DNA structure and replication
The central dogma of molecular biology
Francis Crick
Key features
How we know it today
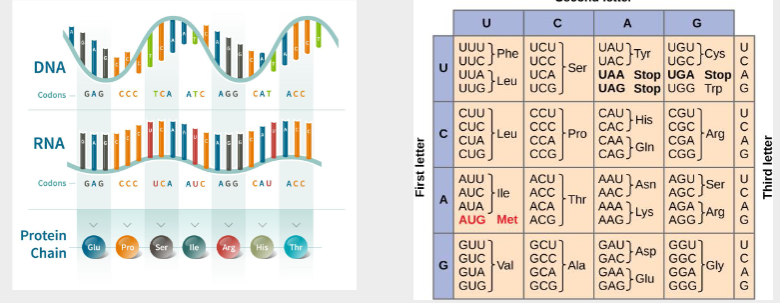
The genetic code
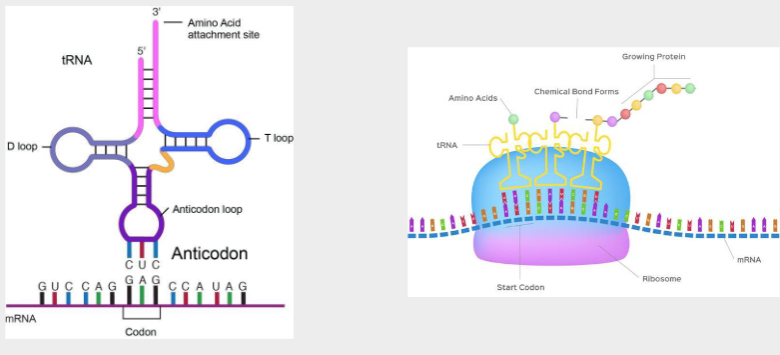
Codons and anticodons
Genes
Coding vs non-coding RNAs and genes
From DNA to proteins
Intro to gene expression
Prokaryotes to eukaryotes and viruses
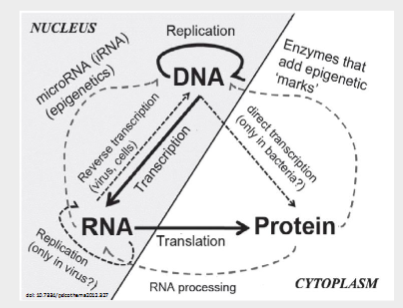
The central dogma of molecular biology
1957- Francis Crick
1953- Watson and Crick
Can go
DNA—> DNA
DNA—>RNA
DNA—> Protein
RNA—>RNA
RNA—> Protein
Can never go
Protein—> Proteins
Protein—>RNA
Protein—>DNA
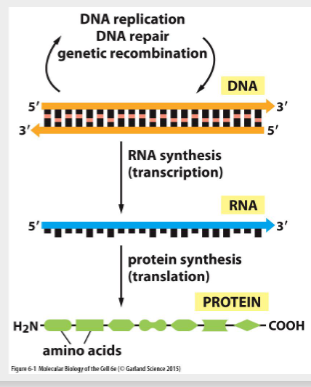
The information in DNA is conserved with high fidelity by DNA replication- ability to contain and transmit all the information needed to produce a whole organism.
During gene expression, the information is passed from DNA to mRNA (transcription)
Then the information passes from mRNA to protein by translation (3 nucleotides contain the information for 1 amino acid)
If one base changes in the DNA a corresponding change will happen in the mRNA and the protein code changes aswell. Proteins can be changed by changing the gene that codes them.
Translation- translating the information from a nucleotide language to an amino acid language
Exons- coding regions of DNA
Introns- non coding regions of DNA
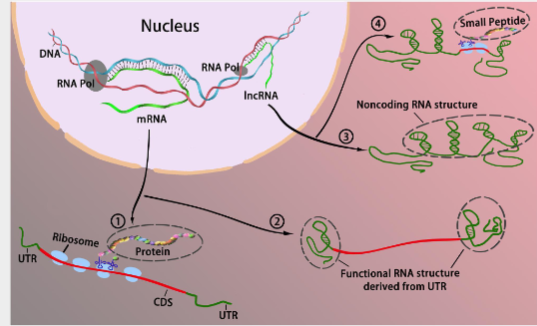
The interchangeable roles between coding and long coding RNAs. Traditionally RNAs could be divided into two categories in accordance with their coding potential, that is, coding RNA and non coding RNAs. Coding RNAs generally refers to mRNA that encodes a protein (1) to act as various components including enzymes, cell structures, and signal transductors. Noncoding RNAs act as cellular regulators without encoding proteins(3). Boundaries blur between coding RNA and noncoding RNA as some coding mRNA can function without translating to proteins via the formation of RNA secondary structure primarily derived from the UTR (2) some incRNA can bind with ribosomes and encode peptides to modulate cellular activities.
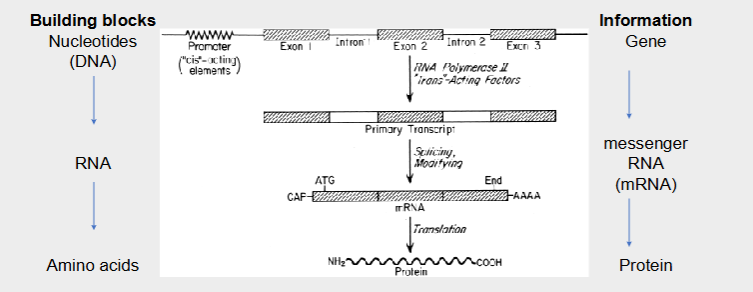
Sequence of base DNA and RNA encodes and transmits all the information required to make the cells and to carry out the very varied functions of different cell types.
DNA structure
The building blocks
Primary structure
Secondary structure
Tertiary structure
Where is genetic information stored?
Prokaryotes vs eukaryotes vs viruses
Nucleus and chromosomes
DNA packaging
Nucleosome and solenoid fibres
Euchromatin vs heterochromatin
DNA the building blocks
Deoxyribonucleotides (nucleotides) are formed by
A five carbon sugar
A phosphate group
A nitrogenous base A,T,G,C
Primary structure
Adjacent nucleotides are linked by phosphodiester bonds between the 5’ phosphate P and the 3’ hydroxyl group OH
A pairs with T - 2 hydrogen bonds
G pairs with C- 3 hydrogen bonds
Hydrogen bonds form between the bases maintaining the two strands of DNA together
The two strands of DNA run in opposite directions 5’ to 3’ and 3’ to 5’
Nucleotides are added by DNA polymerase in the direction of the strand
The final strand will have a P group at the 5’ of the first nucleotide an a OH in the final.
Complementary base pairs in the DNA double helix, the shapes and chemical structures of the bases allow the hydrogen bonds to form efficiently only between A,T and G,C due to the atoms that can form hydrogen bonds
Secondary structure
Double helix
1.5 turns od DNA double helix
Each DNA turn is made up of 10.4 nucleotide pairs and the centre to centre distance between adjacent pairs is 0.34nm
The coiling of the two strands around each other create two grooves, minor and major.
Nucleotides linked together covalently by phosphodiester bonds that join the 3’ OH to the 5’ Oh of the next sugar
Each strand has chemical polarity
Tertiary structure
A and B (right handed double helix) and Z (left handed helix) form DNA
B form is the predominant structure
Where is DNA stored
Eukaryotic cells
DNA stored in the nucleus in linear for
DNA molecules stored and packaged into chromosomes
A chromosome consist of single enormously long linear DNA molecule and proteins
DNA and protein is called chromatin
Chromosomes centromere- two telomere, and a replication origin
Telomere- ends specialised structures required for the stability of the chromosome ends, containing repetitive DNA
Chromosomes
The genome of an organism encompasses all of the genes of that organism
Gene- distinct sequence of nucleotides forming parts of a chromosome, order of nucleotide in gene determines order of AA in a polypeptide or a ribonucleotide in an RNA molecule
Genes are contained in chromosomes
Chromosome are thus structures within cells that contain cells that contain hundreds and thousands of genes
Prokaryotic chromosomes
Circular doble stranded molecule “bacterial chromosome”
Circular DNA is packaged into nucleoid, 50 loops
DNA negatively supercoiled- twisted upon itself
Complexed with several DNA-binding proteins
Packing of nucleosomes
First level of organisation
Second level of organisation is the coiling of the series of the nucleosomes into a helical array to constitute the nucleosome fibre
histones are basic proteins 25% of their AA are lysine or arginine so positively charged AA side chains
Packaging of solenoid fibres
Third level of the organisation is the packaging of the fibre itself
Attached to chromosome scaffold
Euchromativ vs heterochromatin
Lightly stained= euchromatin-active sequence
Darkly stained= heterochromatin- inactive sequence
RNA primary structure
5 carbon sugar
Phosphate group
A,U and G,C
secondary structure- 4 leaf clover
Genome size
Measured as either the number of base pairs or the mass of DNA in a haploid chromosome.
10^6 bp= 1Mbp =1fg (femtogram) DNA
C-value is the amount of nuclear DNA in the unreplicated gametic nucleus, irrespective of the ploidy level of the species
C-value range from less that 10^6 bp for mycoplasma (simple bacteria) 10^11 for plants and amphibians
DNA in bacteria has little wasted coding sequence 90% of DNA present is coding.
In higher eukaryotes only about 1% of DNA present is needed to code for proteins
DNA replication
Needed each time cell divides
Two resulting daughter cells must contain the same genetic information as the parent cell
Double-stranded DNA molecule is copied to produce two identical DNA molecules
During replication each DNA strand acts as template for the production of new DNA strand
Complementary base pairs
Main components
A DNA template strand to be copied- original DNA strand
A DNA polymerase- Enzyme that synthesises DNA by joining nucleotides together
4 deoxyribonucleotides dNTPs
Physiological pH and ionic conditions
A primer - piece of DNA or RNA with a free 3’ hydroxyl, hybridised to the template
Helicase- enzyme that uses ATP to move along DNA strand and as such open the DNA helix to allow replication to occur
Topoisomerase
Replication fork- Localised region of replication, where the two strands are separated allowing replication to occur
Leading strand- newly synthesised strand that is synthesised in a continuous manner from DNA polymerase in 5’ to 3’
Lagging strand- strand is synthesised in small fragments 5’ to 3’ direction
3 main steps
Initiation-involves recognition of the position on the DNA molecule i.e where replication will begin
Elongation- concerns the event on the replication fork where the parent polynucleotides are copied
Termination-which is not very well understood occurs when the parent molecule are completely copied. In E.coli genome seven terminator sequences. have been identified, each one acting as the recognition site for a sequence-specific DNA-binding protein called Tus. Little is known on termination of DNA replication in eukaryotes
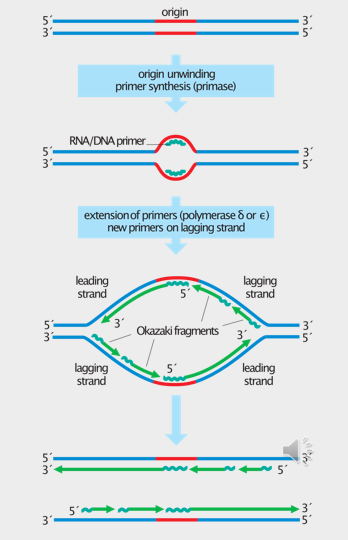
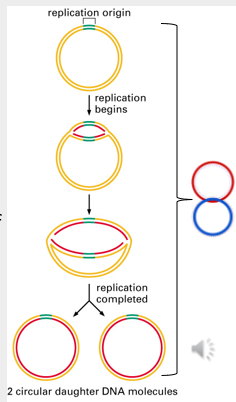
Initiation of DNA replication in prokaryotes
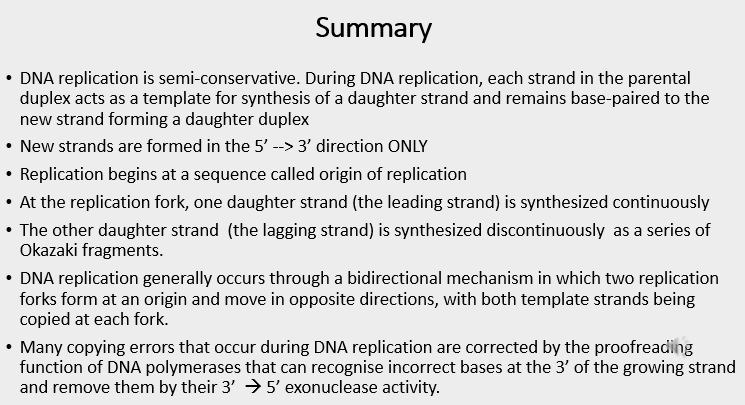
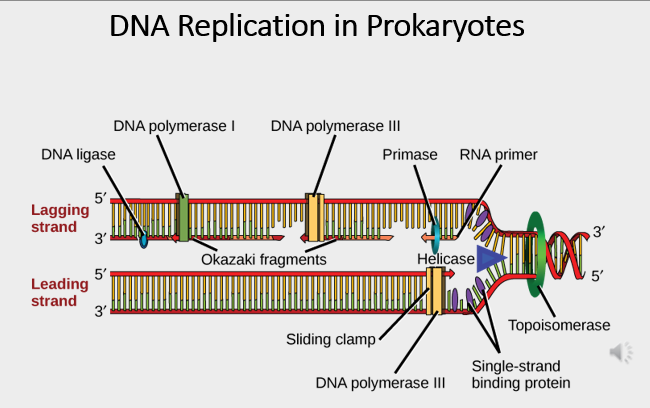

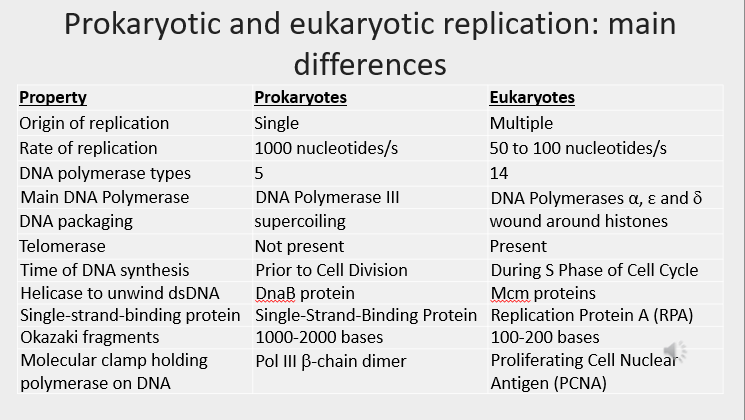

DNA mutations
changes in the base sequence of DNA
Can be neutral beneficial or harmful
Essential for generating variants for Darwinian selection
Molecular and specied driven evolution