Identification of Bacterial Species by 16S rRNA gene sequencing
16S ribosomal RNA PCR
16S rRNA
all bacteria contain 16S rRNA
9 variable regions interspersed among conserved regions along 16S
sequence of variable regions can be determined → identification of species
Application of 16S rRNA gene sequencing
clinical diagnosis
NIH Human Microbiome Project
characterise microbial communities found at multiple human body sites
look for correlations between changes in microbiome and human health
metagenomics: bacterial diversity in environment
Sterilisation
Heat
Steam sterilisation
performed in autoclave
used for sterilising agar media or other equipment
121°C for 15-30 min at 15 psi
╳ solutions containing organic solvent
autoclave tape is used to monitored autoclaving condition
white: before sterilisation
black: after sterilisation
saturated steam under pressure→ denature protein and enzyme at high temp
Dry-heat sterilisation
performed in oven
used for sterilising metalware
╳ suitable for plasticware (melt)
160°C for 3 hr; 170°C for 1 hr; 180°C for 30 min
Filtration
pore size of filter: 0.22μm (smaller than most micro-organism) → retain microbes on membrane
used for heat labile (不穩定) or flammable reagent
╳ applicable to viruses as too small
UV irradiation
DNA or RNA damage → inactivate and inhibit growth of bacteria
wavelengths: 200-280 nm
performed in a biological safety cabinet
Gram stain
Purpose
classify bacteria into Gram positive, Gram negative and Gram variable bacteria
visualise the shape and arrangement of the bacterial cells
Principle
Differences between Gram positive and Gram negative
Gram positive bacteria
single membrane
thick peptidoglycan layer
lower lipid content of cell wall
Gram negative bacteria
double membrane
thin peptidoglycan layer
higher lipid content of cell wall
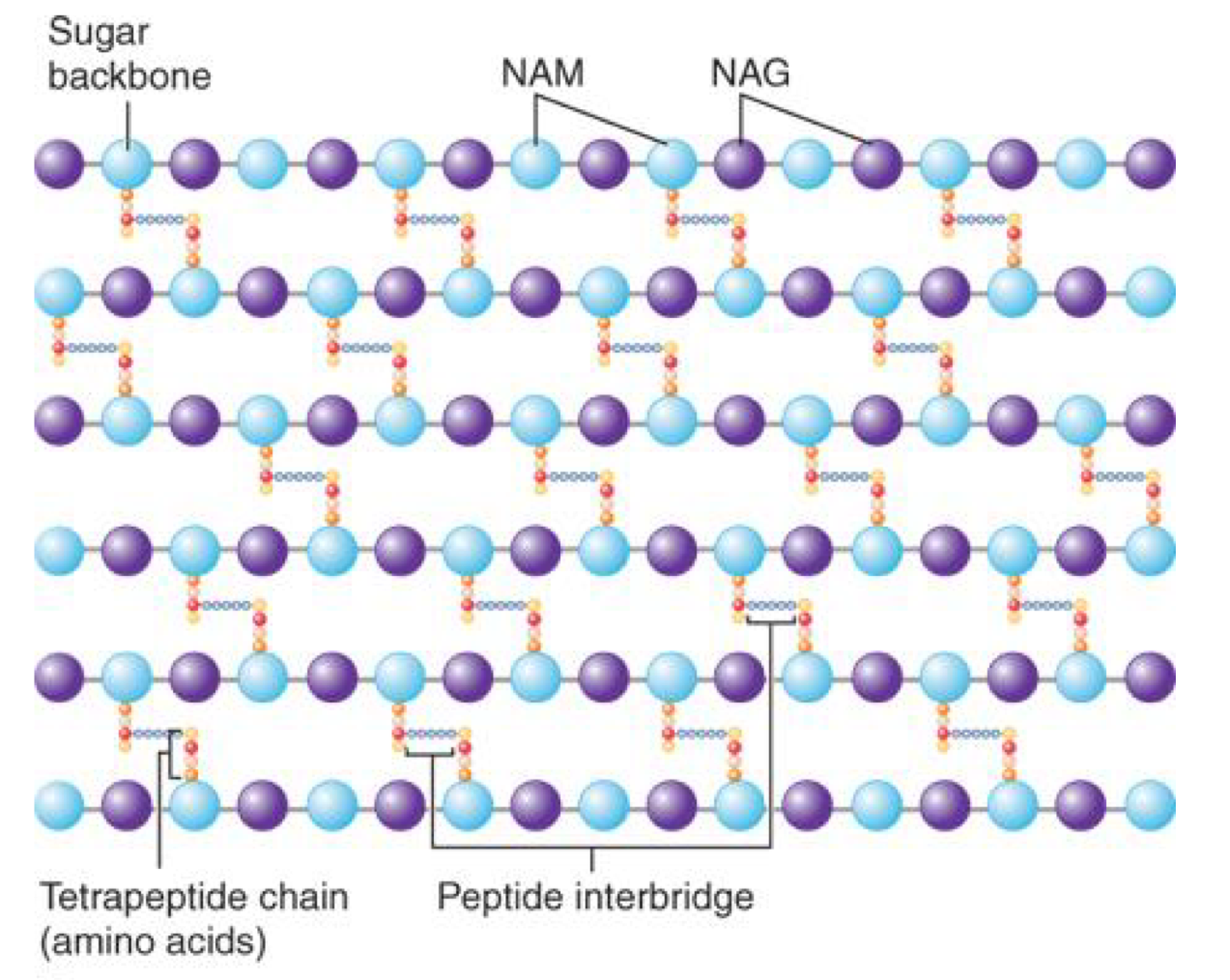
Peptidoglycan
a carbohydrate backbone of alternating units of NAG (N-acetylglugosamine) and NAM (N-acetylmuramic acid) residues linked to peptide
peptide cross bridges linking the tetra peptide on a NAM to an amino group of a terapeptide on a neighbouring NAM
→ forming a larger polymer network
 Principle of Gram stain
Principle of Gram stain
stain with crystal violet dye
then add Gram’s iodine solution (iodine & potassium iodide)to form a complex between crystal violet and iodine
complex is a larger molecule than iodine and crystal violet and insoluble in water
use ethyl alcohol or acetone to decolorize the sample
dehydrate the peptidoglycan layer → shrinking and tightening the layer
large crystal violet-iodine complex: ╳penetrate the tightened peptidoglycan layer
→ trapped in cell in Gram positive bacteria
outer layer of Gram negative bacteria is degraded
→ thinner peptidoglycan layer ╳ retain the complex→ colour lost
counter stain using weakly water soluble safranin (red)
Gram positive bacterial cells remain purple (safranin is lighter than crystal violet)
Gram negative bacterial cells are stained red
Using light microscope
magnification: 10X (eyepiece) & 100X (objective)= 1000X
oil immersion lens
oil has the same refractive index as glass
more light is gathered→ brighter image and higher resolution
only suitable for specific objective lens (e.g. 100X)
Morphology of bacteria
Shape
coccus (sphere)
bacillus (rod)
spiral
vibrio
spirillum
spirochete
Arrangement
diplo-
strepto- (forming a line)
tetrad
sarcinae
staphyl- (a cluster)
Determination of nucleic acids concentration by spectrometry
Beer-Lambert Law
Formulas
T = I/I0
A (Absorbance or extinction or optical density) = log(1/T) = log(I0/I) = ελcℓ
T = transmittance
I0 = intensity of incident light
I = intensity of transmitted light
ελ= molar absorbance coefficient at wavelength λ( M^-1·cm^-1 or dm^3·mol^-1·cm^-1)
0.02 for double stranded DNA
ℓ = path length of sample
1cm for a cuvette
0.05 cm for microplate ot μDrop plate
c = concentration of absorbing solution ( M or mol·dm^-3)
Deviation
Beer-Lambert law only valid for low concentration
higher concentrations → association of molecules (they have different light absorption characteristics)
Determining nucleic acid concentration
measure absorbance at 260 nm, 280 nm and 230 nm
determine A260/A280 and A260/A230 ratio to check purity of nucleic acid
purity of nucleic acid
A260/A280
A260/A280 ≈ 1.8 for pure DNA
A260/A280 ≈ 2.0 for pure RNA
lower A260/A280 indicates protein contamination
ratio also depend on sample pH, wavelength accuracy and nucleic acid composition
A260/A230
A260/A230 > 2.0 for pure DNA or RNA
A260/A230 < 2.0 → possible contamination with phenol, guanidium salt