D1.1 DNA replication Notes
D1.1.1 dna replication
DNA Replication: Producing Exact Copies of DNA
Imagine you're building a replica of a complex structure, like a Lego model.
Each piece must fit perfectly to create an exact copy.
DNA replication works similarly, ensuring that every cell in your body has an identical set of genetic instructions.
Why DNA Replication Matters
DNA replication is essential for:
Reproduction: Offspring inherit DNA from their parents.
Growth and Tissue Replacement: Multicellular organisms rely on cell division to grow and repair tissues.
Tip
Before a cell divides, it must replicate its DNA so that each daughter cell receives a complete set of genetic instructions.
The Mechanism of DNA Replication
Definition
DNA ReplicationDNA replication is the biological process by which a cell copies its DNA to ensure that each daughter cell receives an identical set of genetic instructions during cell division. It is essential for growth, repair, and reproduction in living organisms.
It is a highly coordinated process involving several key steps and enzymes.
">Process of DNA Replication
1. Unwinding the Double Helix
The first step is to unwind the DNA double helix, which is tightly coiled like a spring.
This task is performed by an enzyme called helicase.
Helicase breaks the hydrogen bonds between complementary base pairs (A-T and C-G), unzipping the two strands of DNA.
Analogy
Think of helicase as a zipper puller, separating the two sides of a zipper to open it up.
2. Complementary Base Pairing
Once the strands are separated, each serves as a template for building a new complementary strand.
Free nucleotides in the cell align with their complementary bases on the template strand:
Adenine (A) pairs with thymine (T).
Cytosine (C) pairs with guanine (G).
Example
If the template strand has the sequence A-T-C-G, the new strand will be T-A-G-C.
3. Synthesizing the New Strand
The enzyme DNA polymerase plays a critical role in assembling the new DNA strand.
It adds nucleotides one by one to the growing strand.
This ensures that each base is correctly paired with its complement on the templatestrand.
DNA polymerase also forms covalent bonds between the sugar and phosphate groups of adjacent nucleotides, creating a continuous sugar-phosphate backbone.
Warning
A common mistake is thinking that DNA polymerase can start a new strand from scratch. It cannot, it requires a short RNA primer to begin synthesis.
4. Proofreading and Error Correction
DNA replication is remarkably accurate, with an error rate of about 1 in 10 billion bases.
DNA polymerase has a proofreading function that detects and corrects mismatched bases during replication.
This ensures that the new DNA strand is nearly identical to the original.
Note
In a human cell, which has about 6 billion base pairs, this level of accuracy means there are, on average, only 0.6 errors per replication cycle.
Semi-Conservative Replication
DNA replication is described as semi-conservative because each new DNA molecule consists of:
One original strand (from the parent molecule).
One newly synthesized strand.
This method ensures that the genetic information is preserved across generations.
Example
The Meselson-Stahl experimentin the 1950s provided evidence for semi-conservative replication. By using isotopes of nitrogen to label DNA, they demonstrated that each daughter DNA molecule contains one strand from the parent molecule and one newly synthesized strand.
The Role of Complementary Base Pairing
Complementary base pairing is the foundation of DNA replication.
It ensures that the new DNA strands are exact copies of the original.
If a nucleotide with the wrong base tries to pair, it is rejected because the hydrogen bonds cannot form.
This specificity guarantees that the base sequence of the new strand matches the template strand.
Analogy
Complementary base pairing is like a lock-and-key mechanism.
Only the correct key (base) can fit into the lock (its complementary base).
Biological Significance of DNA Replication
DNA replication is vital for:
Reproduction: Ensures offspring inherit genetic information.
Growth: Supports cell division for growth in multicellular organisms.
Tissue Replacement: Enables the replacement of worn-out or damaged cells.
Example
When you cut your skin, DNA replication allows new cells to form and repair the wound.
Reflection and Broader Connections
DNA replication is a cornerstone of life, ensuring genetic continuity and stability.
Tok
How does the precision of DNA replication influence our understanding of genetic inheritance and evolution? Could errors in replication ever be beneficial?
D1.1.2 semi-conservative nature of dna replication
DNA Replication is a Semi-Conservative Process
Definition
Semi-conservative replicationWhen each new DNA molecule consists of one original (parental) strand and one newly synthesized strand.
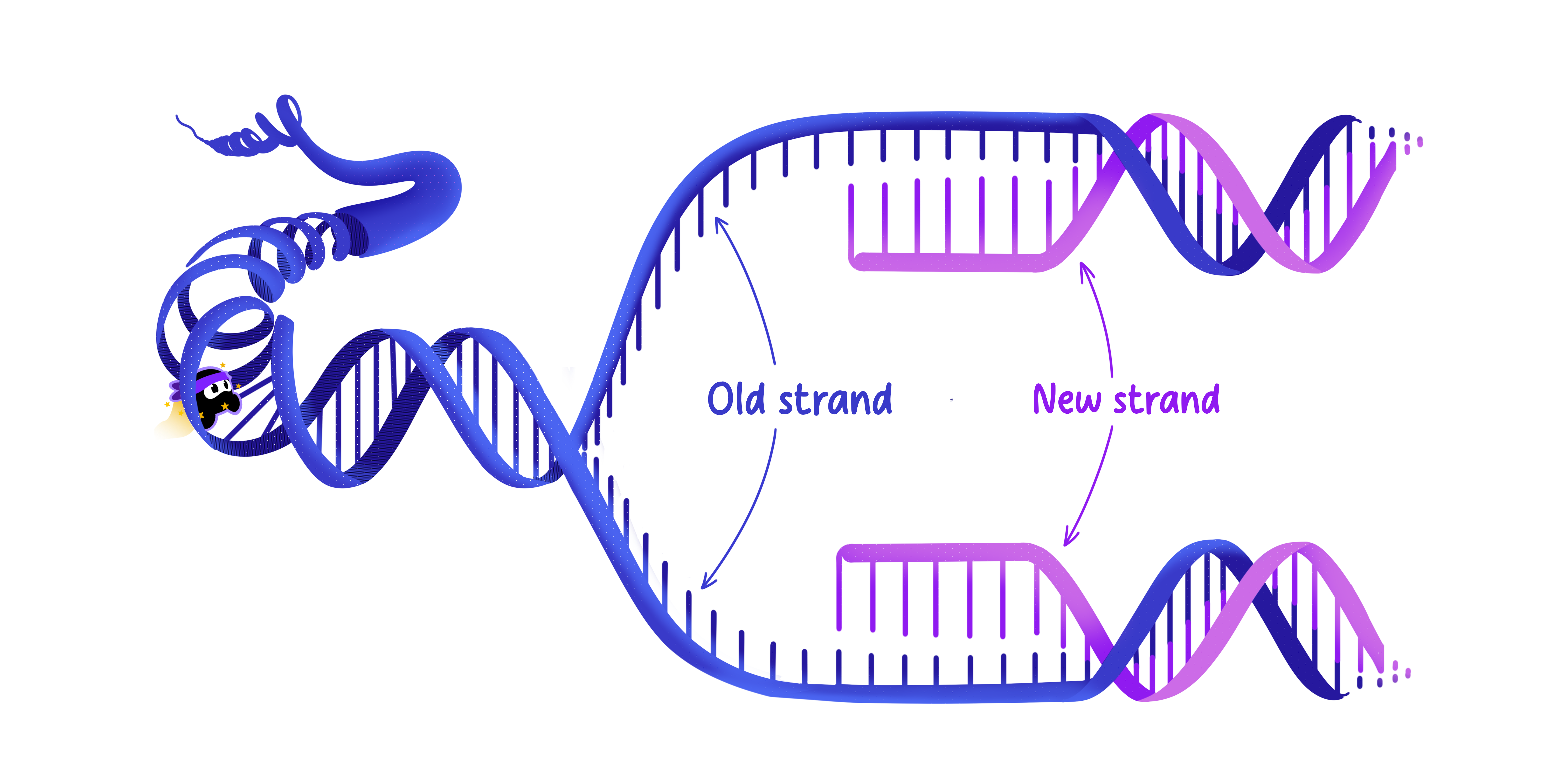
When DNA replicates, the two strands of the double helix separate.
Each original strand serves as a template for a new complementary strand, which results in two identical DNA molecules, each containing one old strand and one new strand.
This method ensures that genetic information is preserved with high fidelity during cell division.
Tip
"Parents share, kids pair."
Each parent strand is shared with a new complementary strand in the daughter molecules.
How Complementary Base Pairing Ensures High Accuracy
Template-Driven Synthesis
DNA polymerase reads the template strand.
For each base on the template, DNA polymerase adds the complementary nucleotide to the new strand.
Hydrogen Bonding as a Quality Check
Correct base pairs form stable hydrogen bonds (A-T or C-G).
Incorrect base pairs (e.g., A-C) do not form proper hydrogen bonds.
If hydrogen bonding is weak or absent, the incorrect nucleotide is rejected.
Proofreading by DNA Polymerase
DNA polymerase has a proofreading function.
If an incorrect nucleotide is added, DNA polymerase detects the mismatch.
The enzyme removes the incorrect nucleotide and replaces it with the correct one.
This dramatically reduces replication errors.
Example
If the template has adenine (A), DNA polymerase adds thymine (T) to the new strand.
Note
This results in a very high fidelity process where the combination of complementary base pairing and proofreading ensures DNA replication is extremely accurate.
In fact, the error rate is approximately 1 mistake per 1 billion nucleotides after proofreading.
Self Review
What does "semi-conservative" mean in the context of DNA replication?
Describe the structure of DNA after one round of semi-conservative replication.
What are the complementary base pairing rules?
How does complementary base pairing ensure accurate DNA replication?
What is the role of DNA polymerase's proofreading function?
What would happen if complementary base pairing did not occur accurately?
D1.1.3 role of helicase and dna polymerase in dna replication
DNA Replication Requires Unwinding and Synthesis
DNA replication copies the entire genome so each daughter cell receives identical genetic information.
Two key enzymes drive this process:
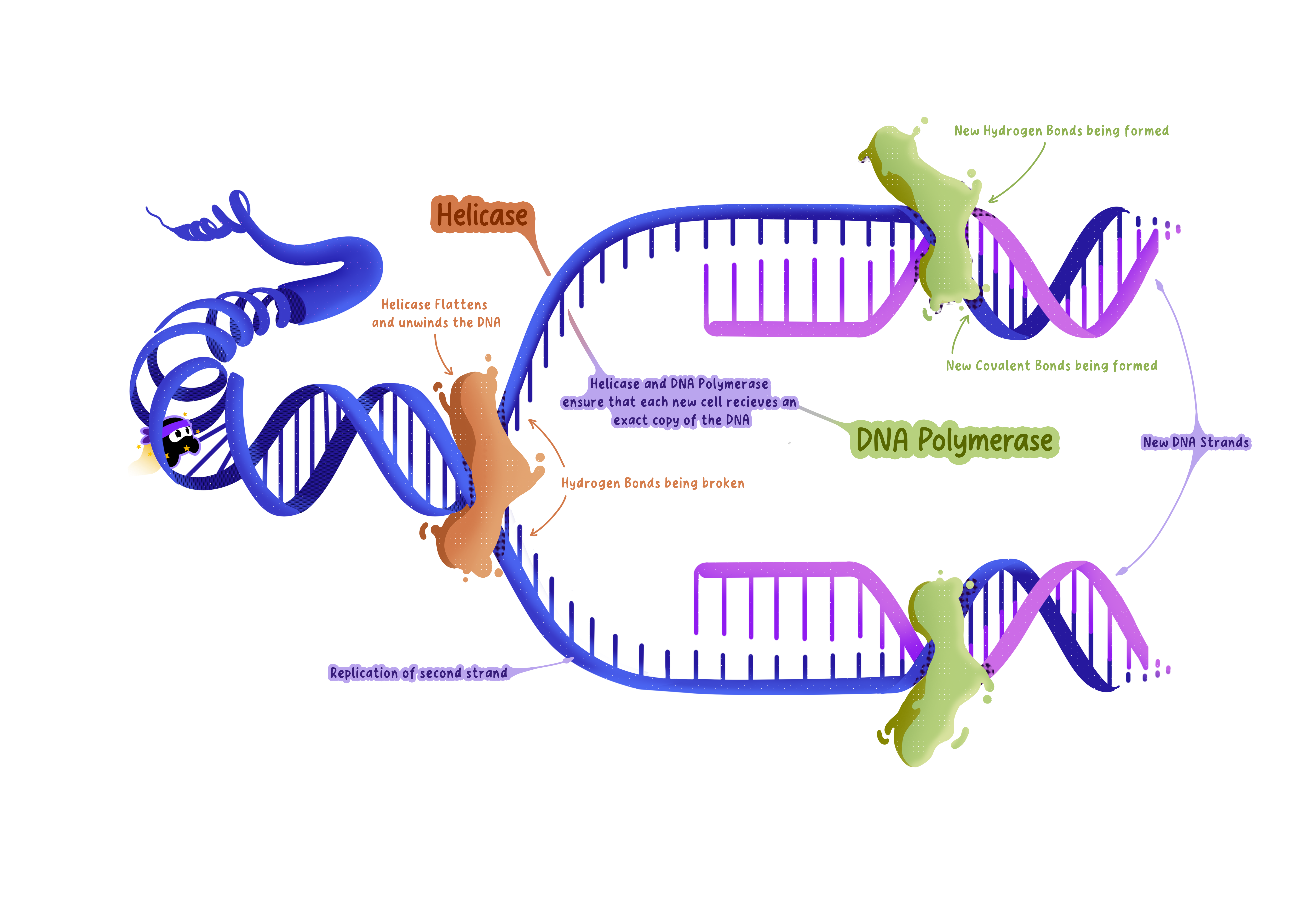
Helicase: Unwinds the DNA double helix.
DNA polymerase: Synthesizes new DNA strands.
Tip
Helicase doesn't break the covalent bonds in the DNA backbone, it only disrupts the weaker hydrogen bonds between the bases.
The Role of Helicase: Unwinding the Double Helix
Definition
Helicase An enzyme that separates the two strands of the DNA double helix.
How Helicase Works
Step 1: Breaking Hydrogen Bonds
Helicase moves along the DNA molecule.
It breaks the hydrogen bonds between complementary base pairs (A-T and C-G).
This separates the two strands.
Step 2: Forming the Replication Fork
As helicase unwinds the DNA, it creates a Y-shaped structure called the replication fork.
The two separated strands serve as templates for new DNA synthesis.
Key Features
Ring-shaped enzyme: Helicase encircles one strand of DNA and moves along it.
Only breaks hydrogen bonds: It does NOT break the covalent bonds in the sugar-phosphate backbone, only the weaker hydrogen bonds between bases.
Directional movement: Helicase moves in one direction (typically 3' to 5' on the template strand), continuously unwinding DNA ahead of the replication machinery.
Analogy
Helicase is like unzipping a jacket.
The zipper (helicase) separates the two sides (DNA strands) by breaking the connections (hydrogen bonds) between the teeth (base pairs).
The Role of DNA Polymerase: Synthesizing New DNA Strands
Definition
DNA PolymeraseAn enzyme that synthesizes new DNA strands by adding nucleotides to a growing chain, using the original strand as a template.
How DNA Polymerase Works
Step 1: Reading the Template
DNA polymerase reads the template strand (one of the separated strands).
It determines which nucleotide to add next based on complementary base pairing:
A pairs with T
C pairs with G
Step 2: Adding Nucleotides
DNA polymerase adds free nucleotides to the growing strand, one at a time.
Each nucleotide is added to the 3' end of the growing strand.
Step 3: Forming Covalent Bonds
DNA polymerase catalyzes the formation of covalent bonds between the sugar of one nucleotide and the phosphate of the next.
This creates the sugar-phosphate backbone of the new DNA strand.
Directionality: 5' to 3' Synthesis
DNA polymerase can only add nucleotides in the 5' to 3' direction.
This has important consequences:
Leading strand: Synthesized continuously in the 5' to 3' direction as helicase unwinds the DNA.
Lagging strand: Synthesized discontinuously in short segments called Okazaki fragments, which are later joined by the enzyme DNA ligase.
Proofreading Function
DNA polymerase has a proofreading ability.
It can detect when an incorrect nucleotide has been added.
It removes the incorrect nucleotide and replaces it with the correct one.
This dramatically reduces replication errors.
Example
In humans, DNA polymerase adds approximately 50 nucleotides per second and has an error rate of less than 1 mistake per billion nucleotides after proofreading.
Comparing Helicase and DNA Polymerase
Enzyme | Function | Bonds Involved |
|---|---|---|
Helicase | Unwinds the DNA double helix | Breaks hydrogen bonds between base pairs |
DNA Polymerase | Synthesizes new DNA strands | Forms covalent bonds in the sugar-phosphate backbone |
Hint
Helicase separates strands; DNA polymerase builds new strands.
Self Review
What is the role of helicase in DNA replication?
What type of bonds does helicase break?
What is the replication fork?
What is the role of DNA polymerase in DNA replication?
In which direction does DNA polymerase synthesize DNA?
D1.1.4 polymerase chain reaction and gel electrophoresis
Polymerase Chain Reaction (PCR): Amplifying DNA
Definition
PCR (Polymerase Chain Reaction)A technique that amplifies DNA, creating millions of copies of a specific DNA sequence in just a few hours.
This allows scientists to study tiny DNA samples from crime scenes, fossils, or other sources.
Key Components of PCR
Template DNA: The DNA you want to copy.
Primers: Short DNA sequences that bind to the template, marking the starting point for replication.
Taq Polymerase: A heat-resistant enzyme that synthesizes new DNA strands.
Nucleotides (dNTPs): The building blocks of DNA.
Thermal Cycler: A machine that precisely controls temperature changes.
Tip
Primers are essential because Taq polymerase can't start building DNA without them.
They are designed to match the beginning and end of the target sequence.
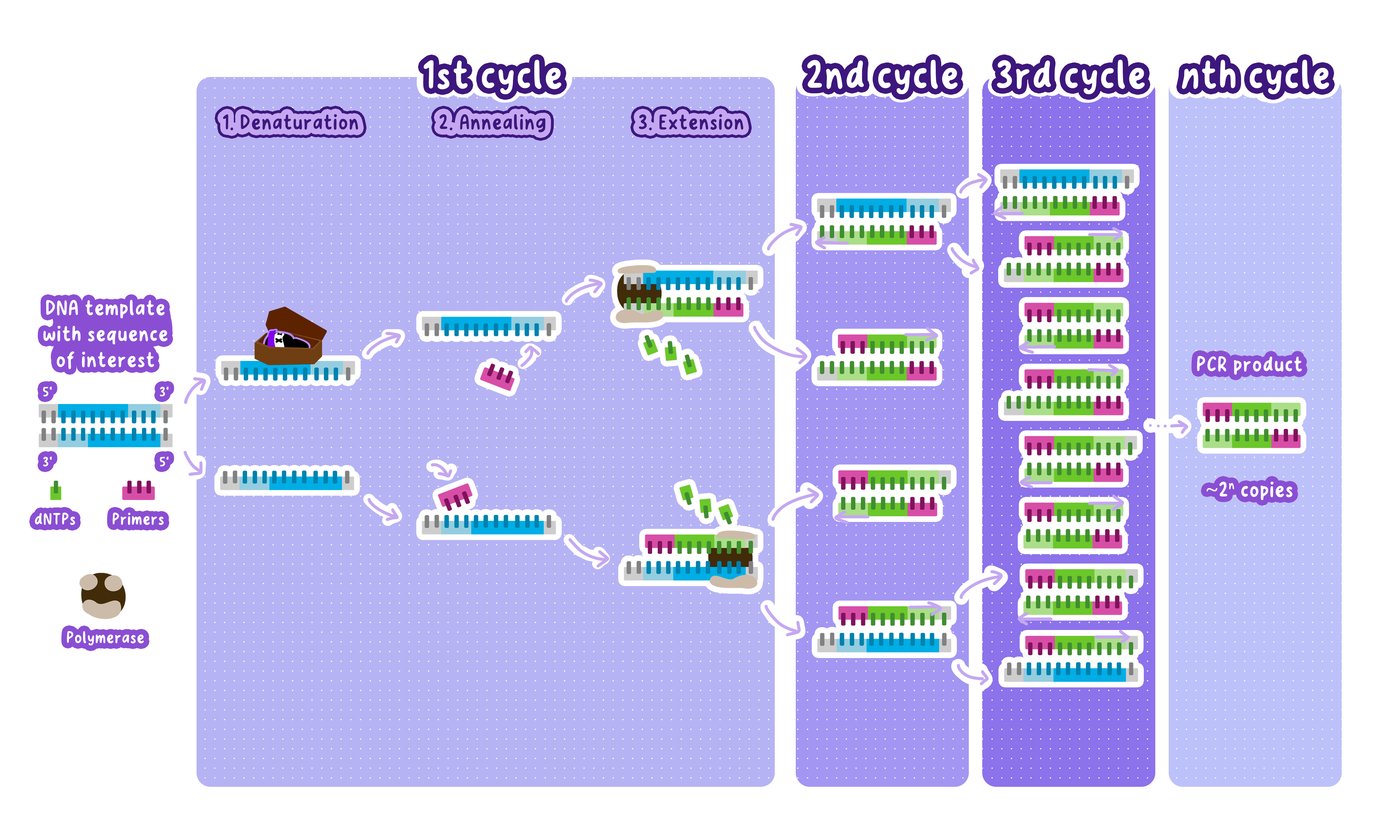
Steps of PCR (DNA Amplification)
PCR consists of a cycle of three main steps, repeated 20-40 times to exponentially amplify the target DNA sequence.
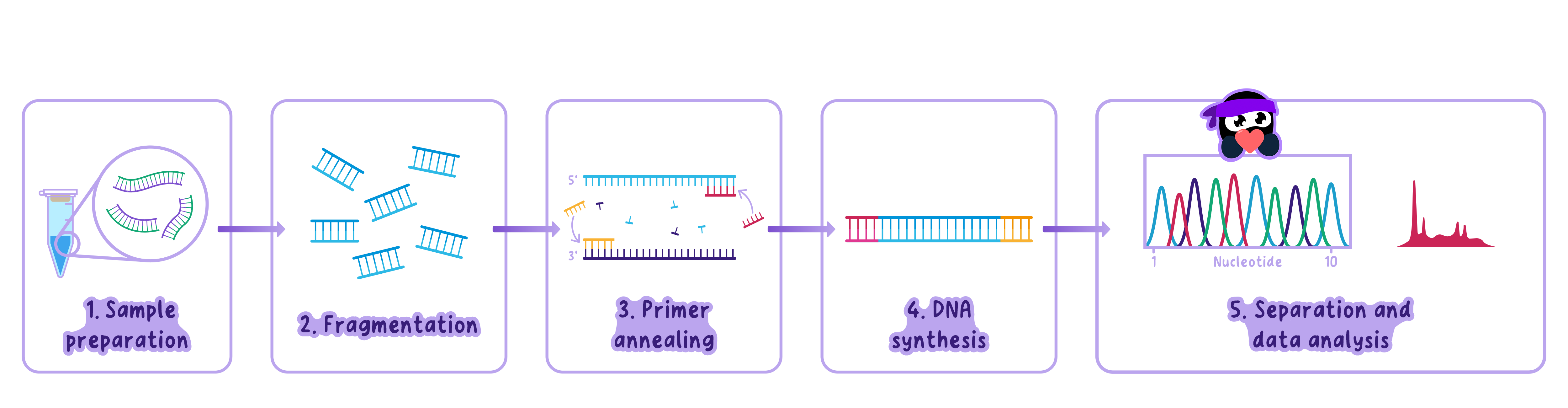
Sample Preparation
The DNA sample is extracted and purified.
Template DNA (the DNA to be copied) is prepared along with other PCR components.
Denaturation (~94-96°C) -> Fragmentation
The DNA is heated to ~95°C.
This breaks the hydrogen bonds between complementary base pairs, separating the double-stranded DNA into two single strands.
These single strands serve as templates for new DNA synthesis.
Annealing (~50-65°C) -> Primer Annealing
The temperature is lowered to 50-65°C.
Short DNA sequences called primers bind (anneal) to complementary sequences on each single-stranded DNA template.
Primers mark the starting point for DNA synthesis and are essential because Taq polymerase cannot start building DNA without them.
Extension (~72°C) -> DNA Synthesis
The temperature is raised to 72°C, the optimal temperature for Taq polymerase.
Taq polymerase is a heat-resistant enzyme that adds nucleotides (dNTPs) to the primers, synthesizing new DNA strands in the 5' to 3' direction.
After one cycle, the amount of DNA doubles.
Separation and Data Analysis -> Gel Electrophoresis
Once DNA is amplified by PCR, gel electrophoresis is used to separate and analyzethe DNA fragments.
Example
After 30 cycles, one DNA molecule becomes over 1 billion copies.
Why Taq Polymerase?
Taq polymerase remains stable at high temperatures (~95°C), unlike most enzymes that would denature.
This allows it to survive the repeated heating cycles in PCR.
Gel Electrophoresis: Separating DNA Fragments by Size
Definition
Gel ElectrophoresisGel electrophoresis is a laboratory technique used to separate and analyze molecules such as DNA, RNA, or proteins based on their size, charge, or shape.
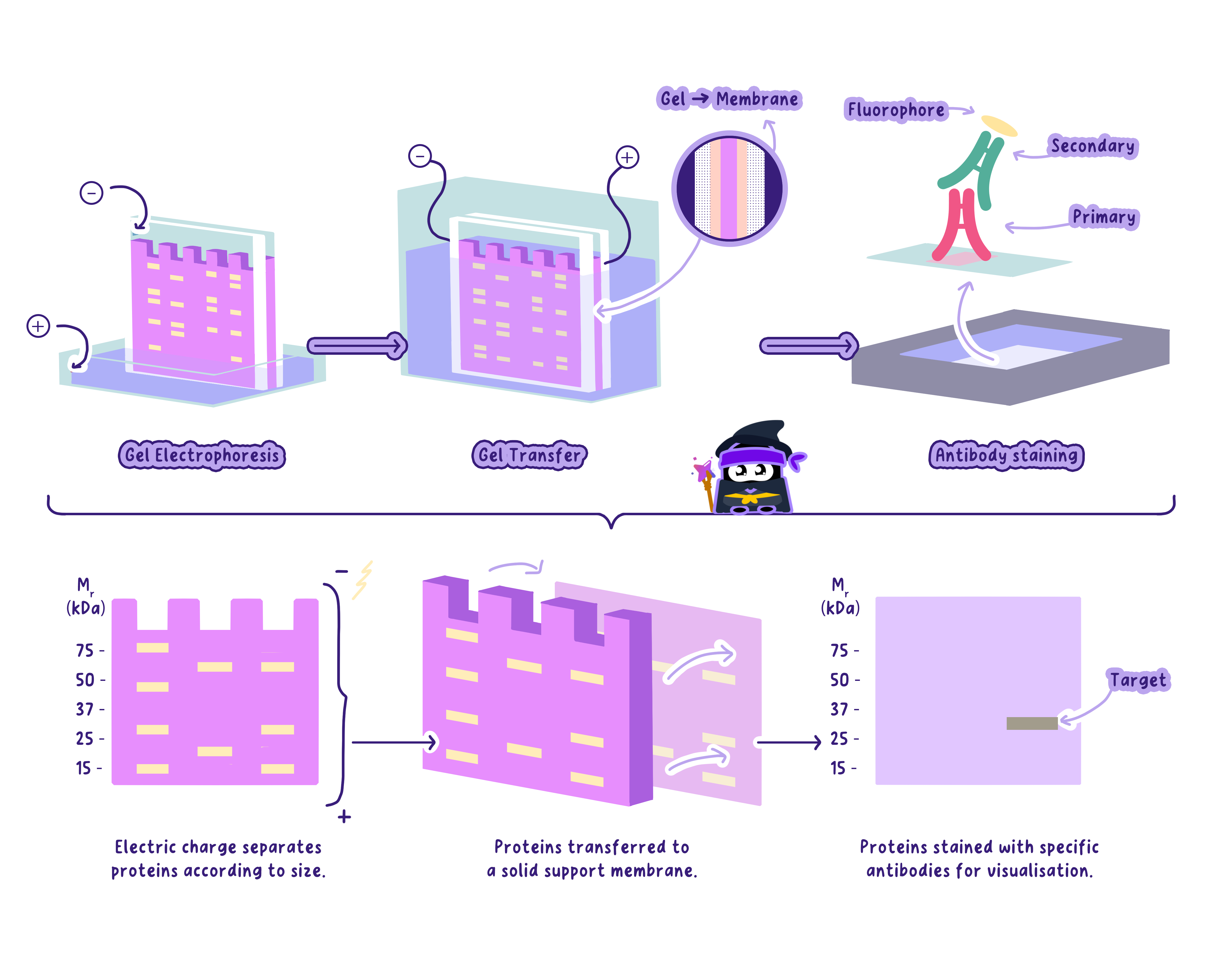
How Gel Electrophoresis Works
Preparation of the gel: A gel (usually agarose) is poured into a mold with wells at one end.
Loading the DNA: DNA samples are mixed with a dye and loaded into the wells.
Applying an electric field: The gel is placed in a buffer solution, and an electric current is applied. • The negative electrode is near the wells. • The positive electrode is at the opposite end.
Migration of DNA: DNA is negatively charged (due to its phosphate backbone), so it moves toward the positive electrode. • Smaller fragments travel faster and fartherthrough the gel. • Larger fragments move slower.
Visualization: The gel is stained with a dye (e.g., ethidium bromide) that binds to DNA, making fragments visible under UV light.
Why DNA Separates By Size
The gel acts like a sieve, slowing down larger fragments more than smaller ones.
This allows DNA fragments to be separated and analyzed based on their size.
Analogy
Imagine a race where smaller runners (DNA fragments) move faster through a crowded field (the gel) than larger ones.
Self Review
Why is Taq polymerase used?
How does gel electrophoresis link to PCR?
D1.1.5 applications of polymerase chain reaction and gel electrophoresis
DNA Profiling Reaveals Unique Patterns
Definition
DNA profilingDNA profiling is a technique used to identify individuals based on unique patterns in their DNA.
DNA profiling is a cornerstone of forensic science, used to match DNA from crime scenes to suspects.
Even a tiny sample, like a drop of blood or a strand of hair, can be amplified and analyzed to identify a suspect or exonerate the innocent.
This is done by relying on regions called short tandem repeats (STRs) short sequences of DNA that repeat multiple times.
The number of repeats varies greatly between individuals, making STRs ideal for distinguishing one person from another.
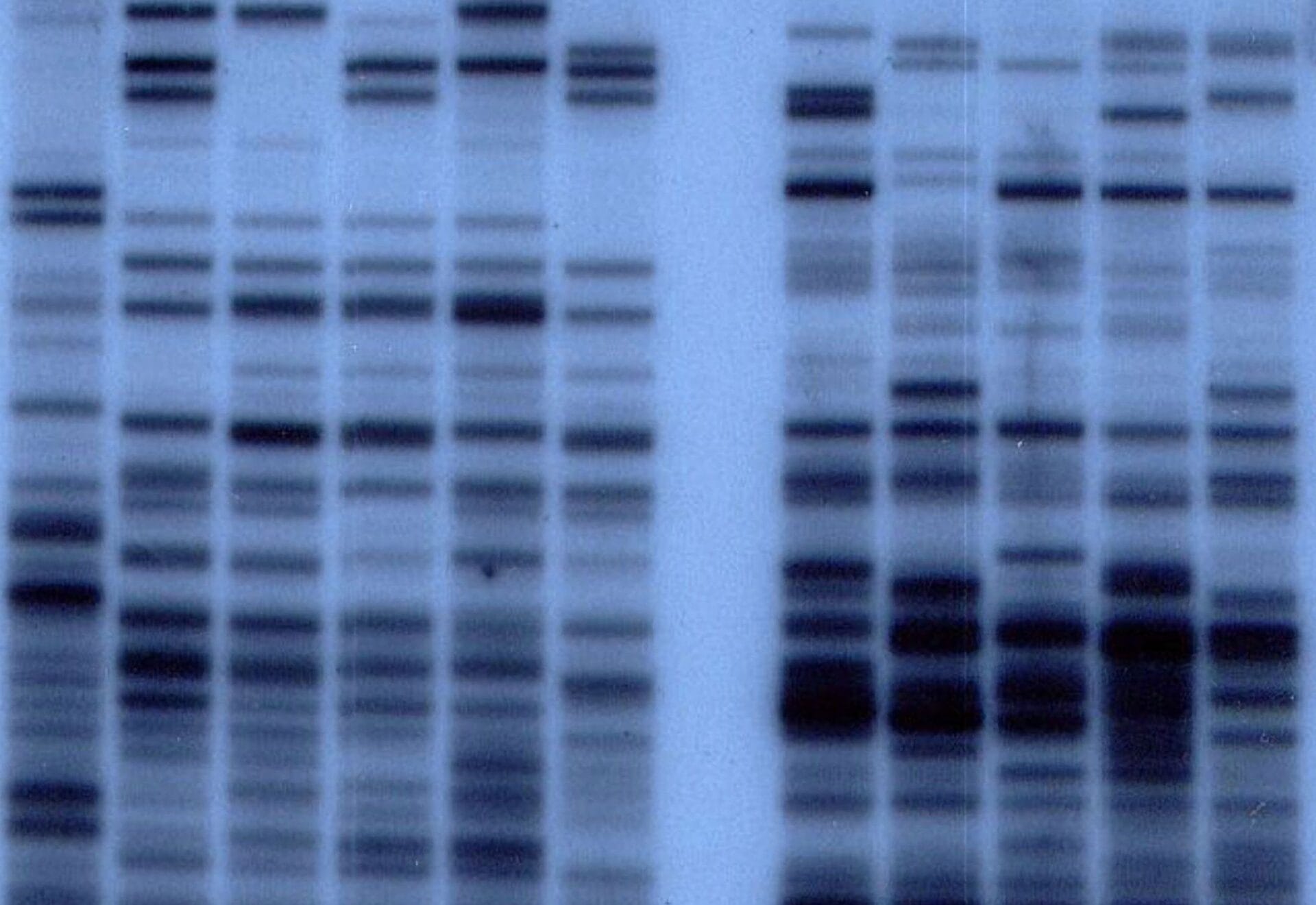
How DNA Profiling Works
Sample Collection: DNA is obtained from sources like blood, saliva, or hair.
Amplification with PCR: Specific STR regions are amplified using PCR, producing enough DNA for analysis.
Separation by Gel Electrophoresis: The amplified DNA fragments are separated based on size, creating a pattern of bands unique to the individual.
Hint
The reliability of DNA profiling increases with the number of STR markers analyzed.
Using more markers reduces the probability of a false match, enhancing the accuracy of the results.
Example
In a paternity test, the child's DNA profile is compared to that of the mother and the potential father.
If the child has bands not present in either the mother or the alleged father, another person must be the biological parent.
Note
In forensic investigations, analyzing 13 or more STR markers is standard practice, making it extremely unlikely for two unrelated individuals to have identical profiles.
Other Applications
1. Disease Diagnosis
PCR is used to detect genetic mutations linked to diseases like cystic fibrosis or sickle cell anemia.
It can also identify pathogens in infectious diseases, such as detecting viral RNA in COVID-19 testing.
Example
In COVID-19 testing, PCR amplifies viral RNA (converted to DNA) to detect the presence of the virus, even in very small quantities.
2. Evolutionary Biology
Gel electrophoresis helps compare DNA from different species to study evolutionary relationships.
By analyzing similarities and differences in DNA fragments, scientists can construct phylogenetic trees.
3. Environmental Monitoring
PCR and gel electrophoresis are used to detect DNA from endangered species in environmental samples, aiding conservation efforts.
These techniques can also identify microbial communities in soil or water, helping monitor ecosystem health.
Tip
Remember that PCR amplifies specific DNA sequences, while gel electrophoresis separates them by size. These complementary techniques are the backbone of many modern biological applications.
Self Review
Can you explain how PCR and gel electrophoresis work together in DNA profiling? What are some other applications of these techniques?
D1.1.6 directionality of dna polymerases (hl)
The Directionality of DNA Polymerases
Reading a book upside down and backward would be confusing and probably really hard.
DNA polymerases face a similar challenge during replication: they can only work in onedirection.
Tip
DNA polymerases always add nucleotides to the 3' end of a growing DNA strand.
The 5' and 3' Ends of DNA: What Do They Mean?
DNA strands have a specific directionality, defined by the structure of their nucleotides.
Nucleotide Structure
Each nucleotide consists of three parts:
A phosphate group
A sugar (deoxyribose in DNA)
A nitrogenous base (adenine, thymine, cytosine, or guanine)
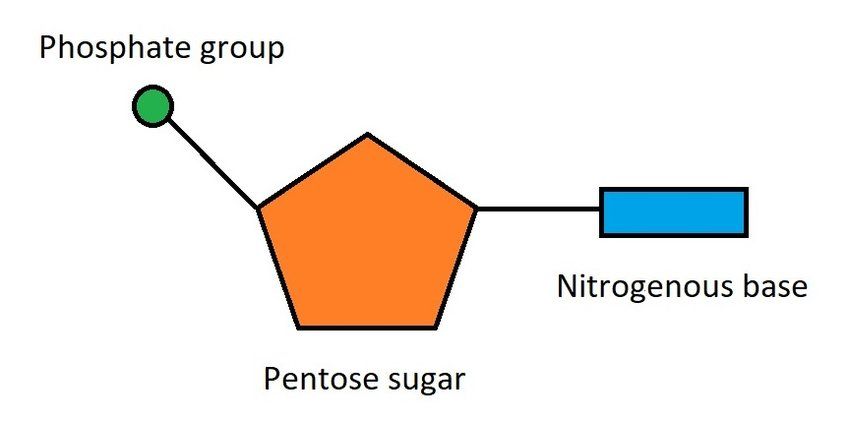
Note
The sugar molecule in DNA has carbon atoms numbered 1' to 5'.
These numbers help define the directionality of the DNA strand.
The 5' and 3' Ends
The 5' end of a DNA strand is where the phosphate group is attached to the 5' carbon of the sugar.
The 3' end is where the hydroxyl group (-OH) is attached to the 3' carbon of the sugar.
Analogy
Think of a DNA strand as a train track.
The 5' end is like the starting point of the track, marked by a signal post (the phosphate group).
The 3' end is the open end of the track, where new sections can be added (the hydroxyl group).
How DNA Polymerases Work
DNA polymerases are enzymes responsible for synthesizing new DNA strands during replication.
However, they have a strict rule: they can only add nucleotides to the 3' end of a growing strand.
Why Only the 3' End?
The addition of a nucleotide involves forming a covalent bond between the phosphate group of the incoming nucleotide and the hydroxyl group on the 3' carbon of the existing strand.
This reaction releases energy, making it possible for the bond to form.
Note
It's a common misconception that DNA polymerases can add nucleotides to either end of a strand. Remember, they onlywork in the 5' to 3' direction.
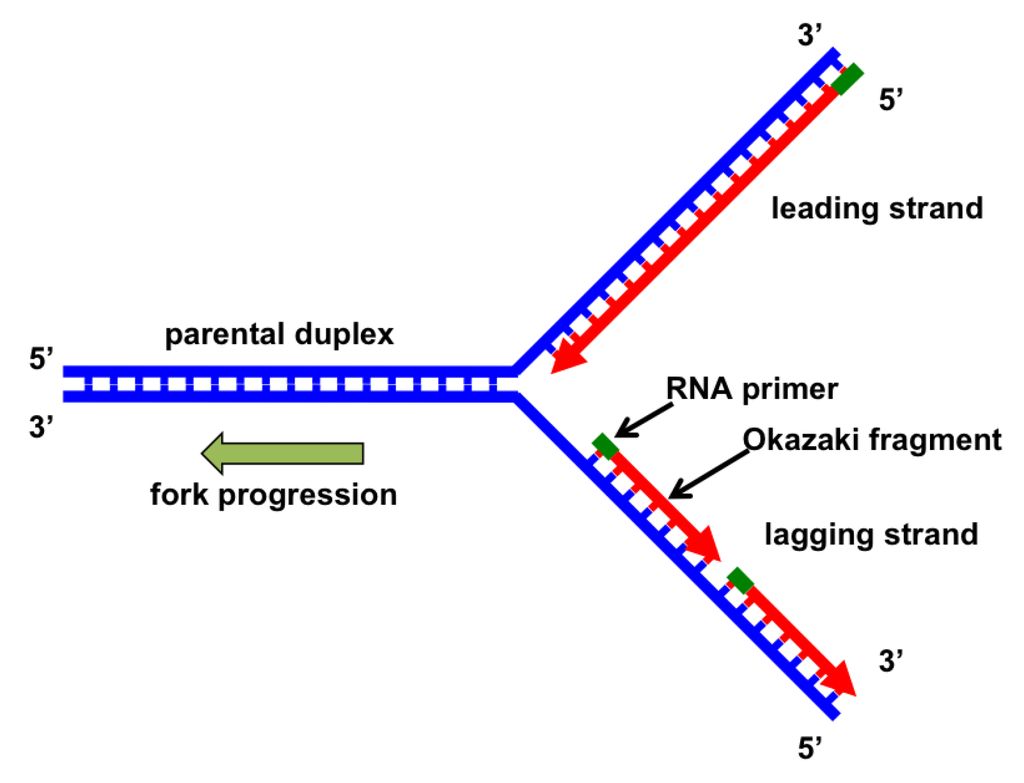
Implications for DNA Replication
DNA replication occurs at a structure called the replication fork, where the two strands of the double helix are separated.
Because the strands are antiparallel (running in opposite directions), DNA polymerases face a unique challenge:
On the leading strand, the polymerase can continuously add nucleotides in the 5' to 3' direction, moving toward the replication fork.
On the lagging strand, the polymerase must work in short segments, called Okazaki fragments, because it can only add nucleotides in the 5' to 3' direction, away from the replication fork.
Example
Imagine a construction crew building a road. One team works smoothly in a straight line (the leading strand), while another team has to build in small sections, moving backward (the lagging strand).
Why Directionality Matters
The directionality of DNA polymerases is crucial for several reasons:
Efficiency: Continuous synthesis on the leading strand ensures rapid replication.
Coordination: The lagging strand's fragmented synthesis requires additional enzymes, such as DNA ligase, to join Okazaki fragments.
Proofreading: DNA polymerases also proofread the newly synthesized strand, correcting errors to maintain genetic fidelity.
Tok
How does the directionality of DNA polymerases reflect the broader principle of structure determining function in biology?
Can you think of other biological processes where directionality is critical?
Key Takeaways
DNA strands have a 5' end (phosphate group) and a 3' end (hydroxyl group).
DNA polymerases can only add nucleotides to the 3' end of a growing strand.
This directionality leads to continuous synthesis on the leading strand and fragmented synthesis on the lagging strand.
Self Review
Why can DNA polymerases only add nucleotides to the 3' end of a DNA strand?
How does this limitation affect replication on the lagging strand?
D1.1.7 differences between replication on the leading strand and the lagging strand (hl)
Differences Between Replication on the Leading Strand and the Lagging Strand
DNA polymerase, the enzyme responsible for synthesizing new DNA, can only add nucleotides in the 5′ to 3′ direction.
This creates two distinct replication processes:
Continuous replication on the leading strand and discontinuous replication on the lagging strand.
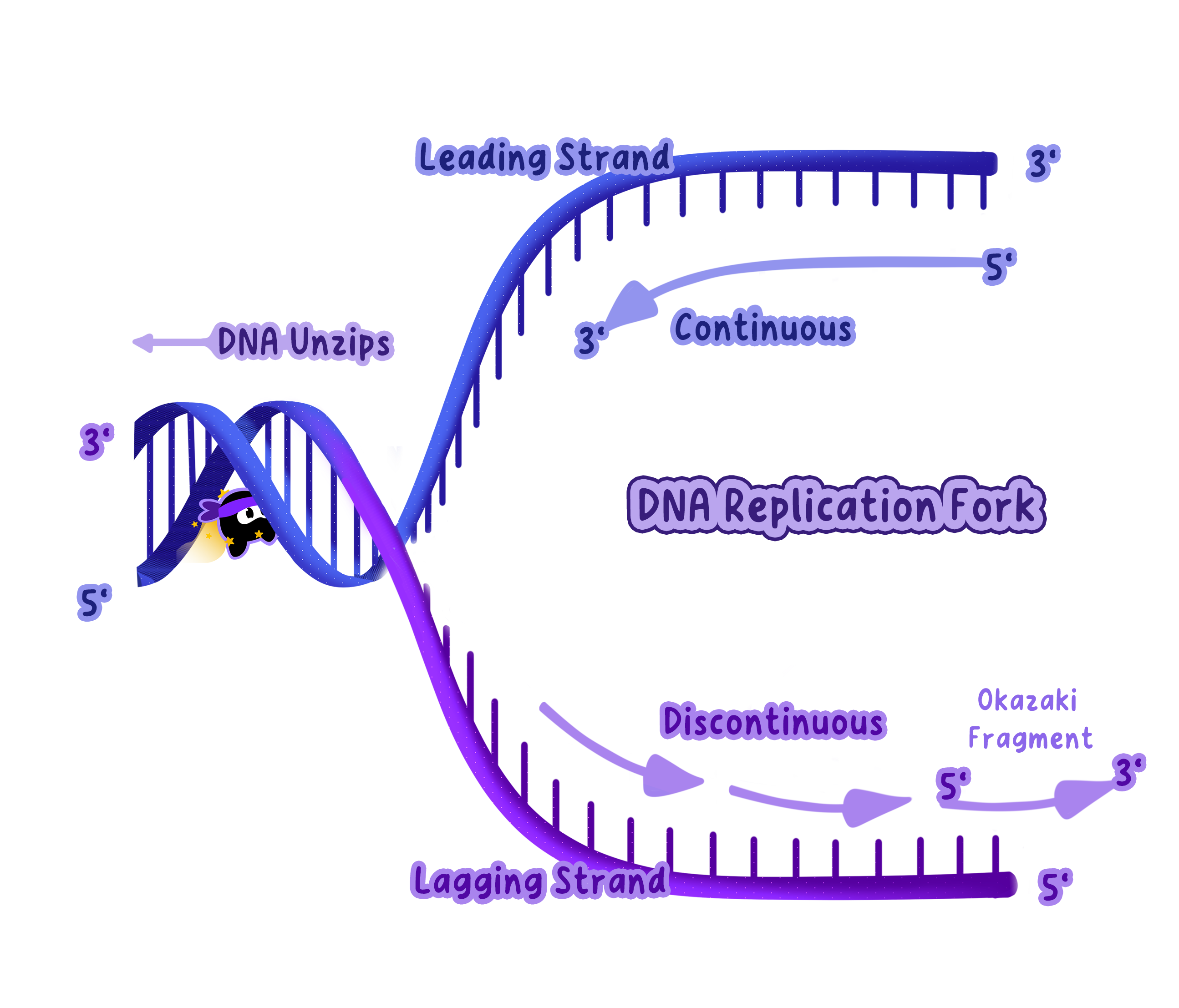
Note
DNA strands run in opposite directions, one in the 5′ to 3′direction and the other in the 3′ to 5′direction.
Continuous Replication on the Leading Strand
The leading strand is synthesized in the same direction as the replication fork's movement.
Single Primer Initiation: DNA primase adds an RNA primer at the start of the strand.
Continuous Synthesis: DNA polymerase III adds nucleotides without interruption, creating a smooth, unbroken strand.
Tip
Think of the leading strand as a car driving on a straight highway, moving steadily without stops.
Discontinuous Replication on the Lagging Strand
The lagging strand is synthesized in the opposite direction of the replication fork's movement, requiring a more complex process.
Multiple Primers: RNA primers are added at intervals along the strand.
Okazaki Fragments: DNA polymerase III synthesizes short segments of DNA, called Okazaki fragments, between primers.
Fragment Joining: DNA polymerase I replaces RNA primers with DNA, and DNA ligase connects the fragments into a continuous strand.
Tip
The lagging strand is like a car navigating a series of stop-and-go traffic lights, moving in short bursts.
Key Differences Between Leading and Lagging Strands
Feature | Leading strand | Lagging strand |
|---|---|---|
Template Strand | Uses the 3' to 5' template strand of DNA. | Uses the 5' to 3' template strand of DNA. |
DNA Polymerase Activity | DNA polymerase moves in the same direction as the replication fork. | DNA polymerase moves in the opposite direction to the replication fork (backstitching). |
Continuous/Discontinuous | Continuous | Discontinuous |
Okazaki Fragments | Not formed | Formed |
Warning
A common misconception is that the lagging strand is synthesized more slowly.
In reality, both strands are synthesized simultaneously, but the lagging strand's discontinuous process makes it appear slower.
Tok
How does the antiparallel structure of DNA reflect the broader theme of symmetry and asymmetry in biology?
Can you think of other biological processes where directionality plays a critical role?
D1.1.8 dna primase, dna polymerase i, dna polymerase iii and dna ligase in replication (hl)
Functions of DNA Primase, DNA Polymerase I, DNA Polymerase III, and DNA Ligase in Replication
DNA replication is a highly coordinated process involving multiple enzymes, each with a specific role.
Note
This section focuses on the prokaryoticsystem, where replication is simpler but follows the same fundamental principles as in eukaryotes.
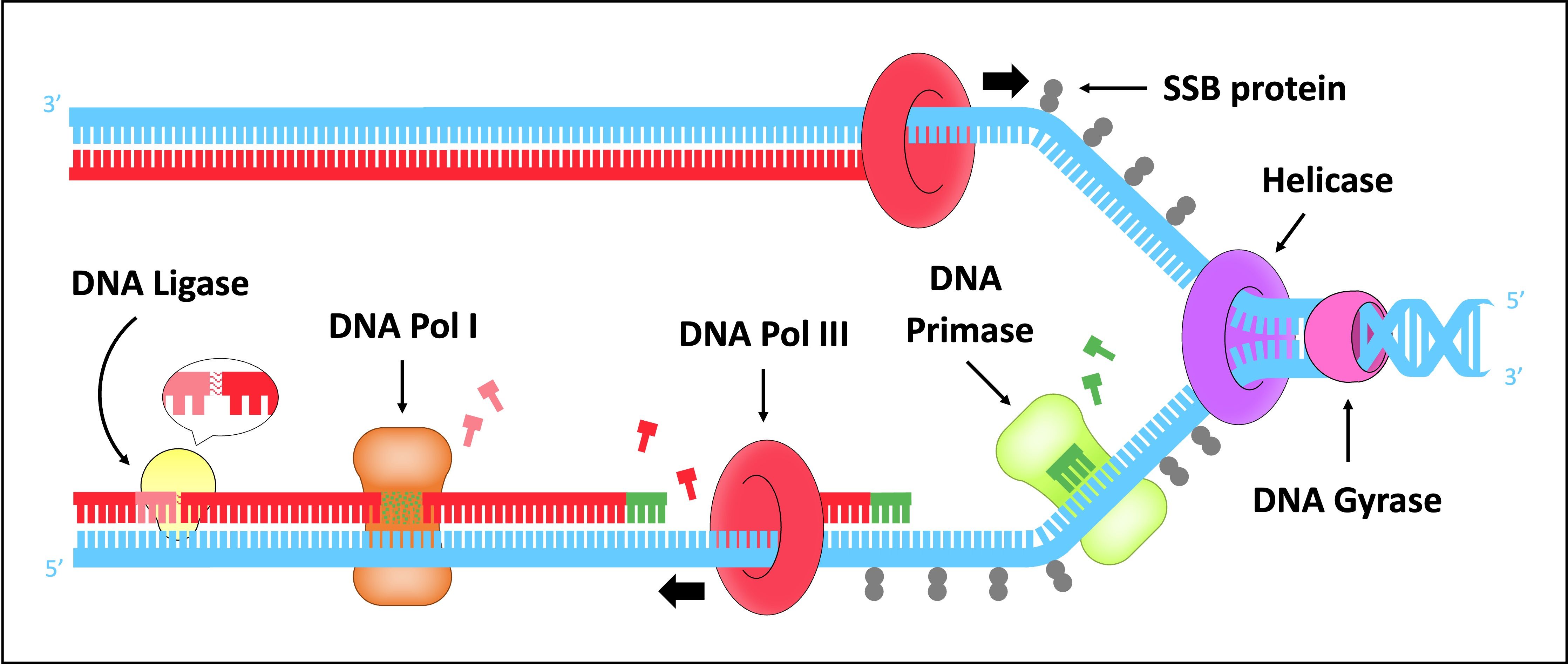
DNA Primase: Laying the Foundation
DNA polymerases can't start synthesizing a new strand on their own.
They need a starting point, a short segment of RNA called a primer.
Definition
PrimasePrimase is an enzyme that synthesizes a short RNA primer to provide a starting point for DNA polymerase III during replication.
Key Functions of DNA Primase
Synthesizes RNA Primers: DNA primase creates a short RNA primer (about 10 nucleotides long) on the template strand.
Provides a 3′-OH Group: The RNA primer offers a free 3′-OH group, which is essential for DNA polymerase III to begin adding DNA nucleotides.
Multiple Primers on the Lagging Strand: On the lagging strand, DNA primase adds a new primer for each Okazaki fragment.
Tip
Remember, DNA polymerases can only extend a strand, they cannot start one from scratch. This is why primase is essential.
DNA Polymerase III: The Main Builder
DNA polymerase III is the workhorse of DNA replication, responsible for assembling the majority of the new DNA strand.
It is the primary enzyme responsible for synthesizing new DNA strands during replicationin prokaryotes.
Key Functions of DNA Polymerase III
Synthesizes DNA in the 5′ to 3′ Direction: Adds nucleotides to the growing DNA strand, using the template strand as a guide.
High Processivity: Can add thousands of nucleotides without detaching, ensuring efficient replication.
Proofreading Ability: Checks each nucleotide after adding it, removing and replacing incorrect ones to minimize errors.
Note
Don't confuse DNA polymerase III with DNA polymerase I.
While both are involved in replication, their roles are distinct.
DNA Polymerase I: The Cleaner
Once DNA polymerase III has synthesized the new DNA, the RNA primers need to be removed and replaced with DNA.
This is where DNA polymerase I comes in.
An enzyme that removes RNA primers and replaces them with DNA nucleotides during replication.
Key Functions of DNA Polymerase I
Removes RNA Primers: Acts as an exonuclease to remove RNA primers from the newly synthesized strand.
Replaces RNA with DNA: Fills in the gaps left by the removed primers with DNA nucleotides.
Works in the 5′ to 3′ Direction: Like DNA polymerase III, it adds nucleotides in the 5′ to 3′ direction.
Example
Imagine a construction crew building a bridge.
DNA polymerase III lays down the main structure, while DNA polymerase I replaces temporary scaffolding (RNA primers) with permanent materials (DNA).
DNA Ligase: The Final Connector
Definition
LigaseLigase is an enzyme that joins Okazaki fragments on the lagging strand by forming covalent bonds between adjacent DNA nucleotides.
After DNA polymerase I replaces the RNA primers with DNA, there are still small gaps in the sugar-phosphate backbone of the DNA strand.
DNA ligase seals these gaps.
Key Functions of DNA Ligase
Seals Nicks in the DNA Backbone: Forms phosphodiester bonds between adjacent DNA nucleotides, completing the sugar-phosphate backbone.
Joins Okazaki Fragments: Ensures that the short DNA fragments on the lagging strand become a continuous strand.
Requires Energy: Uses ATP to catalyze the formation of these bonds.
Analogy
Think of DNA ligase as a glue that holds the pieces of a puzzle together, ensuring the DNA strand is complete and functional.
Coordination of Enzymes in DNA Replication
Primase creates RNA primers to initiate synthesis.
DNA polymerase III extends the DNA strand from the primer.
DNA polymerase I removes RNA primers and replaces them with DNA.
DNA ligase seals the gaps, forming a continuous DNA strand.
Self Review
Can you explain the role of each enzyme in DNA replication? How do they work together to ensure the process is efficient and accurate?
Why These Enzymes Matter
Understanding the roles of these enzymes is crucial for grasping how DNA replication maintains genetic continuity.
These enzymes ensure that DNA replication is accurate and efficient, allowing life to persist across generations.
Tok
How does the interplay of these enzymes reflect the broader concept of collaboration in biological systems? Can you think of other processes where multiple components work together to achieve a common goal?
D1.1.9 dna proofreading (hl)
DNA Proofreading: Ensuring Accuracy in Replication
Imagine copying a 6-billion-letter book by hand.
Even a single typo could change the meaning of a sentence.
DNA replication faces a similar challenge: copying billions of base pairs with near-perfect accuracy.
Analogy
Think of DNA polymerase III as a meticulous editor, scanning each word (or nucleotide) for errors and correcting them on the spot.
The Role of DNA Polymerase III in Proofreading
DNA polymerase III is the primary enzyme responsible for synthesizing new DNA strands.
But it does more than just add nucleotides, it also proofreads its work.
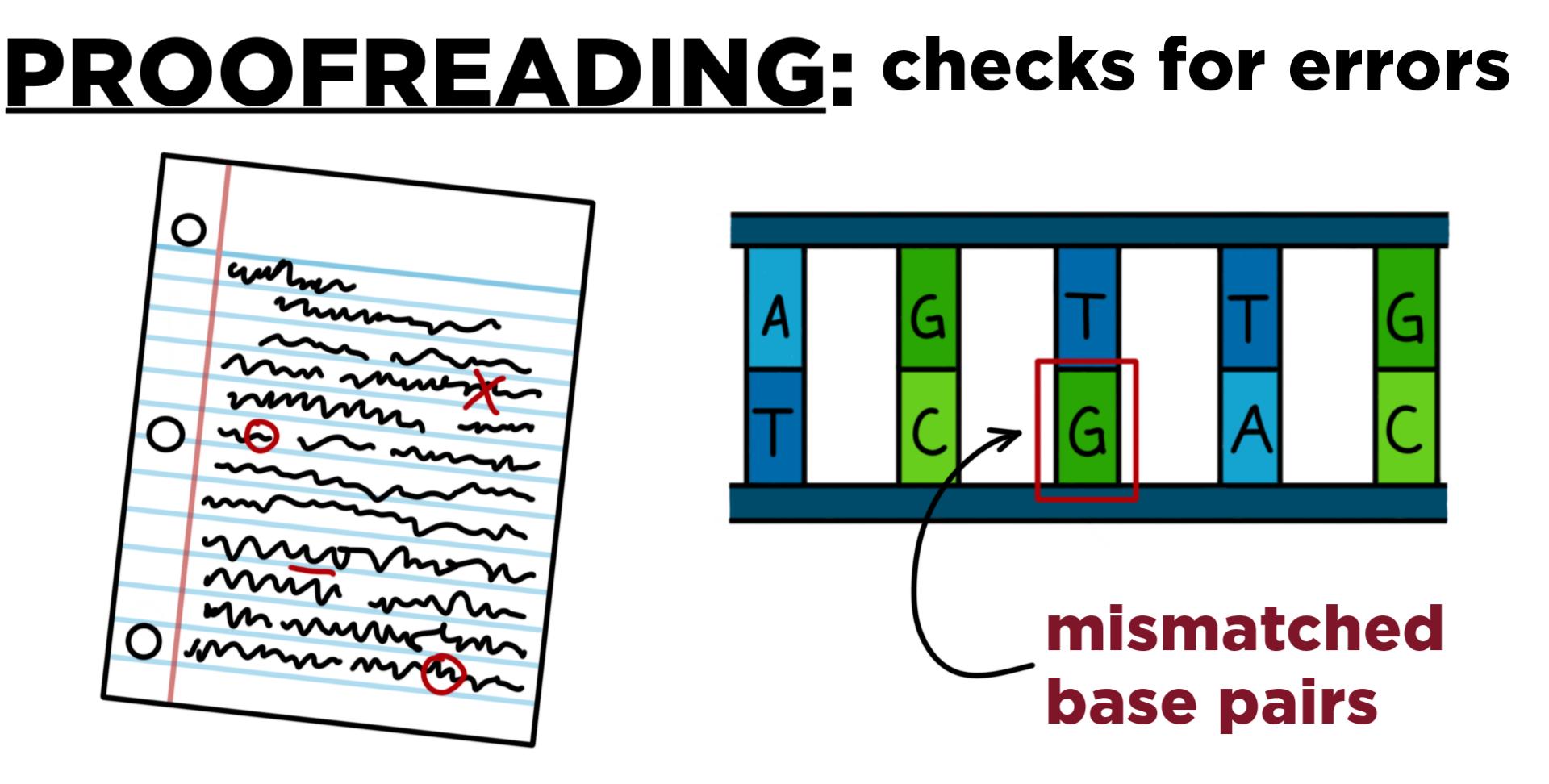
How Proofreading Works
Mismatch Detection
DNA polymerase III checks each newly added nucleotide.
If the base pairing is incorrect (e.g., adenine paired with cytosine instead of thymine), the hydrogen bonds between the bases are unstable, signaling an error.
Exonuclease Activity
DNA polymerase III has a built-in exonuclease function that removes nucleotides from the 3′ end of the growing strand.
When a mismatch is detected, the enzyme pauses, reverses its direction, and removes the incorrect nucleotide.
Correction and Resumption
After removing the incorrect nucleotide, DNA polymerase III adds the correct one and resumes DNA synthesis in the 5′ to 3′ direction.
Tip
Remember: DNA polymerase III can only remove nucleotides from the 3′ end, which is why replication always proceeds in the 5′ to 3′ direction.
Why Proofreading Matters
Without proofreading, errors in DNA replication could lead to mutations, which might disrupt protein function or cause diseases like cancer.
By correcting mismatches, DNA polymerase III ensures that the new DNA strand is nearly identical to the template strand.
Example
In humans, the proofreading activity of DNA polymerase III reduces the error rate to about 1 in 10 million base pairs.
Additional repair mechanisms further decrease this rate to 1 in 10 billion.
The Limits of Proofreading
While DNA polymerase III is highly effective, it isn't perfect.
Some errors escape detection, especially if they occur after the enzyme has moved past the mismatch.
However, other repair mechanisms, such as mismatch repair, can correct these errors after replication.
Note
Don't confuse DNA polymerase III with DNA polymerase I. While both enzymes are involved in replication, DNA polymerase I primarily replaces RNA primers with DNA and does not play a major role in proofreading.
Applications and Implications
The accuracy of DNA replication, supported by proofreading, is essential for genetic stability across generations.
Tok
How does the ability of DNA polymerase III to proofread relate to the broader concept of error correction in other fields, such as computer science or linguistics?
Self Review
Can you explain how DNA polymerase III detects and corrects a mismatched base during replication?