MCB 150 Chapter 7.4-7.5
7.4 Analyze the process of transporting molecules across the nuclear envelope
The nuclear envelope separates the nucleus from the rest of the cell. The structure is supported by an internal fibrous nuclear lamina and bounded by 2 membranes.
The nuclear envelope is punctured with openings that are approximately 60 nm in diameter.
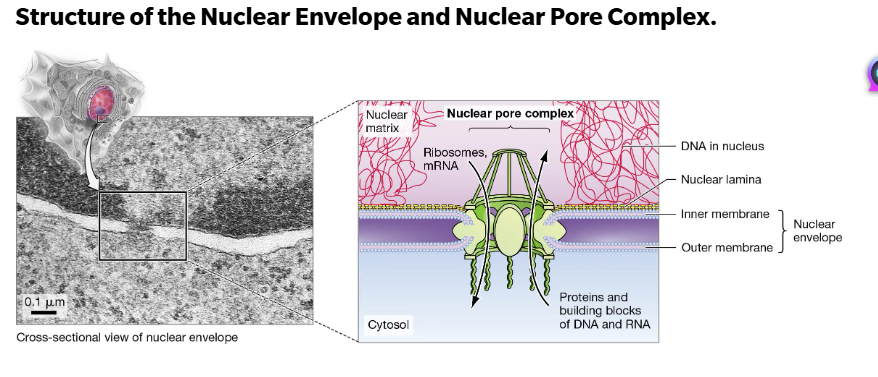
Chromosomal DNA does not travel but remains in the nucleus as long as the nuclear envelope is intact.
Several types of RNA molecules are produced in the nucleus, each different in size and function. These must be transported from the nucleus to the cytosol, where protein synthesis takes place. How is it regulated and directed?
Small molecules, like nucleotides, can freely diffuse through nuclear posed into the nucleus along their concentration gradients. The import of larger molecules, such as proteins and RNA, appears to be more selective: some are allowed to enter, others are excluded.
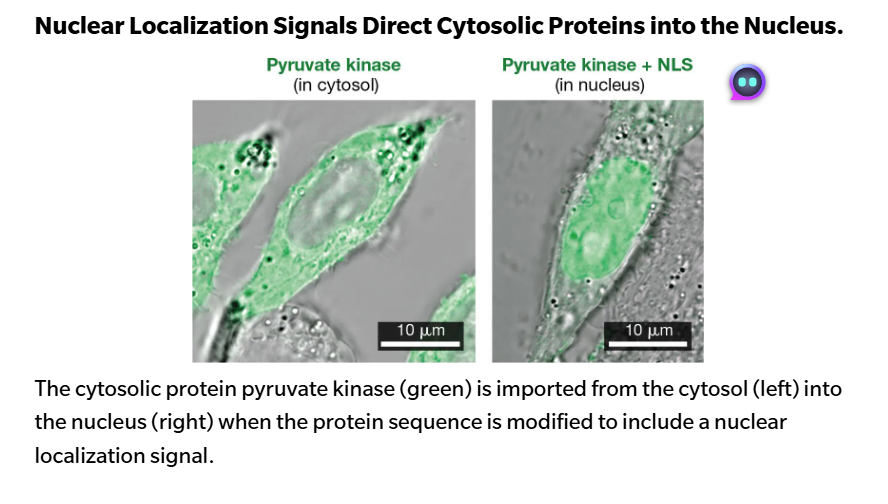
Nuclear proteins contain a “code” that marks them for transport through the nuclear pore complex. It allows them to pass into the nucleus. This code is in their amino acid sequences that directs their transport. This common sequence is called the nuclear localization signal (NLS).
The site of translation determines the fate:
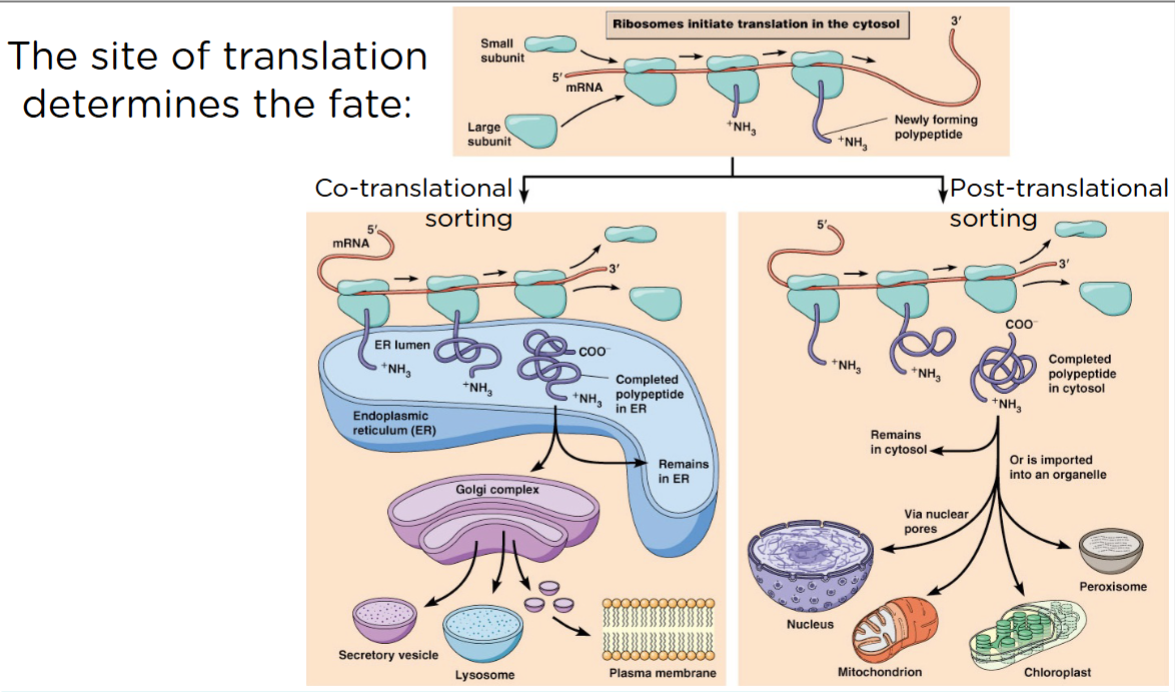
What type of ribosome is translating you, dictated your only possible locations. A ribosome is going to continue to do translation until it is told otherwise, or it has reached a stop codon.
On the right, you see a lot of proteins that aren’t told to take a time out or pause. The starter on cytowali ribosomes and they finish on ribosomes that are still just floating around in the cytoplasm. The protein reached a stop codon and terminated in the cytoplasm. If this is how the protein were made, then there are only certain places in the cell that you can even possibly end up. One is staying in the cytoplasm. You could get taken to the nucleus, the mitochondria. You CANNOT get into the Golgi apparatus if you were translated in a free cytosolic ribosome. So what about the proteins that are supposed to work in the cytoplasm? Do they need to present a signal? No, so they just don’t.
What if you are supposed to get into the nucleus? You need to tell the cell that it needs to go to the nucleus.
After a few amino acids have poked their heads out, they may need a time out. They pause and they go to the ER and pick up where they left off (signal is given). You take a time out on translation, you don’t dissociate your ribosomal subunits. This protein gets pushed into a channel in the rough ER. If this is how you were made, you were not synthesized fully on a free cytosolic ribosome, you were synthesized on a membrane bound ribosome (rough ER membrane), and where can you end up? You are never going to see the inside of a nucleus or the mitochondria. You could end up staying in the rough ER, you could go to the smooth ER, you could go to the golgi apparatus, can get exported (secreted), or a transmembrane protein, or in a lysosome.
When did we determine to move the protein somewhere else? Was it after translation was finished (post translational sorting)? This is what happens to the proteins that were made on free cytosolic ribosomes. On the other side where you need to take a time out, that is called co-translational sorting.
Only talking about eukaryotes
If we don’t say otherwise, you stay where you are
The absence of a signal gives enough information
Traffic through the nuclear pore complex:
Nuclear pore complexes have the ability to open really wide. Don’t want to be open all the time, so it closes but it can never close all the way. bc of this, small molecules can get through the nucleus via passive diffusion, where there didn’t need any energy and did not have to be told that they can or cannot get through (less than 20 kDa). What if you are bigger (more than 60 kDa)? (don’t worry about in between those numbers). Active transport is for vesicles more than 60 kDa and it uses energy from GTP not ATP.
What needs to get into the nucleus? Polymerases, histone proteins, transcription factors, etc. How do they identify that they are not supposed to stay in the cytoplasm?
They present a signal that takes them to the nucleus. signal - sequence of amino acids that is compatible in its shape with some other molecule.
Signaling process: present a signal, something recognizes that signal, and then you take the appropriate action.
How to proteins indicate that they are supposed to go into the nucleus?
Nuclei have the ability to open pretty big and can never close all the way.
Passive diffusion
If you are a bigger than average - active transport which uses energy to make it happen. Energy in this comes from GTP.
what is the signal to make the protein go in the nucleus
Proteins that need to get in the protein - transcription factors and many more - they present a signal (sequences of amino acids)
In a signal process:
Theres a signal, someone recognizes it, and then it takes action
What keeps cytoplasmic proteins out of the nucleus?
only the appropriate proteins (that need to get into the nucleus) have the right signal
That signal is called a nuclear localization signal or NLS
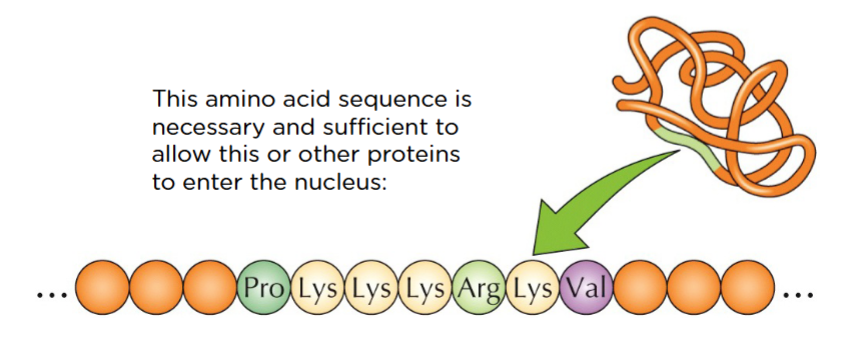
Lys and Arg are really positive - rich in basic amino acids. this is the signal
Another NLS was later found to be bipartite:
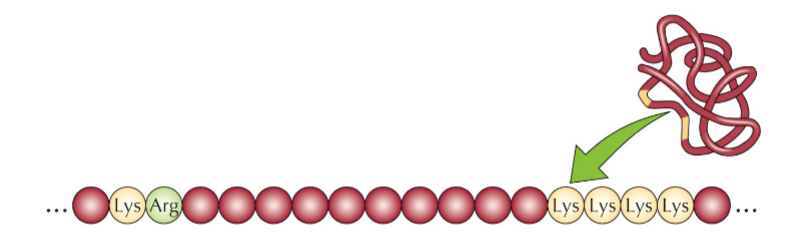
Other NLS’s are similar to the 2 described above and still other NLS’s are created by proper folding
This is in two different sections. both are positively charged. Proteins can fold so that it creates the NLS
What keeps nuclear proteins out of the cytoplasm?
Only the appropriate proteins have the right signal
that signal is called a nuclear export signal or NES
What keeps proteins in the nucleus? You have to have a signal that says get me out of the nucleus. If you don’t have that signal, you don’t get out of the nucleus.
Lot of Leu - NES
Nuclear protein import and export:
Whose job is it the recognize the NES or NLS? Two different signals so two different molecules. The ones that assist in getting into the nucleus and recognize the NLS are called importins. An exportin is a molecule that recognizes an NES and their either do or do not have affinity for the protein that needs to be moved.
Importin - purple protein out in the cytoplasm and needs to get into the nucleus. How do we know that? the round nose represents the NLS. Importin recognizes the signal. Importin binds to the NLS. Anything that is with the NLS or NES can go through the nuclear pore complex. Once you get into the nucleus, we need something to make the importin let go of the protein. That something is called a ran-GTP. ran-GTP binds to the other side of the importin, so the importin lets go and ran-GTP is bonded to the importin. Importin cannot bind to another NLS unless ran-GTP is gone. So when it leaves the nucleus and it hydrolyzes the GTP in the cytoplasm. ran-GTP turns into ran-GDP and now importin does not want to hold on anymore.
Exportin - we need a protein to get out of the nucleus. physically impossible for an RNA to have a NES bc it is made up of nucleotides not amino acids. The only way an exportin has affinity for an NES is if it’s partnered with ran-GTP.
Mitochondria revisited:
Mitochondria have their own genomes.
Human mitochondrial genome:
most of the mitochondrial genome has been migrated to the nucleus of the host cell. What do the 16,500 base pairs code for?
16.5 kb of DNA
13 protein-coding genes (all part of Complex I, III, IV and ATP synthase. all have something to do with the ETC or oxidative phosphorylation)
22 tRNA genes (makes those 13 proteins)
2 rRNA genes
ori
what can a mitochondria make protein wise? it makes proteins necessary for the ETC. but can it make them all? These proteins are necessary but NOT sufficient. everything else has to be imported.
Most mitochondrial proteins are not coded for by genes on the mitochondrial genome
Where are they transcribed?
Where are they translated?
How are they transported?
An electrochemical gradient exists in mitochondria:
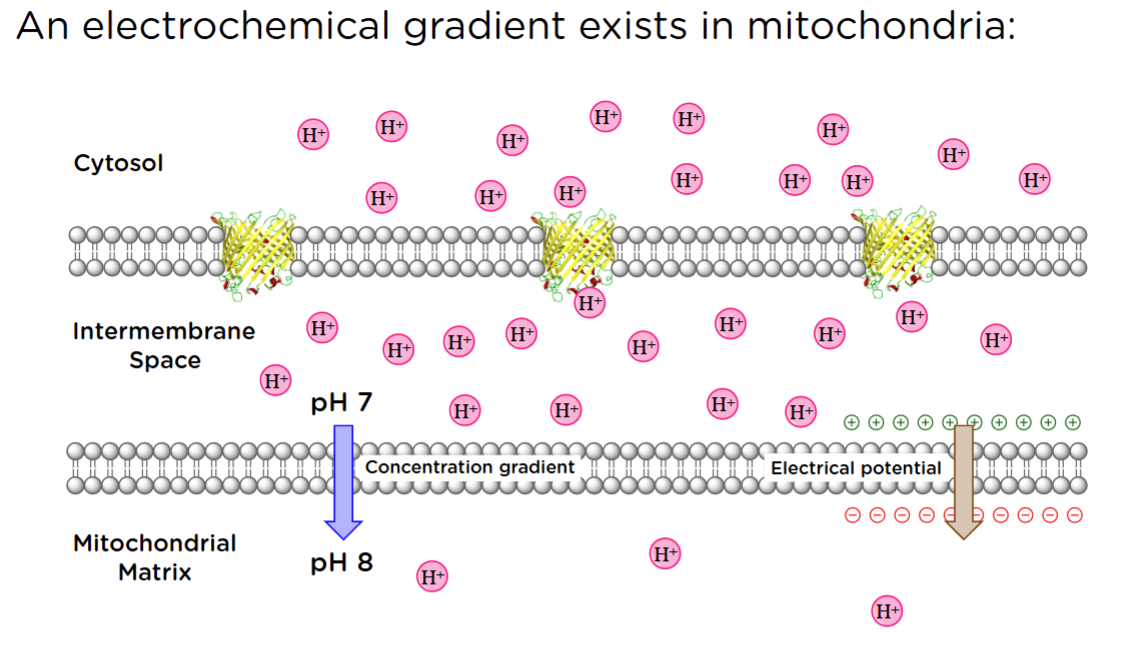
If you were positively charged and you approached the mitochondria, where would you rather be? In the matrix bc there are fewer other positive charges around you.
Mitochondrial protein import:
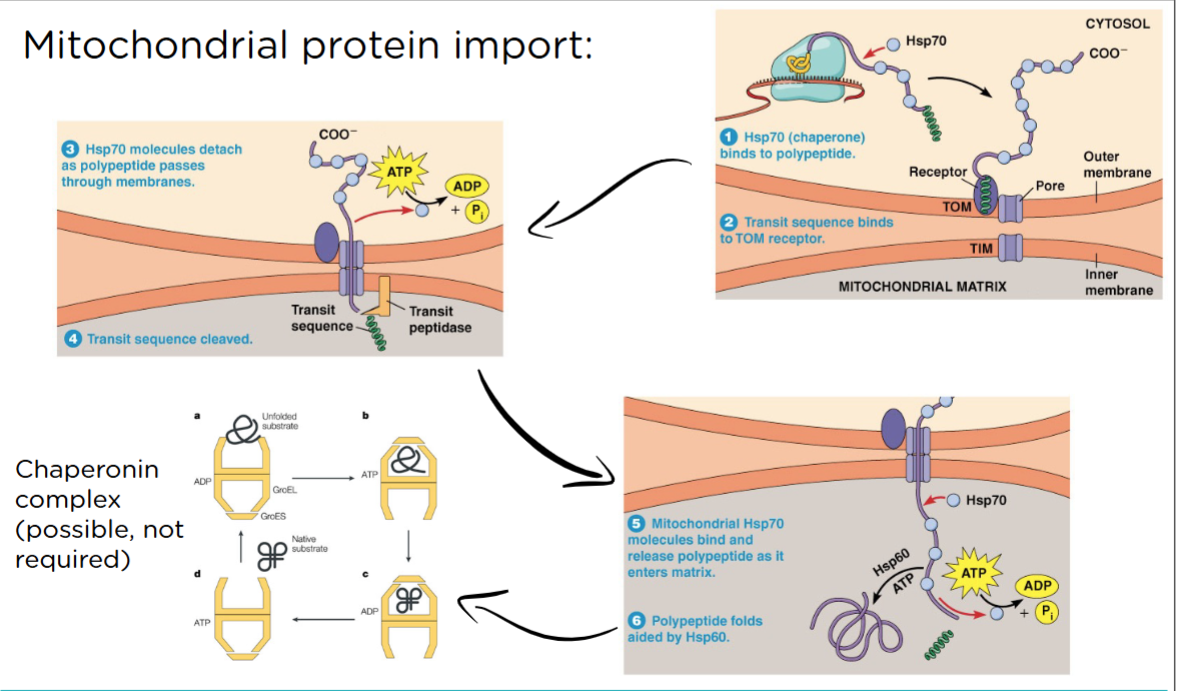
Certain channels can only open wide enough for an unfolded protein to get through. Every other transport channel that is embedded in a membrane and allows for the movement of a protein from one side to another is called a translocon. These can close all the way. they will only open wide enough for an unfolded protein to be roped through. There are chaperones that keep the proteins unfolded in the cytoplasm. The translocon of the outer membrane is called TOM. the translocon of the inner membrane is called TIM. You bind to a tom complex, it recognizes that transit sequence and 2 things start happening. 1 the head moves into the channel that just opened. 2 This opening also causes the nearest tim complex to open. If you just let go nothing would happen but bc the matrix is less positively charged and your head is in a more positively charged area, it gets pushed through. once it gets through 2 things happen. 1 peeling off those chaperones (ATP gets used) 2 don’t need the head anymore and transit sequence gets cut off. Matrix chaperones pull the rest in. They pull you in and help you fold properly. if those chaperones are not enough to get you folded the right way, we might have to put the protein in a protein folding canister, a chaperonin complex. ATP gets spent here. Once you are in the mitochondria you stay in there.
RER protein import:
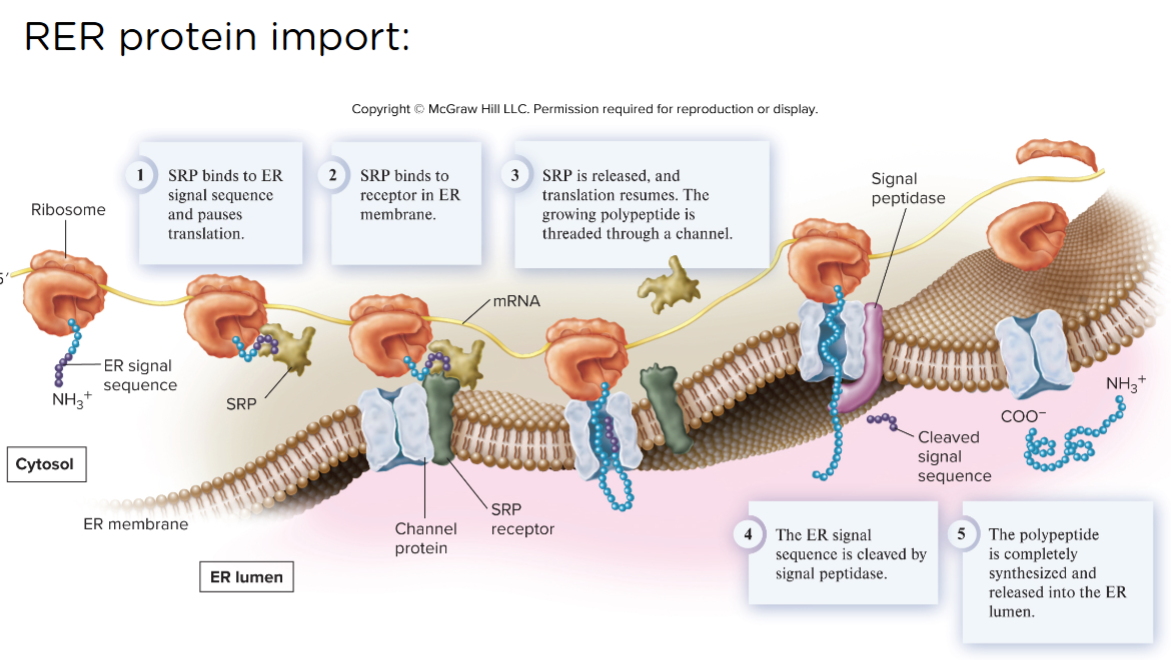
Something is presented that makes translation stop and goes to the ER. the first 25 amino acids that are destined for the co translational sorted side are called an ER signal peptide. It’s a set of amino acids that says don’t leave me where i am right now. This signal gets recognized by a particle by SRP (signal recognition particle, which has an RNA and a protein component). This SRP binds to the signal peptide and does 2 things. 1 pauses translation by blocking the a site 2 causes everybody to make their way to the ER.
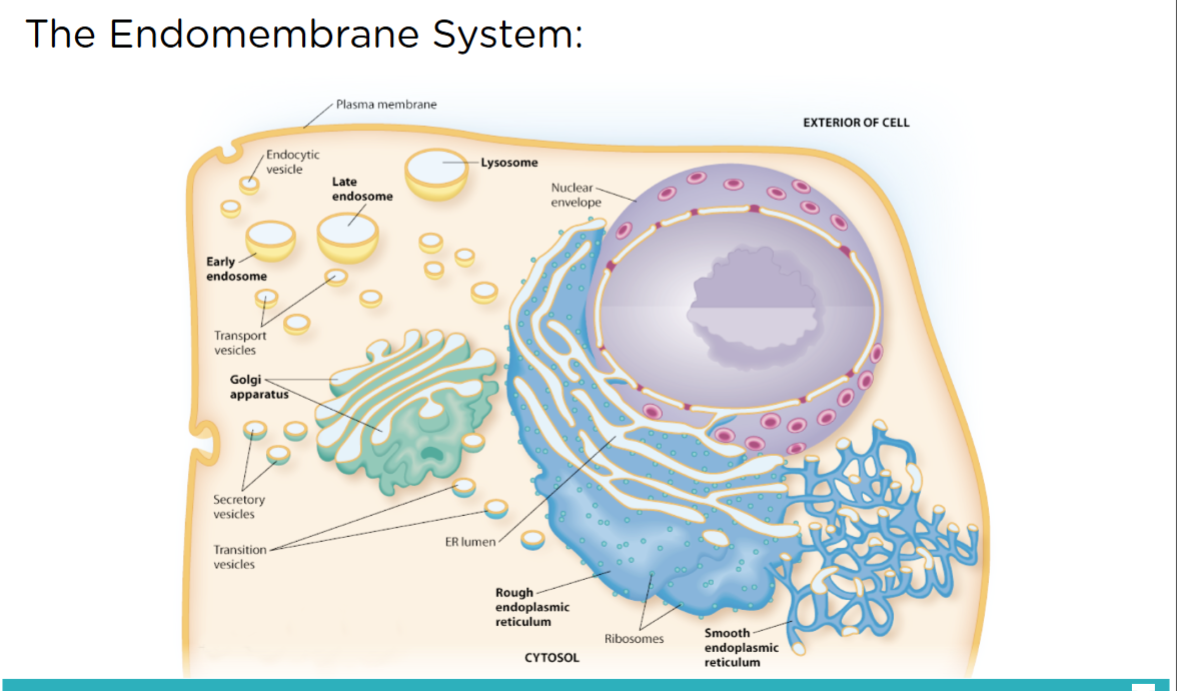
7.5 Analyze the process of transporting molecules into and through the endomembrane system.
Cells that secrete digestive enzymes or other proteins have particularly large amounts of rough ER and Golgi. This correlation led to the idea that these organelles may participate in a “secretory pathway” that starts in the rough ER and ends with products leaving the cell.
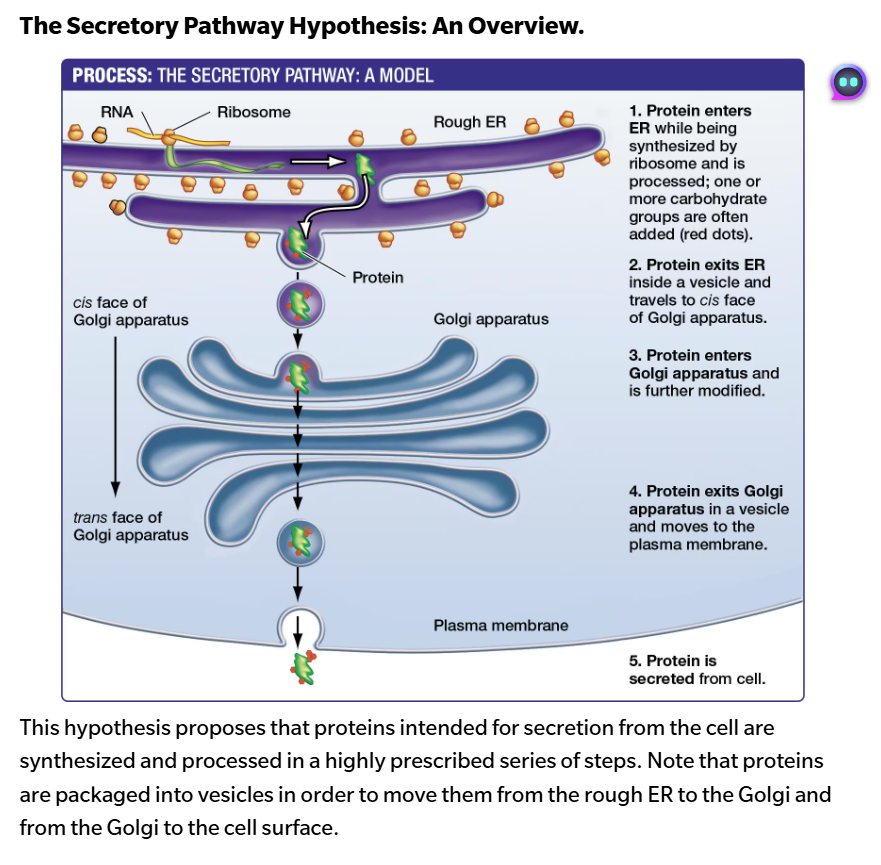
Proteins are trafficked through the secretory pathway in a highly organized and directed manner. A secretory pathway exists and the rough ER and Goldi apparatus function together as parts of an integrated endomembrane system.
The synthesis or proteins destined to be secreted or embedded in membranes begins in ribosomes free in the cytosol. The “signal hypothesis" predicts that proteins bound for the endomembrane system have a molecular code that serves a similar role to the nuclear localization signal in the nucleoplasmin.
There is an ER signal sequence that guides the growing protein and associated ribosome to the rough ER. The ER signal sequence typically is present in the first 20 amino acid residues and is removed when protein synthesis is complete.
Key steps involved for a protein that will eventually be shipped to the inside of an organelle or secreted from the cell:
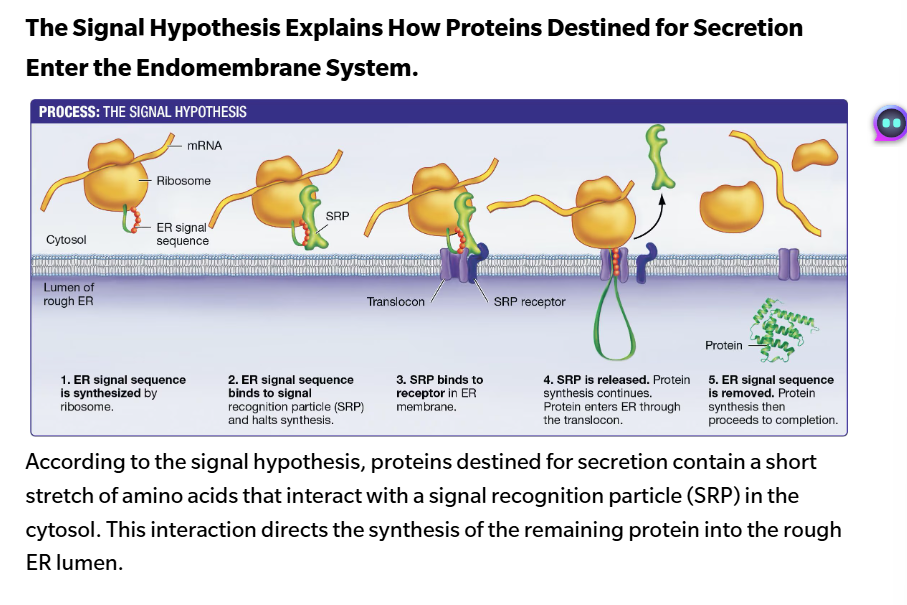
protein synthesis begins on a free ribosome in the cytosol. the ribosome synthesizes the ER signal sequence, using information carried in an mRNA
The signal sequence binds to a signal recognition particle (SRP)—a complex of RNA and protein. the attached SRP causes protein synthesis to stop
the ribosome + signal sequence + SRP complex moves to the rough ER membrane, where it attaches to the SRP receptor. Think of the SRP as a key that is activated by an ER signal sequence. The SRP receptor in the ER membrane is the lock
once the lock (the receptor) and key (the SRP) connect, the SRP is released, and protein synthesis continues through a channel called the translocon.
The growing protein is fed into the ER lumen, and the ER signal sequence is removed
After cleavage of the signal sequence, the protein may be completely released into the ER lumen. Some proteins, however, remain associated with the membrane as integral membrane proteins. How do such proteins get inserted into the ER membrane? Translocon has molecular “gates” that divert stretches of nonpolar amino acids in the growing protein into the phospholipid bilayer. After pushing these regions into the membrane, the rest of the protein continues to the made in the cytosol or the ER lumen.
Once proteins are inside the rough ER or inserted into its membrane, they fold into their 3-dimensional shape with the help of chaperone proteins.
Proteins are transported from the rough endoplasmic reticulum (ER) to the Golgi apparatus via vesicular transport. Vesicles, which are small, membrane-bound structures, bud off from the ER carrying the proteins. These vesicles travel through the cytoplasm and dock at the cis face of the Golgi apparatus, where they fuse and release their cargo for further processing. This process involves precise molecular mechanisms to ensure proper cargo selection, transport, and delivery.
How do proteins travel from the ER to the Golgi apparatus? Proteins are transported in vesicles that bud off from the ER and move to the cis face of the Golgi apparatus.
The composition of the Golgi apparatus is dynamic. New cisternae constantly form at the cis face of the Golgi apparatus, while old cisternae break apart at the trans face, to be replaced by the cisternae behind it.