNAP holoenzyme forms and binds to promoter DNA, forming the closed promoter complex.
DNA is melted, converting the closed promoter complex to an open promoter complex.
While remaining at the promoter, RNAP polymerizes the first few nucleotides.
At this stage, abortive transcripts are generated.
Eventually, the first few nucleotides of the full-length transcript are synthesized.
As the transcript becomes long enough to form a stable hybrid with the melted template DNA, RNAP moves forward in what is referred to as promoter clearance.
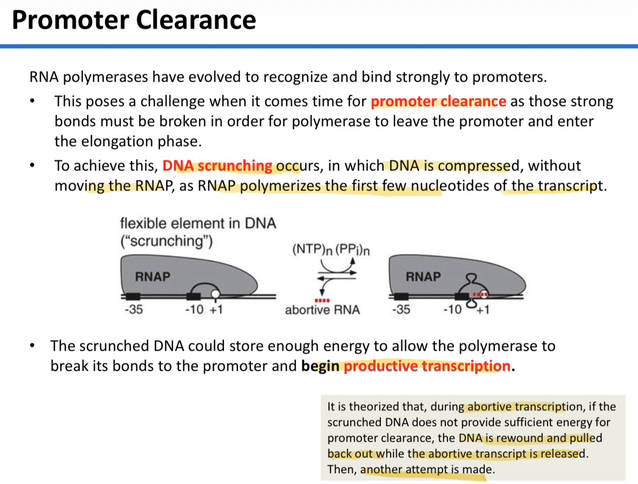
50) Explain the concept of "DNA Scrunching" and its purpose during the initiation phase.
DNA scrunching occurs when RNA polymerase pulls downstream DNA into the enzyme and compresses it while remaining stationary at the promoter; this process is theorized to store mechanical energy needed to break the tight bonds between RNAP and the promoter.
51) Define "Promoter Clearance" and identify the structural requirement for this transition to occur.
Promoter clearance is the stage where RNA polymerase moves forward into elongation; it occurs once the transcript is long enough to form a stable hybrid with the melted template DNA, providing enough energy to overcome the strong initial binding at the promoter.
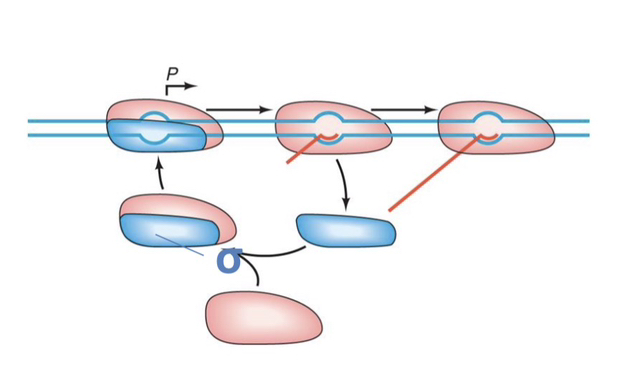
52) Describe the "Sigma Cycle" and explain what happens to the σ factor after transcription initiation.
The Sigma Cycle is the process where the σ factor is released from the RNA polymerase shortly after initiation, leaving the Core RNAP to complete elongation while the free σ factor associates with a new Core enzyme to begin a new round of transcription.
53) Is the sigma (σ) factor required for the elongation phase of transcription?
No, the σ factor is not required for elongation; once initiation is successful and the enzyme clears the promoter, the Core RNAP is fully capable of completing the transcript on its own.
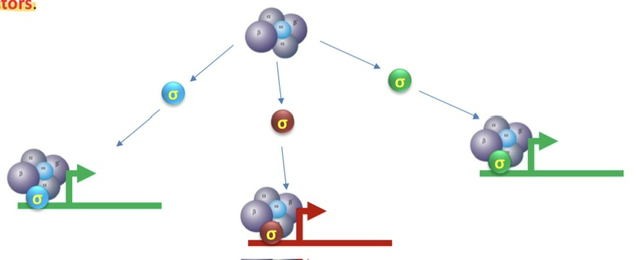
54) Why are sigma factors referred to as "specificity factors" in bacteria?
They are called specificity factors because each type of sigma factor is responsible for directing the RNA polymerase holoenzyme to a specific subset of genes by recognizing different promoter sequences.
55) Identify the primary sigma factor in E. coli and describe its general function.
The primary sigma factor in E. coli is σ70 , and it is responsible for the transcription of genes required for normal, vegetative growth.
56) Provide examples of specialized sigma factors and the environmental conditions that might trigger their use.
Specialized sigma factors are used to transcribe genes for specific stress responses or developmental stages, such as heat shock genes or genes required for sporulation.
57) Compare the primary sigma factor of E. coli with that of Bacillus subtilis.
While E. coli utilizes σ70 for normal growth, Bacillus subtilis utilizes a functionally equivalent primary sigma factor known as σ43.
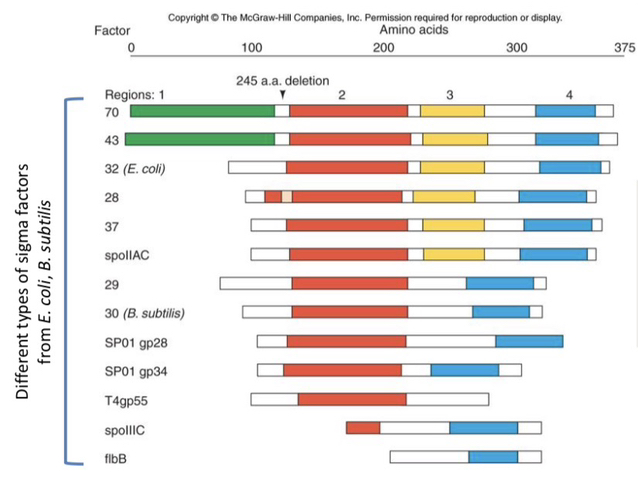
58) Describe the general structural organization of sigma factors as shown in the alignment of different species.
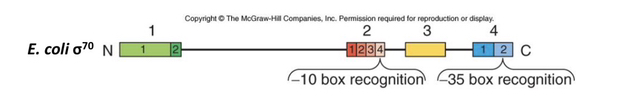
Sigma factors are polypeptides organized into four homologous regions (aligned from the N-terminus to the C-terminus) that have conserved sequences and specific functional roles in binding to either the RNAP core or DNA.
59) What is the specific regulatory function of Homology Region 1 in primary sigma factors?
Homology Region 1 acts as a safeguard that prevents the sigma factor from binding to DNA on its own when it is not part of the RNA polymerase holoenzyme complex.
60) Which specific subregion of the sigma factor is responsible for recognizing the -10 box (Pribnow box)?
The Subregion 2.4 within Homology Region 2 is responsible for binding to the -10 box of the promoter.
61) Describe the dual binding role of Homology Region 3 in the transcription complex.
Homology Region 3 is characterized by its ability to bind to both the RNAP core enzyme and to the DNA template, helping to stabilize the holoenzyme-DNA interaction.
62) Which part of the sigma factor mediates the recognition of the -35 box, and where is it located?
Recognition of the -35 box is performed by Subregion 4.2, which is located within Homology Region 4 near the C-terminus of the polypeptide
63) Describe the competition experiment to confirm interaction:
The goal of this experiment was to prove exactly where Region 4 of the Sigma (σ) factor binds on a promoter. To do this, the researchers used a Competition Filter-Binding Assay.
The Setup: They took Region 4 of the Sigma factor and mixed it with radioactively labeled DNA (the pTac promoter). Because Region 4 binds DNA, the filter became radioactive (high signal).
The Competition: They then added "Competitor DNA" (unlabeled/cold). If the competitor DNA has the "right" sequence, Region 4 will leave the labeled DNA to bind the competitor instead. This causes the radioactivity on the filter to drop.
The Comparison:
When they added competitor DNA lacking a -35 box, the radioactivity stayed high. This means Region 4 didn't care about that DNA; it stayed stuck to the labeled pTac.
When they added competitor DNA containing a -35 box, the radioactivity dropped sharply.
The Conclusion: Region 4 "prefers" the -35 box. If the -35 box is present, Region 4 will bind it. This confirms that Region 4 specifically recognizes the -35 box.
64) What was the primary objective of the competition filter-binding assays involving Region 4 of the sigma factor?
The objective was to determine the binding specificity of Sigma Region 4 and confirm which specific promoter element (the -10 or -35 box) it recognizes.
65) In these competition experiments, what does a decrease in the radioactive signal on the filter indicate?
A decrease in signal indicates that the competitor DNA successfully displaced the labeled promoter DNA, meaning Region 4 has a high affinity for the sequence present in the competitor.
66) How did the presence or absence of the -35 box in competitor DNA affect the experimental results for Region 4?
Competitor DNA containing the -35 box caused a sharp drop in radioactivity (displacement), while DNA lacking the -35 box failed to compete, proving that Region 4 specifically binds to the -35 element.
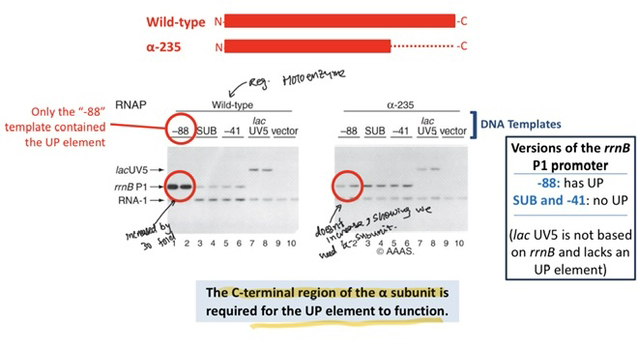
67) Which specific subunit of the RNA polymerase (RNAP) holoenzyme is responsible for recognizing and binding to the UP element?
The α (alpha) subunit of the RNA polymerase is responsible for binding the UP element; experiments show that even purified α subunits can bind to this sequence independently.
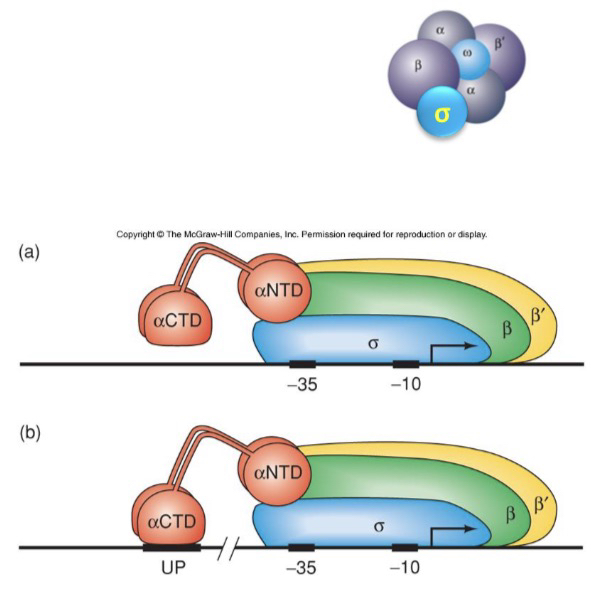
68) Describe the structural domains of the α subunit and how they are connected.
The α subunit consists of two globular domains: the N-terminal Domain (NTD) and the C-terminal Domain (CTD), which are connected by a flexible linker region.
69) Which specific domain of the α subunit is required for the UP element to function in stimulating transcription?
The C-terminal Domain (CTD) of the α subunit is required, as it is the part of the protein that physically anchors the RNAP to the UP element.
70) Define the "Elongation" phase of transcription and identify where it occurs relative to the gene structure.
Elongation is the phase that follows initiation, where nucleotide incorporation and RNA polymerization occur as RNA polymerase (RNAP) travels from the Transcription Start Site (TSS) to the Transcription End Site (TES).
71) Is the rate of RNA polymerase movement constant during elongation? Explain why or why not.
No, the rate is not constant; RNAP often pauses or slows downdue to specific DNA sequences or the influence of auxiliary protein factors that regulate the speed of elongation.
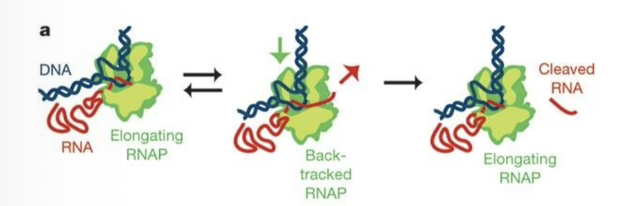
72) Describe the process of "backtracking" and its functional significance during RNA synthesis.
Backtracking occurs when RNA polymerase reverses its directionalong the DNA template, a mechanism primarily used to correct nucleotide incorporation errors (proofreading).
73) Distinguish between the "co-transcriptional" events that occur in prokaryotes versus eukaryotes.
In prokaryotes, the primary co-transcriptional event is the translation of mRNA; in eukaryotes, co-transcriptional events include pre-mRNA processing such as capping, splicing, and 3′ processing.
74) What does the term "nascent RNA" refer to in the context of elongation?
Nascent RNA refers to the newly synthesized, growing RNA strand that is currently being produced by the RNA polymerase as it traverses the gene.
75) What are the two main factors that can cause RNA polymerase to pause during the elongation phase?
The two main factors are specific DNA sequences that inherently confer pausing and auxiliary protein factors that interact with the RNAP to modulate its rate.
76) Compare the extent of DNA melting in the "Open Promoter Complex" versus the "Transcription Bubble" during active elongation.
In the open promoter complex, approximately 10–17 bp of DNA are melted; during active elongation, this increases slightly to about 17–18 bp, forming the transcription bubble.
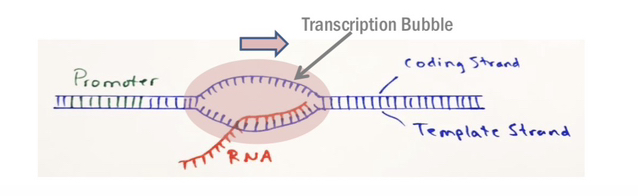
77) Define the "Transcription Bubble" and describe its movement during the elongation phase.
The transcription bubble is the region of melted DNA(approximately 17–18 bp) that is maintained by the RNA polymerase as it traverses the gene, moving continuously along the DNA template.
78) Describe the primary catalytic function of RNA polymerase during the elongation of an RNA transcript.
The catalytic function involves the formation of phosphodiester bonds between the 3′ hydroxyl end of the growing RNA chain and the incoming nucleoside triphosphates (NTPs).
79) Aside from its catalytic role, what enzymatic activity does RNA polymerase possess to ensure the accuracy of the transcript?
RNA polymerase possesses RNase activity, which allows it to perform proofreading functions by removing incorrectly incorporated nucleotides.
80) What are the three primary responsibilities of RNA polymerase while it traverses the length of a gene?
The three responsibilities are moving the transcription bubble, performing catalytic synthesis (forming phosphodiester bonds), and executing proofreading via its RNase activity.
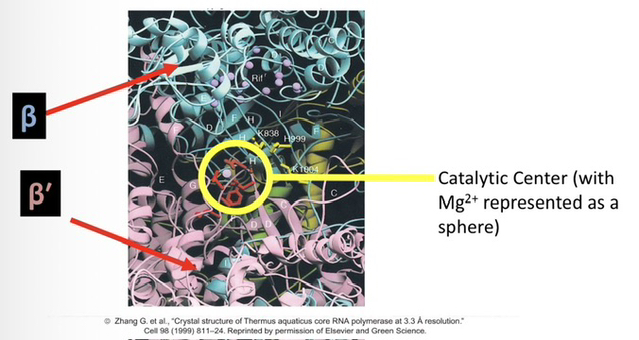
81) Identify the specific amino acid residues and the subunit where the RNAP catalytic center is located.
The catalytic center is located within the β′ (beta-prime) subunitand consists of three aspartate (Asp / D) residues.
82) Explain the role of Magnesium (Mg2+) ions in the catalytic center of RNA polymerase.
One or two Mg2+ ions are coordinated by the aspartate residues to form the active site, where they facilitate the formation of phosphodiester bonds between nucleotides.
83) Describe the physical location of the catalytic center within the overall structure of the enzyme.
The catalytic center is situated within a channel that runs through the enzyme, allowing the DNA template to enter and access the active site for transcription.
84) Define a "ternary complex" in the context of transcription elongation.
A ternary complex refers to the stable association of three different molecules bound together: the RNA polymerase enzyme, the DNA template, and the nascent RNA strand.
85) According to X-ray crystallography of the T. thermophiluselongation complex, what is the length of the RNA-DNA hybrid?
The elongation complex maintains an RNA-DNA hybrid of approximately 9–10 base pairs in length.
86) Describe how the DNA strands are handled by the RNA polymerase as they move through the enzyme during elongation.
Crystal structures reveal that the two strands of DNA feed through separate channels within the RNAP, and only one base pair of the template strand remains unannealed (unpaired) exactly at the active site.
87) Is the rate of transcription elongation constant? Explain the phenomenon of "pausing."
No, elongation is not uniform; RNA polymerase frequently undergoes transcriptional pausing, where it transiently slows down or stops in response to specific sequences or factors.
88) Identify three critical biological functions of transcriptional pausing in bacteria.
Pausing is essential because it coordinates transcription with translation, enables the regulatory process of attenuation, and facilitates proper transcription termination.
89) Why is the coordination of transcription and translation through pausing particularly important in bacteria?
Because these processes are co-transcriptional and translation is generally slower than transcription; pausing allows the ribosome to keep pace with the RNA polymerase.
90) Describe the physical movement of RNA polymerase during the proofreading process.
When an mis-incorporation occurs, RNAP performs backtracking, where it reverses direction along the DNA, causing the 3′ end of the nascent RNA to protrude from the active site.
91) How does RNA polymerase remove incorrectly incorporated nucleotides?
The enzyme utilizes its inherent ribonuclease (RNase) activity to cleave and remove the extruded RNA nucleotides that emerged from the active site during backtracking.
92) What triggers the cleavage of RNA during proofreading, and why is the correct RNA transcript spared?
Cleavage is triggered by the orientation of mismatched RNA nucleotides; RNA that is properly hybridized to the DNA template strand does not adopt this orientation and therefore does not trigger the RNase activity.
93) Describe the step-by-step process of transcription elongation, including the structural components and catalytic mechanism of the ternary complex.
Transcription elongation proceeds through the following coordinated steps and structural features:
Maintaining the Bubble: RNAP maintains a moving transcription bubble of approximately 17–18 bp of melted DNA, ensuring only one nucleotide of the template strand is un-hybridized at the active site at any given time.
Hybrid Formation: A stable RNA-DNA hybrid of 9–10 bp is maintained within the complex to anchor the growing transcript.
Catalysis: Within the β′ subunit, three Aspartate (Asp) residues coordinate with Mg2+ ions to form the catalytic center; this center facilitates the formation of phosphodiester bonds between the 3' end of the RNA and incoming NTPs.
Movement and Regulation: The enzyme traverses the gene, frequently pausing to coordinate with translation or backtracking to allow its inherent RNase activity to proofread and remove mismatched nucleotides.
Strand Management: The DNA strands are fed through separate channels in the enzyme, allowing the template strand to pass through the catalytic center while the non-template strand remains isolated.