sample questions
1. A baby presents at a clinic with a severe neurodevelopmental disorder that you suspect may
be caused by a de novo (sporadic) genetic mutation. Describe how you could identify
candidate Single Nucleotide Polymorphisms (SNP) that may cause the disease, without
having access to samples from the child’s parents or from any other family member. Briefly
describe a different approach that you could use to identify Copy Number Variants (CNV)
Lecture 5: Complex Traits
– Includes methods for identifying SNPs using GWAS, and understanding the use of SNP arrays.
2. Define linkage disequilibrium, haplotype, and Tag-SNP.
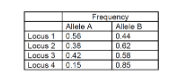
The table below shows the frequency of alleles (called A and B) at four different loci
numbered (1-4) measured in a human population. The frequency of the haplotype ABBA was
observed in 3% of the population (where the alleles at each locus are shown in order,
starting with Locus 1). Do you think these alleles show evidence of LD, and why
Lecture 5: Complex Traits
– Explains Tag-SNPs and includes examples of LD blocks and haplotypes in GWAS.
3. For Genome Wide Association Studies (GWAS), variants present in the tested groups
(patients and controls) can be identified using SNP arrays (often called bead arrays, high
density arrays, microarrays or similar). Describe how these arrays can be used to
characterise up to 1 million SNPs (Single Nucleotide Polymorphisms) in a single individual in
a single experiment. Describe how the method works – that is, how the base present at each
potential polymorphic site can be identified.
Lecture 5 and 6:
– Cover Illumina Bead Arrays, SNP identification using high-density arrays, and how fluorescent signals identify base changes.