8. DNA repair
Major types of DNA repair → Various DNA repair pathways have evolved to maintain genomic stability
Direct reversal
→ Photoreactivation (PR)
Proof-reading
Excision repair
→ Mismatch repair (MMR)
→ Base excision repair (BER)
→ Nucleotide excision repair (NER)
Double-strand break repair
→ Homologous recombination (HR)
→ Single strand annealing (SSA)
→ Nonhomologous DNA end joining (NHEJ)
→ Alternative End Joining (Alt-EJ)
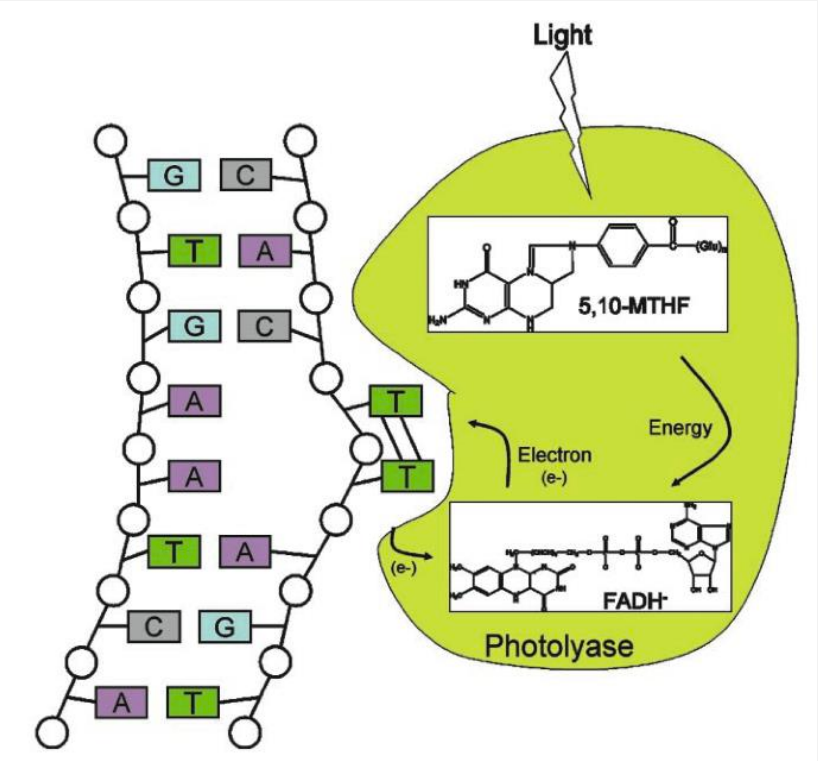
Photoreactivation (PR)
Photolyase binds to DNA containing a pyrimidine dimer and flips the dimer out into the active site pocket.
The cofactor methenyltetrahydrofolate (5,10-MTHF) absorbs a photon of blue light.
The excitation energy is transferred from 5,10-MTHF to the catalytic cofactor FADH-
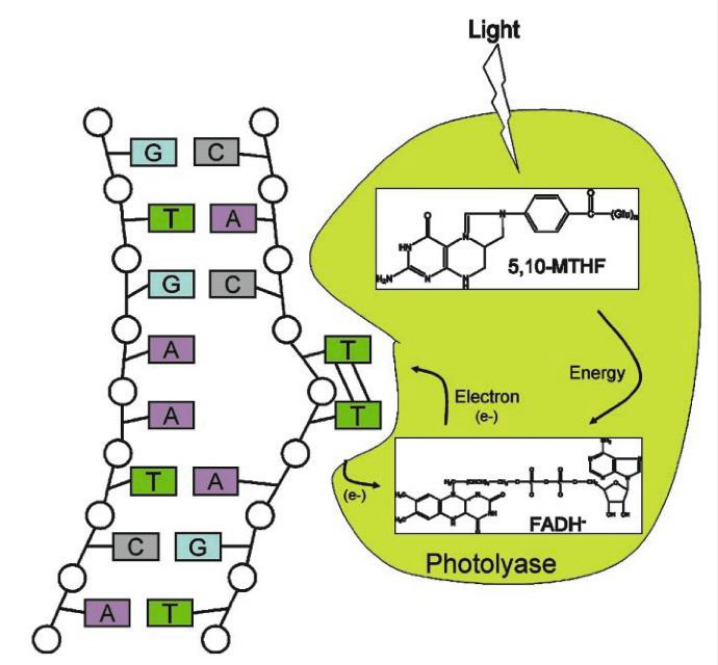
The excited state FADH- transfers an electron to the dimer splitting it.
The electron returns to the flavin radical, regenerating FADH- .
Damage sensing and repair require a single protein.
Photolyases are found in many prokaryotes and eukaryotes but not in placental mammals.
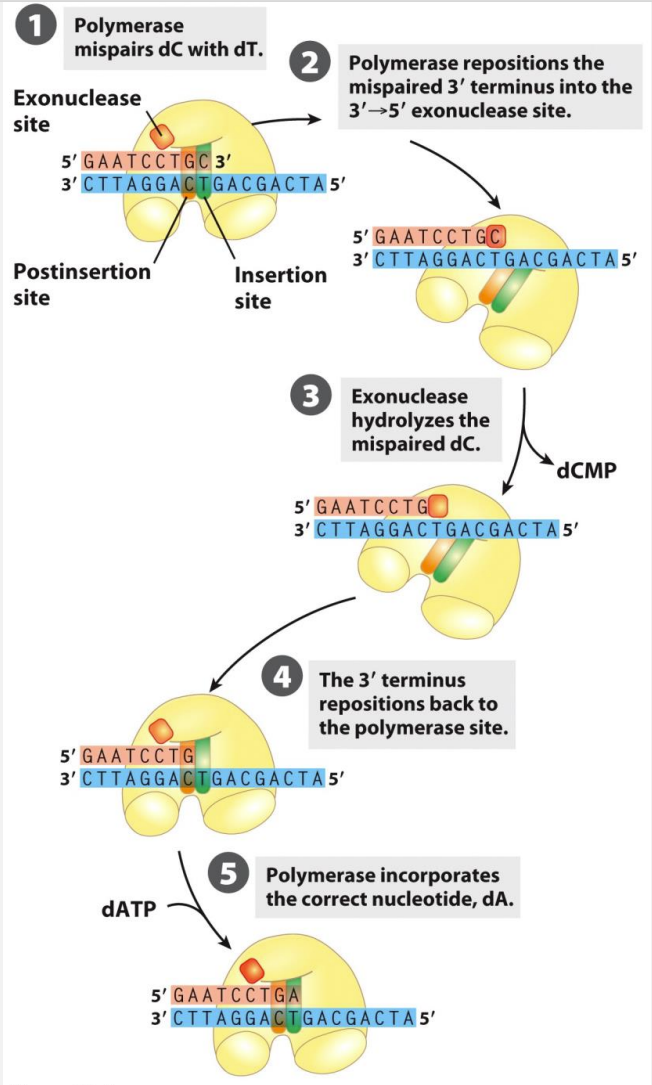
Proof-reading
Process by which DNA polymerase checks for errors during DNA replication and corrects them. It involves the ability of DNA polymerase to recognize and remove mismatched nucleotides that have been incorrectly incorporated into the newly synthesized DNA strand. This proofreading activity helps maintain the accuracy of DNA replication and ensures the fidelity of the genetic code.
Major replication DNA polymerases have 3’→5’ exonuclease activity
3’→5’ exonuclease activity has a ‘proof-reading’ function – removing misincorporated nucleotides
Mismatch repair (MMR)
Cellular process that corrects errors in DNA replication, specifically mismatches between the newly synthesized DNA strand and the template strand. It involves the removal and replacement of the incorrect nucleotides, ensuring the integrity of the genetic information.
Types of DNA damage repaired by MMR:
mismatch
insertion
deletion
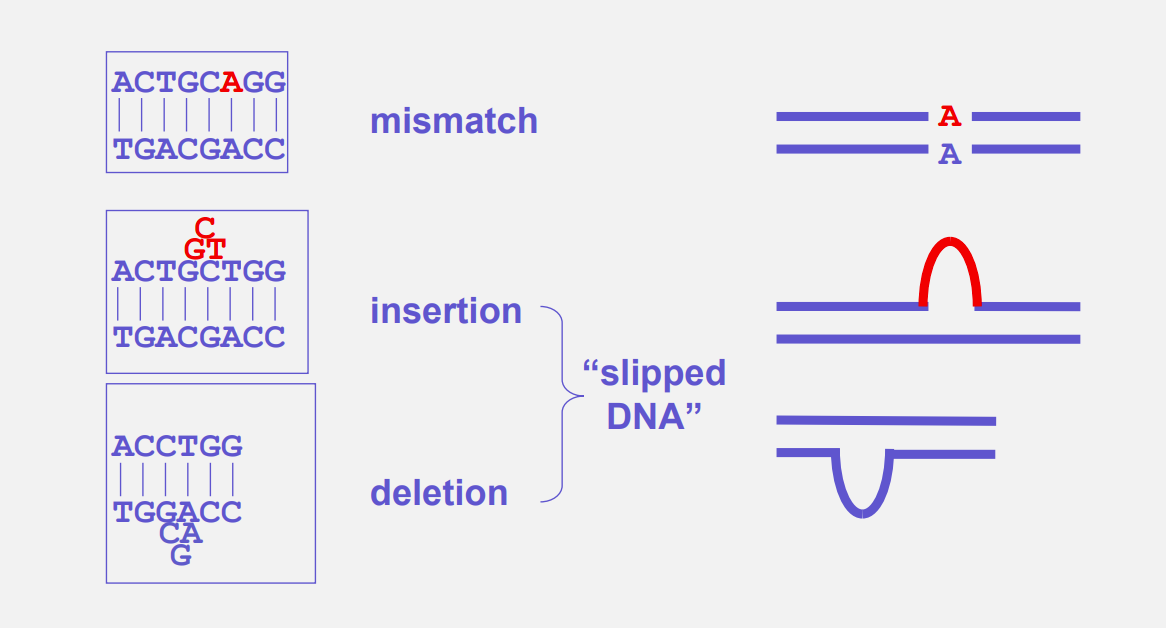
The mismatch repair (MMR) pathway in humans involves several steps:
Recognition: Mismatched bases are recognized by the MMR proteins, such as MutSα and MutSβ, which scan the DNA for errors.
Excision: Once a mismatch is detected, the MutS proteins recruit MutL proteins to form a complex. This complex then recruits exonucleases, which remove the section of DNA containing the mismatched base.
Resynthesis: After the excision, DNA polymerase fills in the gap with the correct nucleotides, using the undamaged DNA strand as a template.
Ligation: DNA ligase seals the nick in the newly synthesized DNA strand, completing the repair process.
Surveillance: The repaired DNA is checked again by the MMR proteins to ensure the accuracy of the repair.
Base excision repair (BER)
DNA repair mechanism that corrects small, non-bulky DNA lesions, such as damaged or mismatched bases. It involves the removal of the damaged base by a specific DNA glycosylase enzyme, followed by the excision of the sugar-phosphate backbone and subsequent replacement with the correct nucleotide sequence.
BER plays a crucial role in maintaining the integrity of the genome and preventing the accumulation of mutations.
An overview of the steps of BER:
Base removal by DNA glycosylase
Removal of abasic nucleotide by AP endonuclease
Resynthesis of DNA
Ligation
Human BER pathways → two main mechanisms:
Short-patch BER (1 nucleotide is replaced)
Long-patch BER, (several, 2-14, nucleotides are replaced)
Nucleotide Excision Repair (NER)
DNA repair mechanism that removes and replaces damaged nucleotides in the DNA strand. It is responsible for repairing a wide range of DNA lesions, including those caused by UV radiation and certain chemicals.
NER involves the recognition and removal of the damaged DNA segment, followed by the synthesis and ligation of a new DNA strand to restore the integrity of the DNA molecule.
NER is well characterised in humans in major part because the human disease xerodema pigmentosum (XP) results from the mutation of NER components.
Two types:
Global genome NER (GG-NER) this pathway scans the genome for helix-distorting lesions and repairs them
Transcription-coupled NER (TC-NER) removes transcription-blocking lesions
GG-NER and TC-NER differ in the identification stages but share excision and repair mechanisms.
Damage recognition in GG - NER:
Damage bound by a complex that includes the XPC, Centrin 2 (CETN2) and RAD23B proteins. XPC is considered main sensor. In vitro it binds to a range of abnormal DNA structures.
UV-DDB binds some types of damage (UV-induced lesions) and promotes XPC binding
Recruitment and action of the TFIIH complex (10 proteins), including the XPB and XPD helicase subunits, leads to the formation of an open duplex around the damage site.
Damage recognition in TC - NER:
Initiated by lesion -stalled RNA Pol II
Cockayne syndrome proteins CSA and CSB are recruited.
CSA and CSB, core NER factors and several TC - NER -specific proteins, including UVSSA, USP7, XAB2, HMGN1 form a complex
Complex backtracks (translocates 5’ to 3’ along DNA)
Excision and repair:
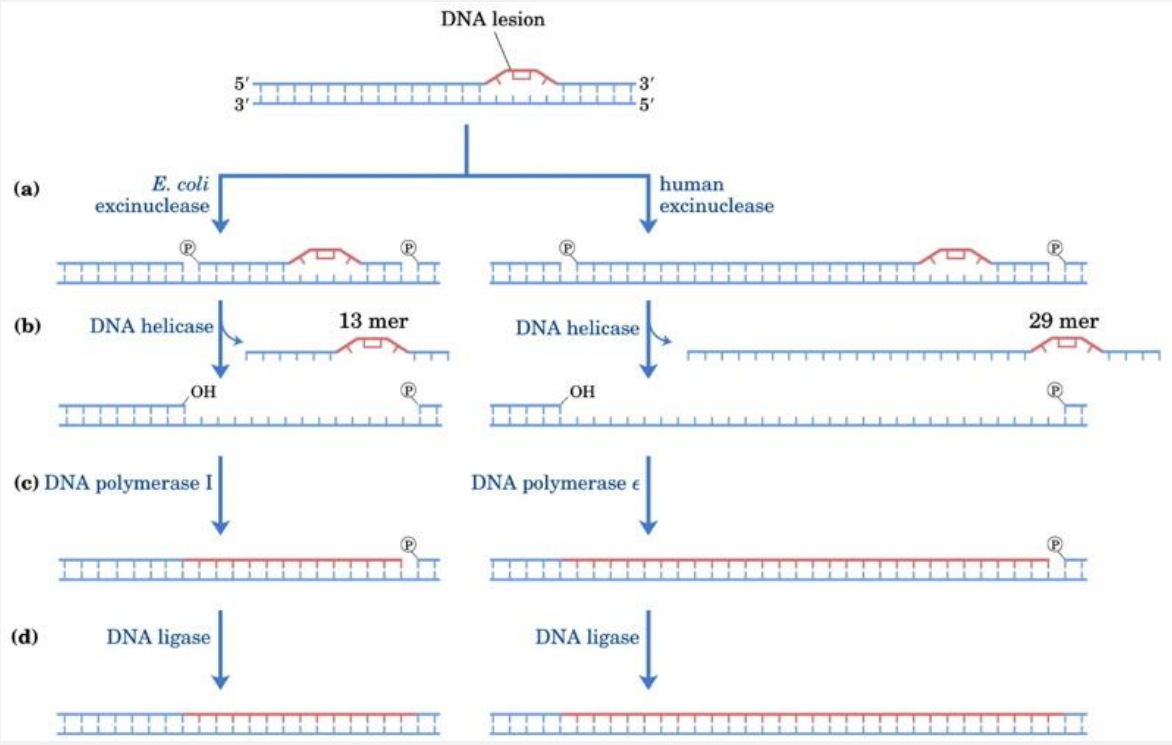
The damaged strand of the open duplex is a substrate for XPG structure -specific nuclease, which cleaves 3’ to the damage, and the XPF:ERCC1 nuclease dimer, which cleaves 5’ to the damage.
XPG cuts 5 to 6 phosphodiester bonds away from the damage and XPF:ERCC1 cuts 22 to 24 phosphodiester bonds away.
This results in the removal of an ~30 residue single -stranded oligonucleotide containing the damage.
The corresponding gap in the double - stranded DNA is filled by DNA synthesis involving RFC, PCNA and DNA polymerases δ, ε or κ and by ligation by DNA ligase I or XRCC1 –DNA ligase 3
What single protein can repair thymine dimers? → Photolyase
What multi-protein process can repair thymine dimers? → Nucleotide Excision Repair (NER)
DSBs & genome stability
DSBs that persist in genome are very dangerous for cell viability – essential genes may be broken or lost.
This observation is supported by essential nature of Lig4/XRCC4, ATR, BRCA1 and BRCA2.
Partial loss of function of some proteins leads to predisposition to cancer in humans, e.g. ATM, BRCA1, BRCA2, NBS1 and LigIV/XRCC4.
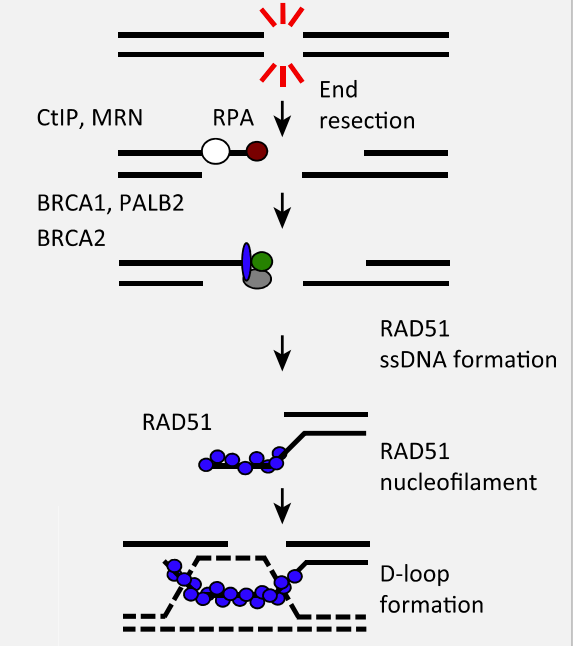
Homologous recombination (HR)
DNA repair process that occurs between two similar or identical DNA molecules, it involves the exchange of genetic material between these molecules, resulting in the formation of new combinations of genes.
This process plays a crucial role in DNA repair, genetic recombination, and the generation of genetic diversity.
The distinguishing feature of HR is that information lost from the broken DNA is retrieved from the homologous duplex. HR thus requires a copy of DNA sequence to undertake repair. This can be difficult to find these pathways occurs primarily in the mid-S and mid-G2 phases of the cell cycle, when the sister chromatid is available.
Three steps:
Strand invasion
Branch migration
Holliday junction resolution
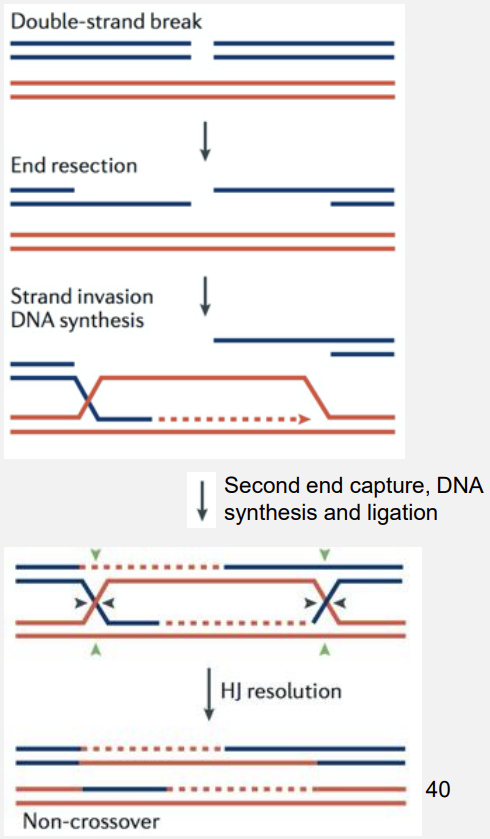
Ends are processed by CtIP and the Mre11-Rad50-Nbs1 (MRN) complex.
Strand invasion is initiated by the Rad51 (or Dmc1) recombinase (eukaryote homologue of RecA)
Single Strand Annealing (SSA)
DNA repair mechanism that occurs in cells, it involves the recombination of two DNA sequences that have similar but not identical sequences. During SSA, the single-stranded regions of the DNA molecules align and form a double-stranded DNA molecule through a process called annealing. This repair mechanism is important for fixing DNA double-strand breaks and maintaining genomic stability.
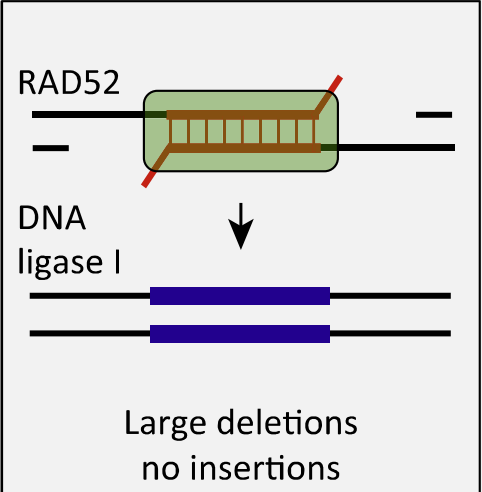
No homologous sequence is required.
No strand invasion.
Independent of Rad51.
End is substantially resected and regions between homologies are lost.
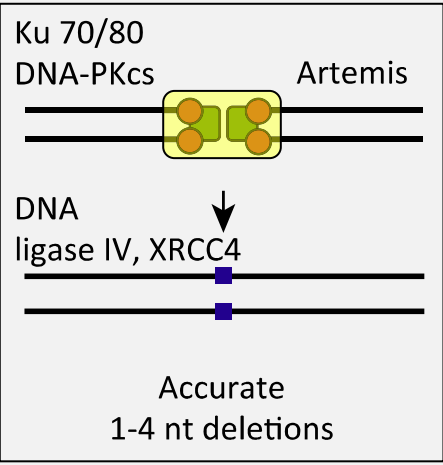
Nonhomologous DNA end joining (NHEJ)
DNA repair mechanism that repairs double-strand breaks in DNA by directly joining the broken ends together. It does not require a homologous template for repair and can result in small insertions or deletions at the site of repair.
NHEJ simply links ends of a DSB, without any template, so can occur at any point in cell cycle. Dominant in G0/G1 and G2.
Steps:
Ku heterodimer binds two termini
End processing by Artemis
MRN holds the ends in close proximity
XRCC4/Ligase IV complex ligates the ends
The XRCC4/Ligase IV complex will ligate the ends irrespective of whether they come from the same chromosome.
In addition, while HR is “ error-free”, NHEJ can add or lose nucleotides.
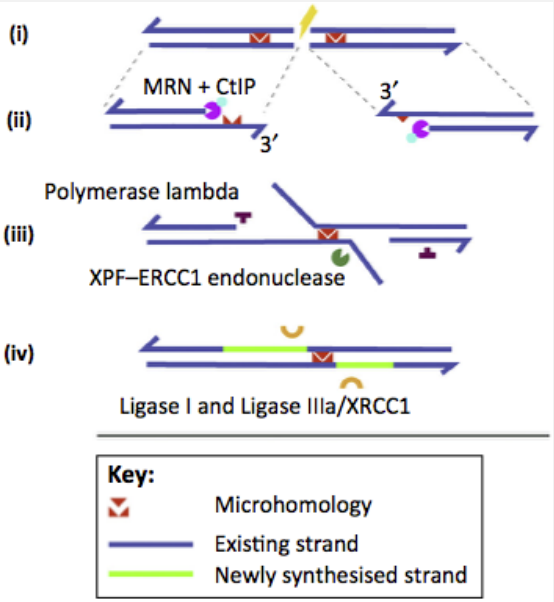
Alternative End Joining (Alt-EJ)
DNA repair mechanism that occurs when double-strand breaks in DNA are repaired by joining together non-homologous DNA ends. It is an error-prone process that can lead to the loss or gain of genetic material.
‘Alternative NHEJ pathway’
Steps:
5’ to 3’ resection by the Mre11– Rad50–Nbs1 (MRN) complex and CtIP (ii)
annealing of the microhomologous sequences, trimming of the noncomplementary 3’ flaps, gap filling (iii)
Ligation (iv)
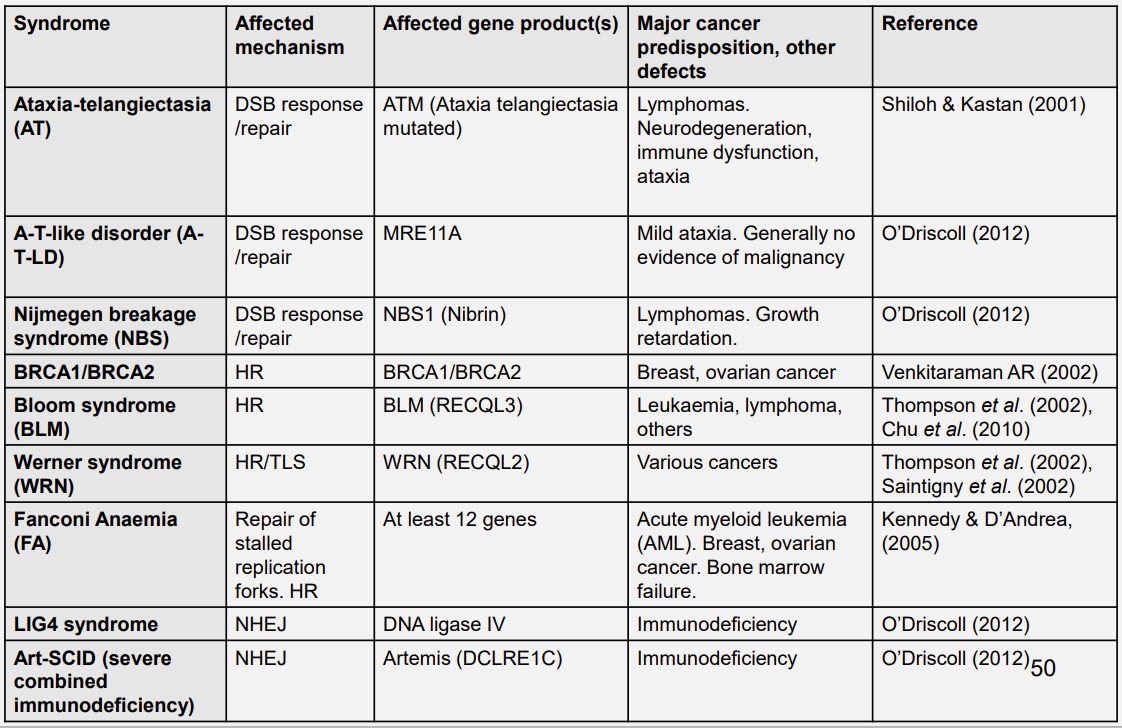
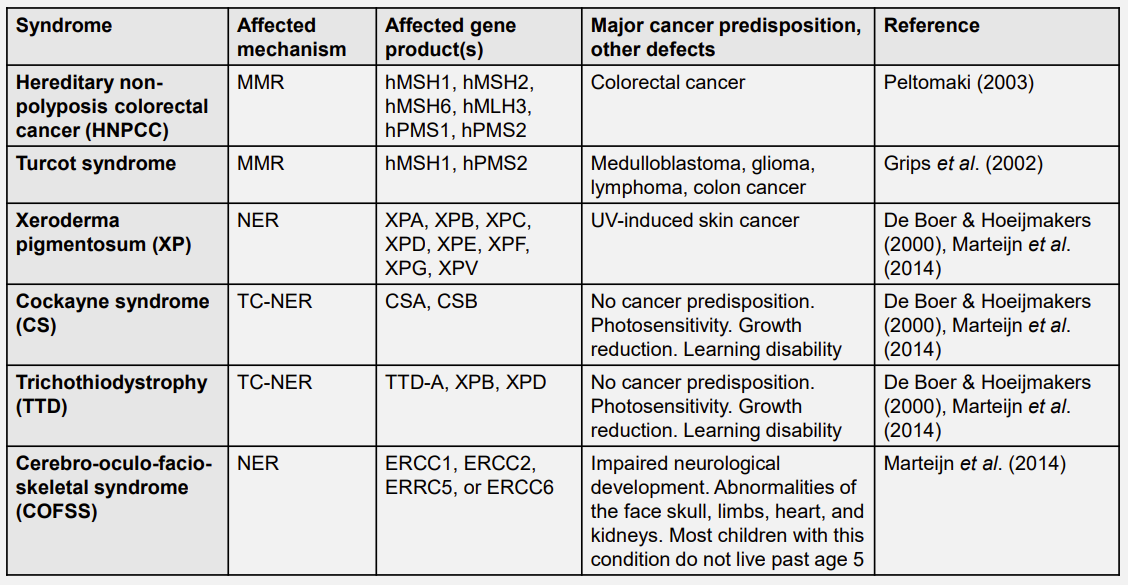
Human syndromes associated with defective genome repair