Microbial Diversity
The tree of life is one of the key organising principles in biology.
Ernst Haeckel discovered, described and named thousands of species and designed a tree of animals, plants and protists.
Robert Whittaker added unicellular organisms such as cyanobacteria to the tree and organised the organisms in the tree into photosynthetic, organisms that absorbed their nutrients and those that took in nutrients by eating.
Carl Woese was able to define archaea which he called the third domain of life, the other two being bacteria and eucarya. He did this through molecular phylogeny, realising that the 16S ribosomal RNA shows evidence of a universal ancestor.
Until recently, we have been unable to study abundance and diversity of microbes in the environment due to lack of tools which are needed. Traditional taxonomy and systematics relied on morphology and biochemistry (appearance etc). However microbes are very small and often look similar, and some have so far been impossible to cultivate.
Phylogenetics is the science of examining evolutionary relationships of a group of organisms eg aligning gene sequences from different species and looking for differences between them. This allows the relationship between two species to be inferred.
When selecting genes for use as a phylogenetic marker, we want them to be orthologous, present in all species you’re interested in comparing, conserved but with observable differences, and slow and steady to evolve (purifying/negative selection → stabilising selection).
Orthologous = descended from the same ancestral sequence and separated by a speciation event - vertical descent.
Vertical descent is the vertical line of descent which occurs when all microbes arise as the result of parental fission.
Many genes can be used as phylogenetic markers with variable results. Whole genome analysis has become much cheaper and quicker since its discovery and is changing the view on single gene use for phylogenetics.
The 16S rRNA gene is widely used for phylogenetic studies. It’s the RNA component of the 30S subunit in the ribosome and recognises the Shine-Dalgano sequences of the promoter. It’s highly conserved across all life forms meaning we can develop universal PCR primers to amplify the gene, although in organisms with introns there’s still conservation. The 16S rRNA gene can be put through high throughput methods using next gen sequencing - amplicon sequencing/metagenomics. The 16S rRNA gene has revealed large amounts of diversity in the environment.
Metagenomics is the isolation of cells to extract DNA, which is then fragmented and sequenced into ‘raw reads’. These raw reads are aligned and then annotated (gene function is named and what part of the gene sequence it is).
Genomics has led microbiologists to a two domain model for the tree of life, in which archaea and eukaryotes are actually part of the same branch. For the study which determined this (Hug et al, 2016), ribosomal protein genes were concatenated (linked) with the 16S rRNA gene. By using candidate phyla and thousands of 16S rRNA genes, it was determined that eukaryotes aren’t too genetically different from archaea and that bacteria actually dominate the tree by a significant amount. This study altered the way that we view the origins of cellular life.
LUCA = last universal common ancestor, meaning every living organism on the planet is a descendent of LUCA. We are unsure of whether LUCA was a single cell or a community of populations, but we know it had all the universal genes that we see today. In the ‘tree of life’, we call LUCA the root but the true roots are the chemistry and cells which LUCA originated from - LUCA is NOT the first cell.
The Earth is 4.6 billion years old. For the first two billion years, all life was microbial. All organisms had anoxic metabolisms such as methanogenesis due to the anoxic atmosphere of mainly nitrogen and carbon dioxide. Anoxic phototrophs evolved 3.5 billion years ago which gave way to oxygenic phototrophic cyanobacteria 2.5 billion years ago. Oxygen dependent metabolisms were then able to develop and eventually multicellular life forms. We have evidence of cellular life 3.8 to 3.9 million years ago.
Viruses are ancient forms but are not included in the tree of life. There are two main hypotheses for the emergence of viruses: genome reduction to the point of an obligate intracellular parasite, or genome escape - aggregations of genes that somehow escaped cellular regulation. Both of these hypotheses have issues and neither is widely adopted. Bacteriophage genome based evidence has suggested that the viral genome structure is ancient, from before bacteria and archaea split apart. The viral genome is mosaic in structure, meaning modules recombine and exchange.
Eukaryotic diversity on Earth was estimated at around 8.7 × 10^6 species total. 10g soil contains 10^10 bacterial and archaeal cells, an estimated 8.3 × 10^6 species.
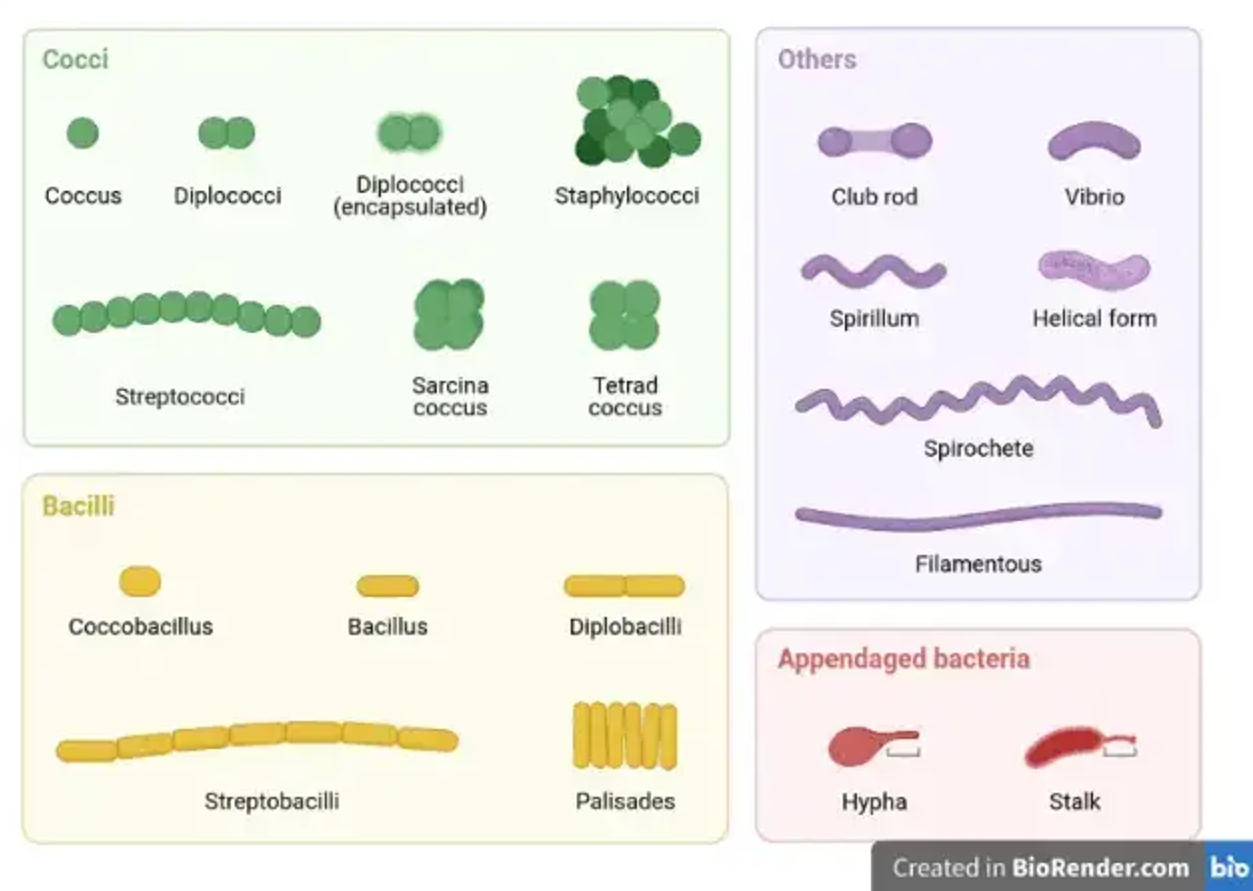 There’s great variety of morphological phenotypes in bacteria and archaea, and there’s often plasticity shown in the morphology. The growth form of the microbe often reflects the organism’s biology and lifestyle. Bacteria may be cocci, bacilli, appendaged or ‘other’. Examples of cocci are streptococcus and staphylococci. Examples of bacilli are bacillus, coccobacillus and palisades. Example other bacteria are club robs, vibrio and helical.
There’s great variety of morphological phenotypes in bacteria and archaea, and there’s often plasticity shown in the morphology. The growth form of the microbe often reflects the organism’s biology and lifestyle. Bacteria may be cocci, bacilli, appendaged or ‘other’. Examples of cocci are streptococcus and staphylococci. Examples of bacilli are bacillus, coccobacillus and palisades. Example other bacteria are club robs, vibrio and helical.
The genome is the full complement of genes for an organism. Haemophilus influenzae was the first free-living organism to have its genome sequenced in 1995. Escherichia coli is the free living organism with the best studied genome. E. coli’s genome is made up of 4.4 million base pairs (4.5 Mbp), which make up around 4300 genes. E. coli’s coding density is 1000 genes/Mbp.
The human genome is 6.2 Gpb, where 1 Gigabase is 1 billion base pairs. We have around 20 to 25 thousand protein coding genes and the same number of non-protein coding genes. Human coding density is 10 genes/Mbp.
The axolotl genome has 32 Gbp with 23,500 protein coding genes.
Bacterial and archaeal genomes are generally one single circular DNA molecule but there are lots of exceptions such as linear or multiple chromosomes and plasmids. Bacterial genomes are generally haploid and encode all genes required for organism assembly. Genes are often in operons and there’s little non-coding DNA between genes.