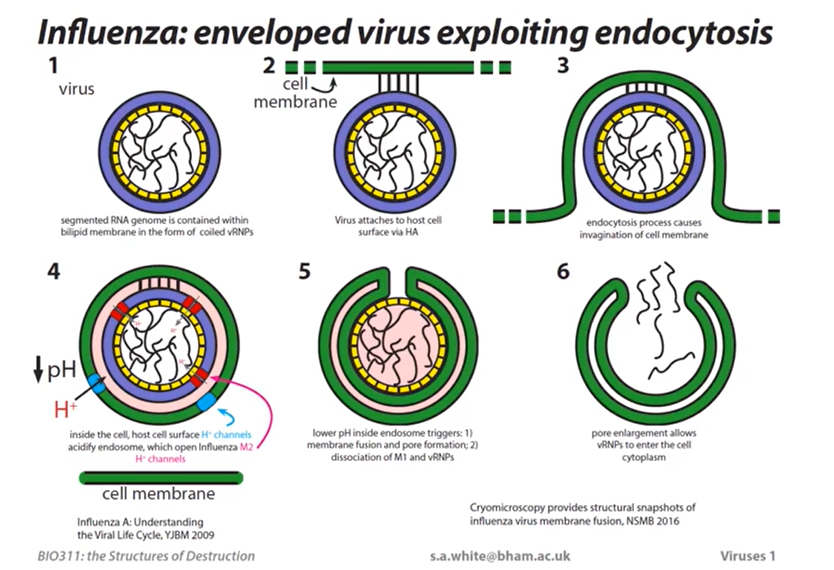
Influenza Virus
Viruses and Viral Proteins
Influenza Virus Structure and Entry
Influenza virus is an enveloped virus with a segmented RNA genome. This means that there is a bilipid membrane that surrounds the nucleic acid inside.
Influenza is a segmented RNA virus.
There are 8 RNA segments, each bound to multiple nucleoproteins (NPs) and a polymerase (P).
RNA + NPs + P = vRNP (Viral Ribonucleoprotein) complexes that are essential for the transcription and replication of the viral genome.

The segmented genome is contained within a bilipid membrane.
The membrane features 3 trans-membrane proteins: Haemagglutinin (HA), Neuraminidase (NA), and M2 protein.
M1 matrix protein (in yellow on image) (underneath the bilipid membrane) binds the ribonucleoprotein segments.
Influenza virus particles can be:
Spherical: spread through aerosol between hosts.
Filamentous: more commonly found: important in infecting neighbouring cells.
Side note: not to be confused with Haemophilus influenza which is caused by a bacteria and gives similar symptoms.
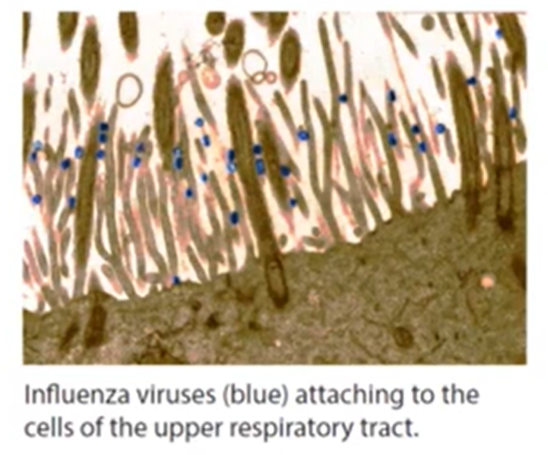
Viral Entry Mechanism
Attachment: Virus attaches to the host cell surface via Hemagglutinin (HA).
Endocytosis: The binding of the virus causes invagination (bending) of the cell membrane. The virus is brought into the cell via endocytosis and is inside an endosome which was once part of the cell membrane.
Acidification:
Host cell (proton) H+ channels pumps protons across into the endosome to acidify the endosome as an immune response. This lowers the pH inside.
Influenza M2 H+ channels open in response to the lower pH on the outside of the endosome and subsequently the pH has now lowered in the inside of the virus envelope.
This triggers a number of events
Triggered Events Due to Lowered pH:
Membrane fusion and pore formation. This means a hole has been made that allows the viral RNA to be released into the host cell cytoplasm, facilitating viral replication and infection.
Dissociation of M1 and vRNPs, which leads to the release of viral ribonucleoproteins into the cytoplasm, thus promoting the transcription and replication processes essential for viral propagation.
Entry into Cytoplasm: Pore enlargement allows vRNPs to enter the cell cytoplasm.
Key Viral Structural Proteins
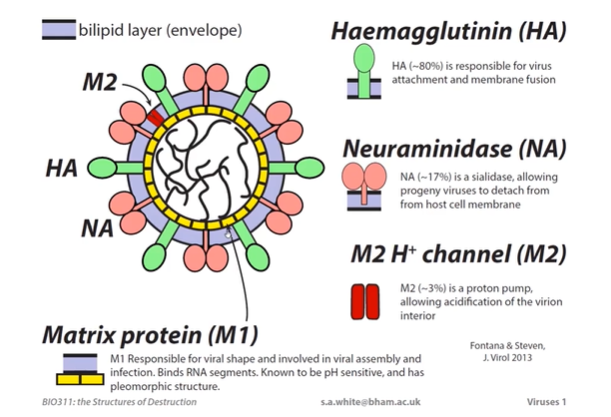
Haemagglutinin (HA)
HA makes up ~80% of the viral proteins.
Transmembrane protein and sits thru the bilipid layer and extents into the viral exterior.
Responsible for virus attachment to the host cell and membrane fusion.
Essential for viral entry and pore formation.
Neuraminidase (NA)
NA makes up ~17% of the viral proteins.
It is an enzyme. Has a catalytic function to breaking bonds in sialic acid.
Sialic acid is a sugar chain that is attached to glycoproteins in the host cell surface. The influenza virus attaches by recognising SA, this is done by HA binding to SA.
It is a sialidase, allowing developing endosomes to detach from the host cell membrane and infect neighbouring cells.
Key role in viral egress (exit strategy) and a target for antivirals.
The sialidase head group catalyses the cleavage of sialic acid residues from carbohydrate sidechains of both HA and glycosylated cellular membrane proteins.
NA is a homotetramer attached to a membrane an chor.
M2 H+ Protein channel
(Influenza) M2 makes up ~3% of the viral proteins.
It is a proton pump, allowing acidification of the virion interior.
Matrix Protein (M1)
M1 is responsible for the viral shape and is involved in viral assembly and infection.
It binds RNA segments.
Known to be pH sensitive and has a pleomorphic structure.
Assembles into a metastable matrix layer attached to the viral envelope during virion formation.
Within the endosome, a trigger converts the matrix layer into a multi-layered matrix.
Threat of Pandemic Flu
A new influenza virus will appear at some time in the future through natural processes or bioterrorism.
The world’s population will likely have no immunity to the new virus, causing rapid spread, misery, economic disruption, and deaths.
Vaccines will take time to develop; antiviral drugs, like neuraminidase inhibitors, are the most effective means of control in the early stages of an epidemic.
Tamiflu (oseltamivir) is an example of a neuraminidase inhibitor.
Neuraminidase Structure and Inhibitors
Neuraminidase is a 6-bladed -propeller.
Tamiflu binds to neuraminidase.
Key residues in the active site include:
R118, R292, R371, E119, D151, A246, I222
Design of antivirals targeting influenza NA:
Initial attempts (1970s-80s) to identify inhibitors by high-throughput screening failed.
Crystallographic studies of neuraminidase crystals soaked in sialic acid led to initial inhibitors like DANA and FANA.
Tamiflu, Relenza, and related drugs were developed with concerted effort (1990s - 2000s) and are now in clinical use.
Antiviral design focuses on:
arginine pocket
conserved acid pocket
neutral pocket
Haemagglutinin (HA) Structure and Function
HA0 is a folded trimeric glycoprotein with a cleavable loop near the viral membrane.
Disulphide bridges link the two halves of the folded protein.
The loop is cleaved extracellularly by trypsin-like protease.
Mature HA (infectious form) consists of two polypeptides (HA1 & HA2) but remains folded in its trimeric quaternary structure.
HA requires cleavage for infectivity.
HA folds as a trimer to form a triple-coiled coil about Helix C.
Significant rearrangement of secondary structure, triggered by low pH, moves the fusion peptide from the viral membrane into the host-cell membrane.
Conformational change brings fusion peptide and transmembrane domains closer, leading to membrane fusion & pore formation.
Mutations in the sialic acid receptor allow binding of both -2,3 (avian) and -2,6 (human) receptors.
Pathogenicity and HA Acid Stability
Pathogenicity is regulated by HA acid stability.
Strains with lower pathogenicity had the same rate of HA cleavage as high pathogenicity H5N1 variants.
The level of pathogenicity correlates with the pH at which conformational change is triggered.
Acid stability is largely mapped to amino acid differences at positions 104 and 115, either end of a helix in an HA1 subdomain, where highly conserved interactions are disrupted.
HAs from highly pathogenic H5N1 strains undergo conformational change at pH 5.8 - 6.0.
Optimal pH stability of HA:
Too high and HA triggers too soon within the endosome.
Too low and influenza has been transferred to the lysosome.
Influenza: Current and Future Challenges
Universal antibody:
Screening of plasma cells identified antibodies that bind to both Group 1 and Group 2 HAs.
Analysis of successful antibodies showed common elements and enabled protein engineering to produce FI6-v3, binding and neutralising many strains from both groups.
Neutralisation mechanism appears to be a combination of cleavage inhibition and conformational change inhibition.
HA2-based synthetic peptide vaccine providing protection in mice against influenza viruses of structurally divergent subtypes has been developed.
A structural explanation for the low effectiveness of the seasonal influenza H3N2 vaccine exists.
L194P is an egg-adaptive substitution in human H3N2 HA.
Egg-passaged human H3N2 isolates are categorized based on the number of passages.
M1 Protein Structure and Function
During virion formation, M1 assembles into a metastable matrix layer that is attached to the viral envelope.
Within the M1-V97K oligomer, matrix proteins interact via the stacked dimer interface while the CTDs and NTDs are in close contact