4. Antibodies
Antibody molecule –
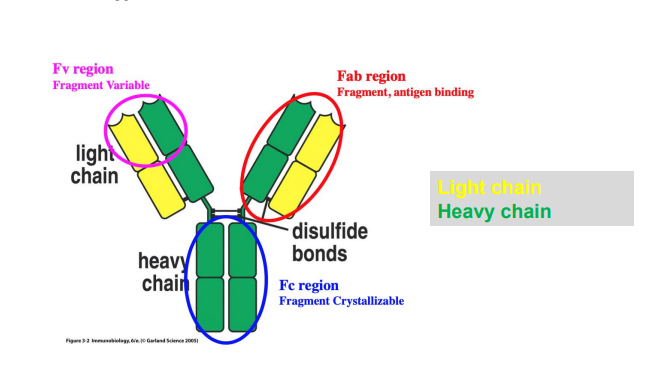
- An antibody unit consists of four polypeptide chains: two identical heavy chains and two identical light chains connected by disulfide bonds.
- Different B cells produce different antibodies.
- A protein of about 150 kDa molecular weight
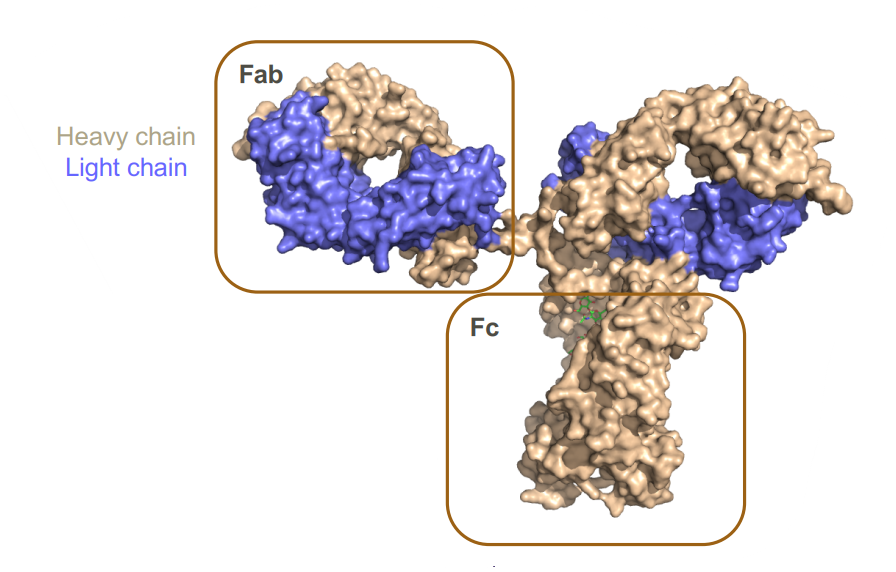
- The fragment antigen-binding region (Fab region) is a region on an antibody that binds to target molecules (antigens). It is composed of one constant and one variable domain of each of the heavy and the light chain (carries the most diversity).
- The fragment crystallizable region (Fc region) is the tail region of an antibody that allows other components to bind (like protein)
- Relatively little diversity
- Defines antibody isotype (IgA, IgD, IgE, IgG, IgM)
- Carries carbohydrates
- Recruits effector function
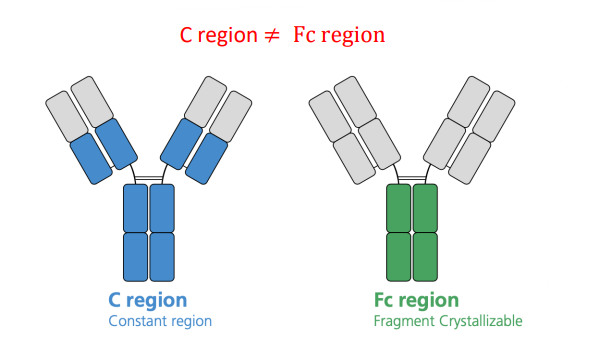
C region is not variable as the variable region
Epitope- An epitope is the part of an antigen recognised by antibodies (or T cells)
- Different antibodies may recognise very similar epitopes
- Kinetics and binding interactions may be different
- Different antibodies may recognise very different epitopes
![]()
Overlapping and non-overlapping epitopes
Overlapping epitopes: two or more epitopes on an antigen that share some portion of their molecular structure. Antibodies that recognize overlapping epitopes may compete for binding (the antibodies block each other’s binding sites).
Non-overlapping epitopes: do not overlap → may be simultaneously recognised by antibodies without interfering with each other’s binding.
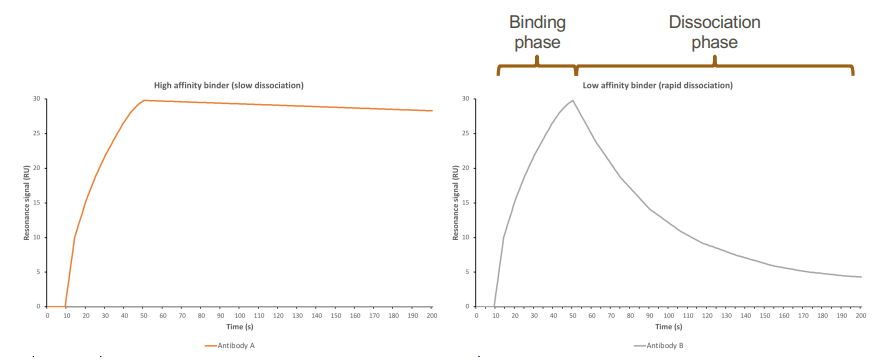
Affinity – reaction rate kinetics
• Typically, stable high-affinity interactions are beneficial to development high-functionality
• SPR (surface plasmon resonance) technology
• KA=kass/kdiss (typical range of t1/2: seconds – many hours)
kass=speed of association
kdiss=speed of dissociation
High-affinity binder → slow dissociation
Low affinity binder → rapid dissociation
Polyclonal antibodies– a mixture of antibodies found e.g. in serum
– origin in many different antibody-producing B cells
– recognize different parts (epitopes) of the target (target is called an antigen).
– antibodies recognise the antigen with different binding strength (affinity). Some will bind more strongly
– different preparations will be variable with respect to each other
• Risk of viral contamination of human antibodies derived from human subjects (eg human proteins and viruses from humans)
• Risk of immune response against animal-derived antibodies
• High batch variability (problematic for production)
• Impossible to standardize completely
• Bind many different epitopes with different functionalities
• Bind epitopes with different affinities and reaction rate kinetics
• May bind many different antigens
There is commonly a need to define one or a small set of antigens to be targeted by one or a small set of antibodies
Monoclonal antibody – a single antibody
– origin in a single antibody-producing B cells
– recognize one part (epitope) of the target (the antigen) with a single, defined binding strength (affinity)
– different preparations have the potential to be very similar to each other
• May be of animal or human origin and the latter is likely to not induce substantial immune response against the antibody during treatment
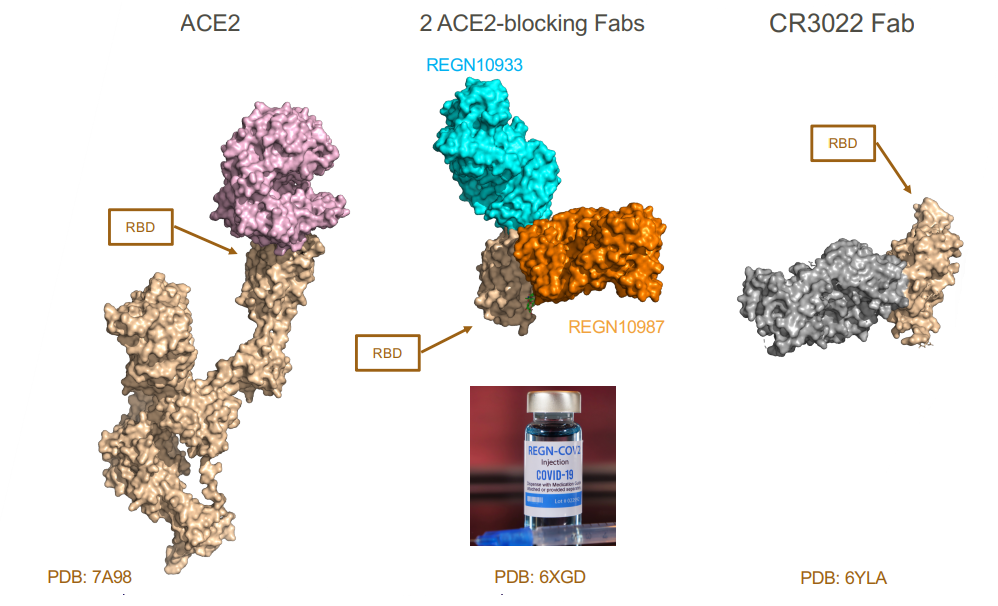
Binding of virus-neutralizing antibodies to the spike protein of SARS-CoV-2 – different mechanisms of action
They bind different epitopes. They prevent viruses from infecting and they neutralise
It changes the structure so it doesn’t function anymore
Defining an appropriate antigen to target in a therapeutic application
- Pre-existing functional knowledge
- can build on the knowledge on receptor binding, function etc.
- Target-based approaches (to decide on potential target)

- Gene expression profiling
- Microarrays
- Next generation sequencing (NGS)
- Proteomics
- to define possible targets
- Gene expression profiling
- Function-based approaches
- if you don’t have detailed knowledge about the target of interest. Look at different antibodies and see which would function.

Antibody-druggable targets
• Antigens exposed to the extracellular environment
– Membrane proteins/receptors
– Extracellular matrix
– Soluble mediators (cytokines, chemokines, etc.)
– Viral and bacterial surface proteins etc.
• In general not intracellular proteins, transcription factors, intracellular domains of membrane proteins etc.
Development and screening cascade
Affinity maturation is the process by which TFH cell-activated B cells produce antibodies with increased affinity for antigen during the course of an immune response.
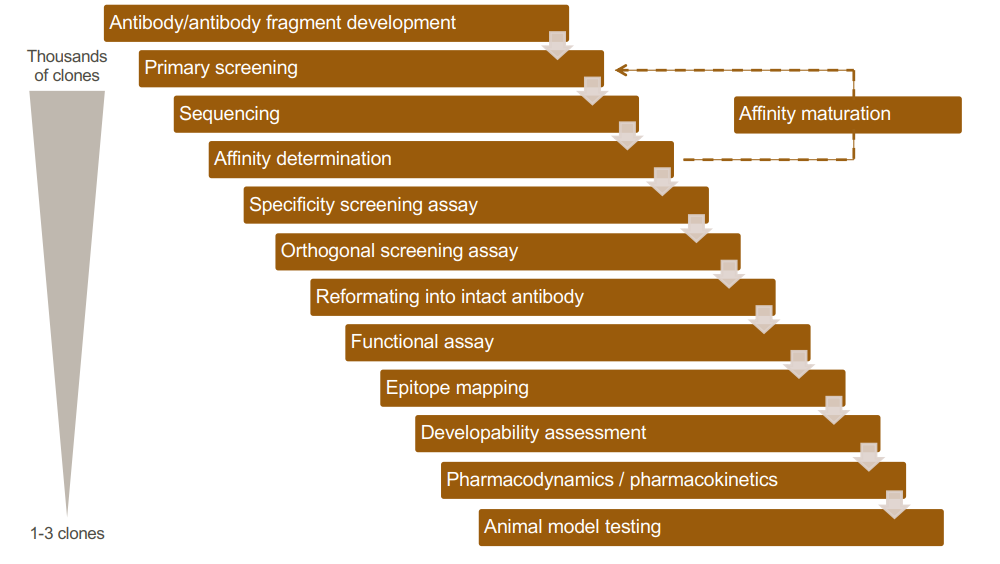
Primary screening phase- Evaluate the binding affinity and specificity of antibodies to the target antigen. This step helps identify lead candidates for further development.
• Immunochemical assays (e.g. ELISA)
• High throughput flow cytometry screening assay (can my antibody bind to these cells that carry my target or not)
• Next Generation Sequencing (look for antibodies that are enriched suggesting that they have some binding ability to the target)
Sequencing- Determine the amino acid sequence of the selected antibodies. This information is crucial for downstream development and manufacturing.
Affinity determination- Measure the strength of binding between the antibodies and the target antigen. Want to identify variants that stay bound very well to the target.
Specificity screening assays- Assess the specificity of antibodies by testing their binding to related or structurally similar antigens to ensure selection for the intended target. The goal is to identify antibodies that specifically recognize the target of interest, thus minimizing the risk of off-target effects.
Cross-reactivity- when an antibody or a T cell receptor can recognize and bind to more than one distinct target. Can be beneficial (can respond to a more wide range of pathogens) but may also lead to unwanted immune responses such as autoimmune reactions.
• Cross-reactivity to model animal’s antigen may be required for
– Functional validation
– Assessment of side effects
» On-target toxicity
» Off-target toxicity (bind to something else that is not expected)
Orthogonal screening assay- Further validation of the binding specificity and characteristics of the antibodies. Performed on antigen on cells by flow cytometry.
Binder to function transition
• Design of format to develop antibody with appropriate functionality
– Antibody with long half life
» binding to FcRn: recycling of antibodies → extended half-life in circulation.
» PEGylation of smaller antibody fragments: (increases size → more difficult to secrete)
– Antibody with short half life
– Antibody that bind cellular Fc receptors for cellular effector function
– Antibody that bind complement (a complex system of protein in the blood that induces inflammation and punching holes in cells)
– Antibody that does not bind cellular Fc receptors or complement: helps minimize unintended immune responses.
– Antibody that carries payload (toxin, 125I,177Lu (radioactive isotop))
Functional assay- Evaluate the ability of the antibodies to elicit the desired biological response or function
Examples:
- Cell killing
- Apoptosis
- Complement fixation
- ADCC (Antibody-dependent cellular cytotoxicity)
- Inhibition of migration
- Checkpoint inhibition
Epitope mapping- Define where on the molecule that antibody binds. Understanding epitope specificity is important for understanding antibody mechanism
• Epitope mapping assays
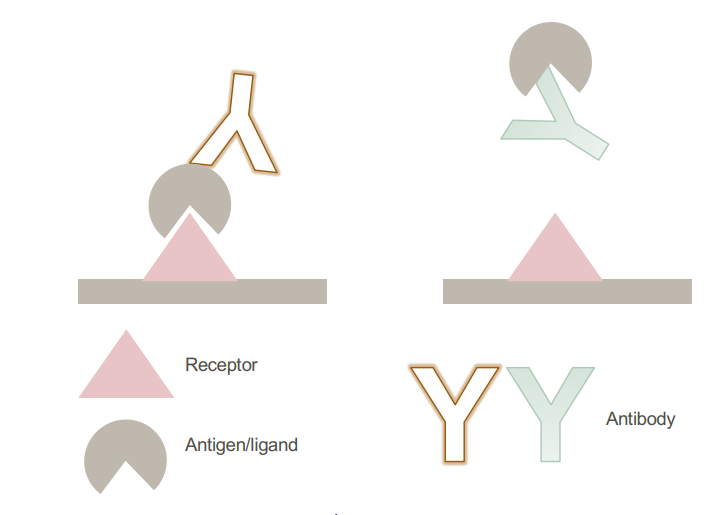
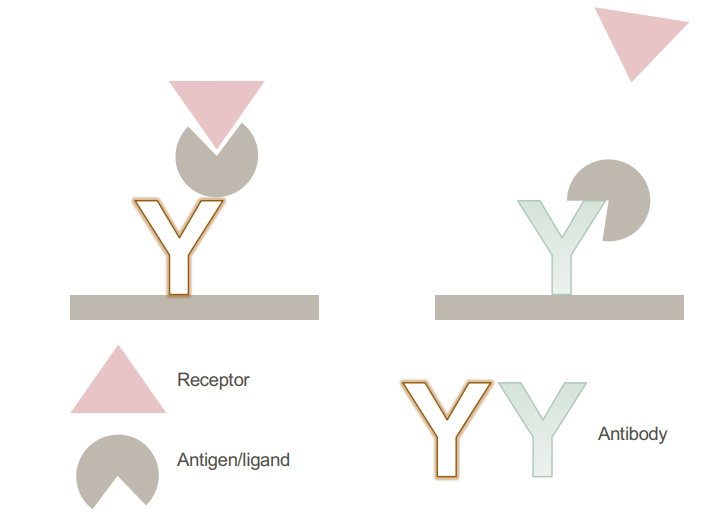
• Sandwich assay
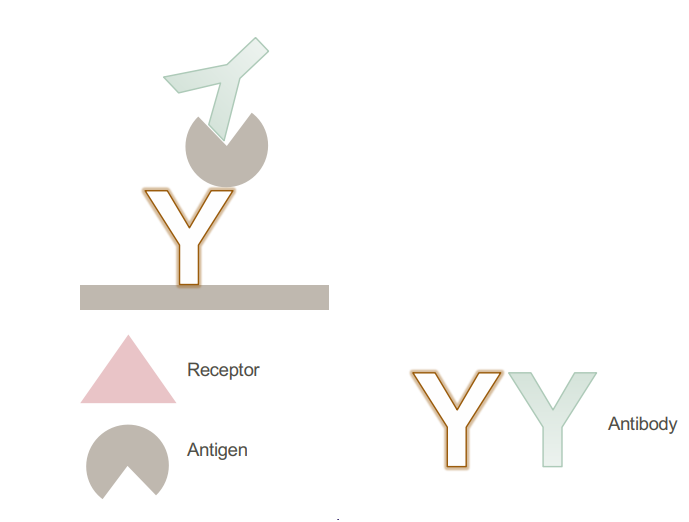
Epitope mapping assays
– Immunoassay (ELISA, FRET assays, blocking of function etc.)
– Label-free assays e.g. surface plasmon resonance
– HDX-MS (hydrogen-deuterium exchange mass spectrometry)
Which residues of protein that are protected from interaction with water and that tells which binds to the antibody
– X-ray crystallography
Developability – products with well behaving properties
- Identifying problematic features for drug development like aggregation. Biophysical problems.
- Evaluation of factors such as stability, solubility and potential immunogenicity.
Animal model testing- Asses the therapeutic potential and safety of the antibodies in relevant animal models.
Antibody mechanisms of action
e.g.
- Blocking
- Signalling
- Payload delivery (e.g. radioactivity, drugs)
- Inhibition of migration
- Induction of apoptosis (a cellular suicide process)
- Recruitment of cell killing immune mediators (NK cells, complement etc.)
- Cell crosslinking by bispecific antibodies (Bispecific antibodies = binds to different targets)
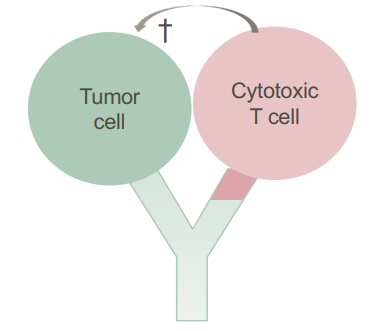
- Cell killing through CAR T cells (will bind to tumor cells)
– an ATMP
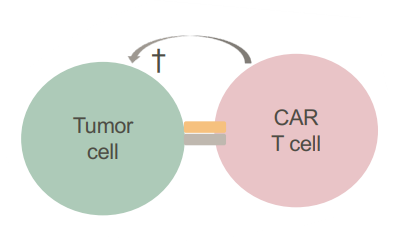
- Checkpoint inhibitors – allowing immune control to be active
Check-point inhibitors- Immune checkpoints are key regulators of the immune system that when stimulated can weaken the immune response to an immunologic stimulus. Tumour cells can stimulate immune checkpoints → weaken immune response to protect themselves from attack
Monoclonal antibodies
• Immunization – to invoke production of specific antibodies (typically mice, now also e.g. rabbits)
• Immortalization of antibody-producing capability through a fusion of antibody-producing B cells with immortal cell lines (survival and antibody-producing properties)
• Animal antibodies will be seen as foreign and induce an antidrug immune response when used in therapeutic setting
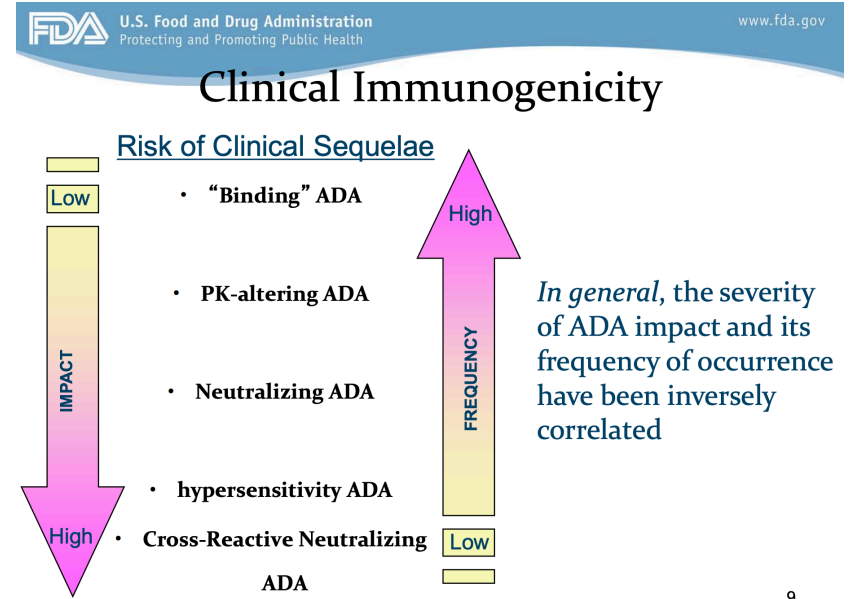
Anti-drug antibodies (ADA)
- May limit drug use and efficacy and induce toxicity
- In general, the severity of anti-drug antibodies (ADA) impact and the frequency of occurrence have been inversely correlated to impact

Human antibodies technology
- The content of non-human sequences may influence an antibody’s ability to induce anti-drug antibody responses that will limit the application of the antibody drug
- Human antibody technology – immortalization of antibody-producing human B cells
– Hybridoma: cell line created by the fusion of two different types of cells (B cells and myeloma cells)
– Single cell sequencing
– High throughput single cell sequencing, e.g. 10X genomics chromium technology in combination with NGS
The animal/human antibody approach
- Humanized mice – immunization of mice that carry human immunoglobulin genes
- Humanization – redesign of animal (mouse) antibody into protein with a human-like sequence
- Chimerization – redesign of animal (mouse) antibody into a molecule with partial human sequence
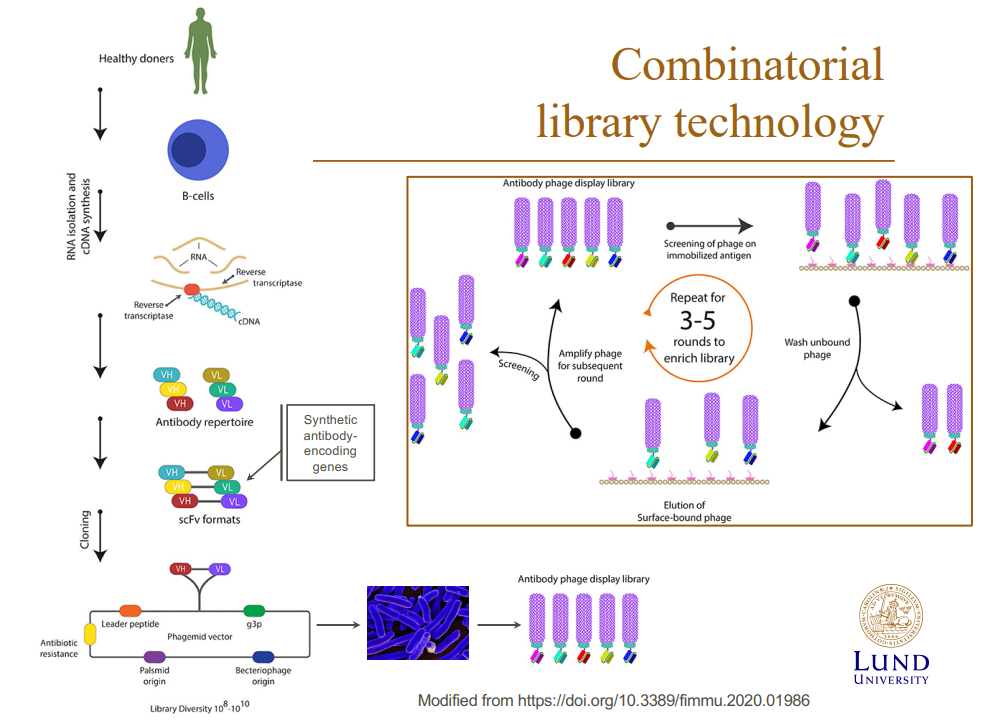
- Combinatorial library approaches
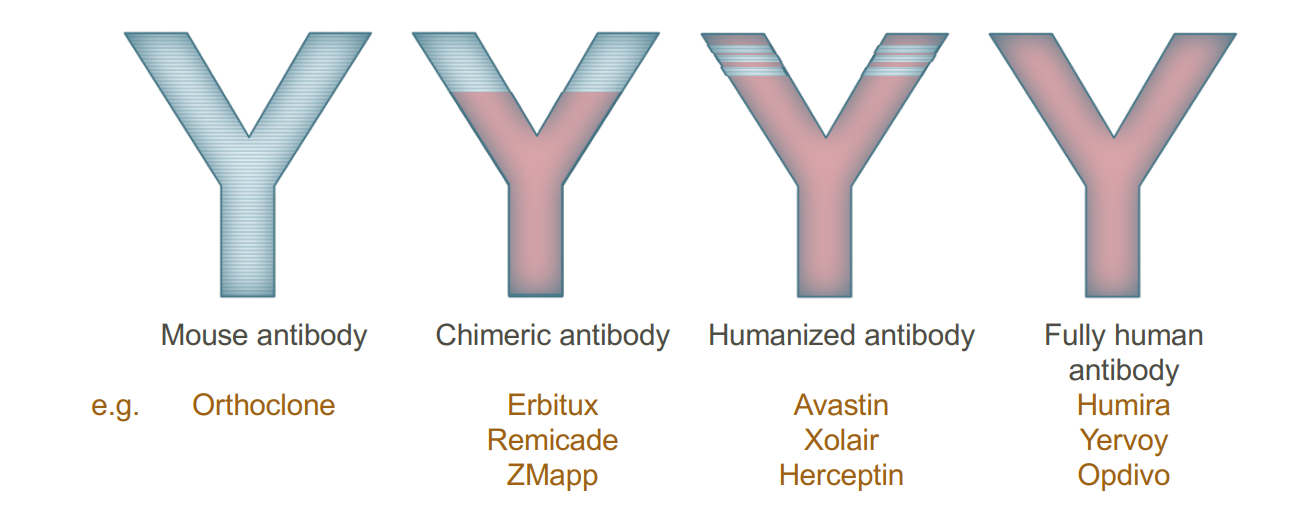
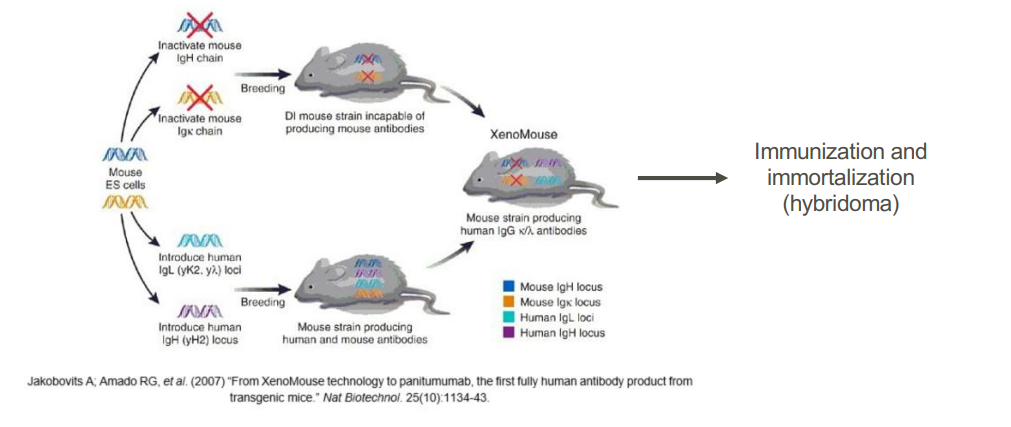
The animal/human antibody approach
• Mouse strains that produce human antibodies
• Hybridoma technology to develop a human antibody producing cell line
• Can be used to produce antibodies specific for targets against which there is no immune response in humans
Antibody phage display library (Combinatorial library technology)-