DNA Modification Enzymes
Purpose of Manipulating DNA Molecules
Gene Cloning: Creating multiple copies of a particular DNA segment for various applications.
Functional Analysis of Genes: Studying the function and regulation of genes by analysing their expression and the effects of mutations.
Adding Known Sequence to Unknown Sequence: Connecting identified genetic sequences with those whose function is not yet understood, facilitating further research.
Amplified Fragment Length Polymorphism (AFLP) Analysis for Genotyping: A method that combines selective PCR amplification of restriction fragments from a digest of the genomic DNA to analyse genetic diversity.
Next-Generation DNA Sequencing: Allows rapid sequencing of large stretches of DNA, providing high-throughput capabilities for genomic research.
Production of Recombinant Genes: Creating new genetic constructs that combine sequences from different sources.
Gene Cloning
Basic Concept
Involves taking a target DNA molecule (for example, a PCR product) and inserting it into a vector, which is a DNA molecule that can replicate within a host cell.
The resulting structure is known as a clone:
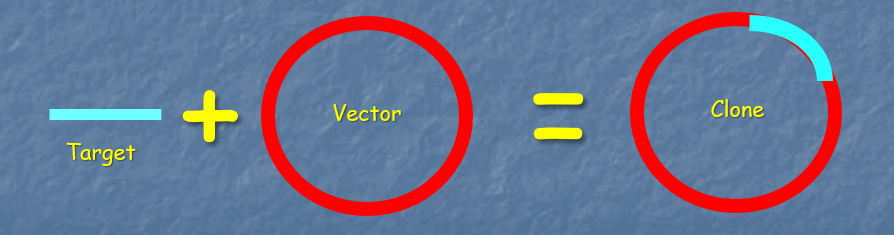

Mechanism of Cloning

Analysing Gene Function
Procedure
Obtain Coding Section of the Gene: This can be done by converting RNA to DNA through reverse transcription and PCR amplification of the desired gene.
Clone into an Expression Vector: Use a vector that contains its own gene promoter to ensure expression in host cells.
Evaluate Gene Function: Overexpress the gene to observe its effects. Alternatively, clone the target gene in reverse orientation to the promoter to create an antisense copy, which will block the function of the normal gene, allowing the study of gene silencing effects.
Producing Recombinant Genes
Fusion of Polymerase Activities
Example: Fusing the sequence encoding Taq polymerase's polymerase activity with the proofreading sequence of Vent™ polymerase.
Isolation by PCR: This step is followed by adding overhangs to the ends of the sequences and joining sticky ends.
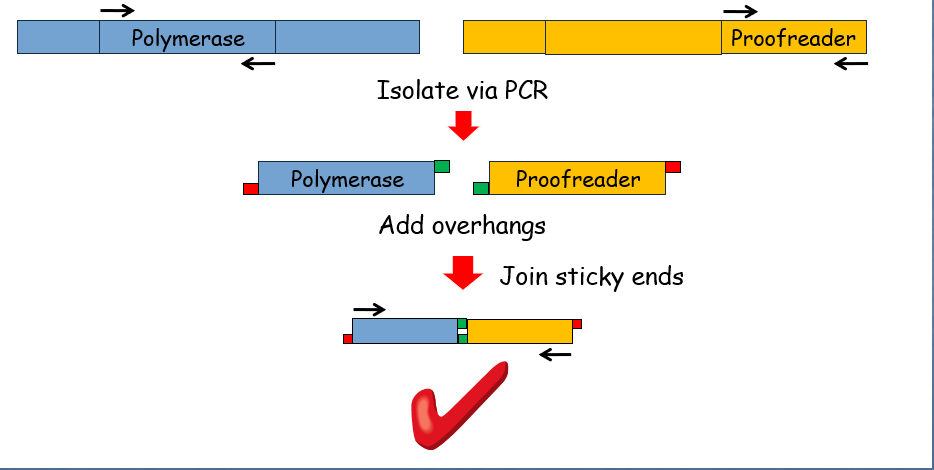
Avoiding Incorrect Fusions
Ensuring that the ends of each sequence have different sticky ends to prevent incorrect fusions during the joining process. The modification is facilitated by precise incorporation into PCR primers.
Connecting DNA Molecules
Requirement for Joining DNA
The fundamental requirement across all applications is the ability to efficiently join two pieces of DNA through a suitable enzyme, acting as a glue for DNA fragments.
DNA Ligase
Function
Definition: DNA ligase is an enzyme that binds two or more DNA fragments together. In nature, it repairs broken DNA in chromosomes.
Mechanism: Utilizes a source of ATP (adenosine triphosphate) to form new phosphodiester bonds between the ends of the DNA fragments, facilitating the joining of DNA strands.
Sources of DNA Ligase
T4 Bacteriophage DNA Ligase: A commonly used ligase in molecular biology.
E. coli DNA Ligase: Less commonly used because it relies on NAD (nicotinamide adenine dinucleotide) instead of ATP and cannot ligate blunt-ended fragments efficiently.
Challenges in Ligation Process
T4 ligase can theoretically ligate any DNA molecule:
Possibility of producing long chains of target DNA due to inefficient ligation controls.
Potential for multiple copies of the target DNA integrating into the vector, which may complicate experiments.
Risk of vector closure inhibits target splicing.
DNA Terminal Phosphorylation
Role of Phosphorylation
Requirement for Ligation: The 5’ ends of each DNA strand must be phosphorylated for DNA ligase activity.
Enzymatic Modification Options
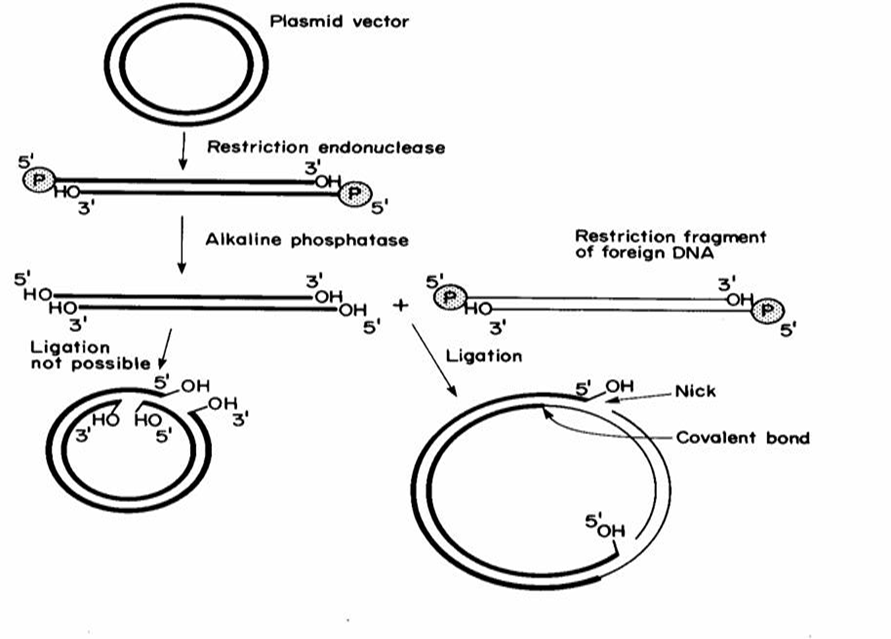
Alkaline Phosphatase: Enzyme that chemically modifies DNA by removing the 5’ phosphate group; a dephosphorylated vector cannot ligate to itself, ensuring only the target fragments can join.
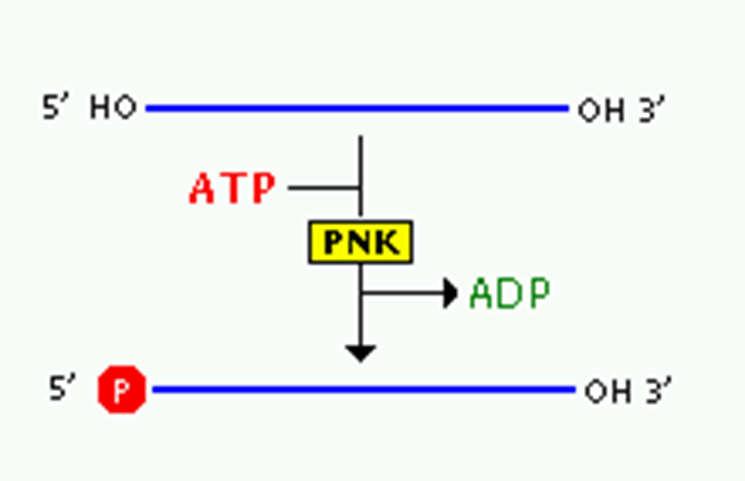
Polynucleotide Kinase: Adds phosphate groups to the 5’ end of DNA strands if not already present, ensuring compatibility for subsequent ligand reactions.
Klenow Fragment of E. coli DNA Polymerase I
Overview
This fragment is a larger component (76 kDa) that retains 5’ -> 3’ polymerase activity, as well as 3’ -> 5’ exonuclease activity but lacks 5’ -> 3’ exonuclease capability.
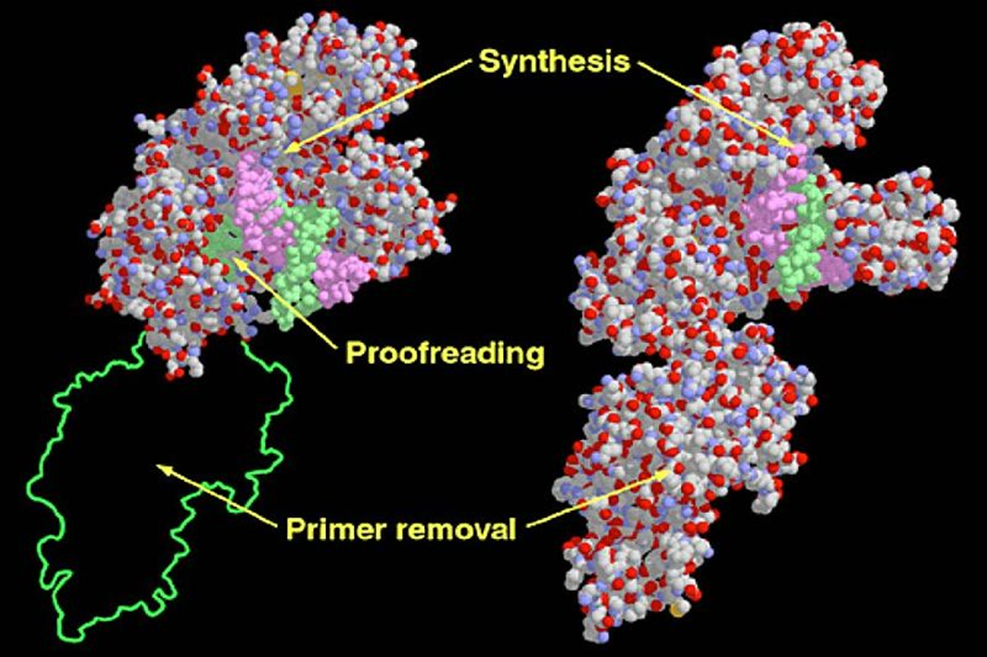
Applications of the Klenow Fragment
Synthesis of Double-Stranded DNA: Converts single-stranded templates into double-stranded DNA, previously used for creating labelled DNA probes.
Filling In Recessed Ends: Adding complementary bases to the recessed ends of DNA fragments to create blunt ends.
Digesting Protruding Overhangs: Removing any undesired protruding 3’ overhangs from DNA fragments.
Exonuclease-Minimized Klenow Fragment
Enhancing Blunt-End Joining
T4 ligase can join blunt-ended fragments, however, efficiency is generally lower compared to sticky ends. By treating blunt-ended DNA with a version of Klenow that lacks exonuclease activities, single-base overhangs can be added, improving compatibility for subsequent ligations with T4 ligase.
Reverse Transcriptase in Replication
Introduction of Concepts
Some viruses possess single-stranded RNA (ssRNA), which must be converted into DNA for replication within host cells.
Reverse Transcriptase: An enzyme that synthesizes DNA from RNA. Discovered in 1970 by researchers who were awarded the Nobel Prize for this work, highlighting that some organisms can indeed reverse the transcription process from RNA back to DNA.
Mechanism of Action
Reverse transcriptase operates similarly to a DNA polymerase and requires a short double-stranded RNA (dsRNA) duplex as a primer. In eukaryotic retroviruses, regions binding specific host tRNA enable duplex formation, which aids in synthesizing a complementary DNA copy in the 5’ -> 3’ direction.
The RNA component of this DNA-RNA duplex is decomposed by RNase H functionality of the reverse transcriptase afterward.
Applications of Reverse Transcriptase
Uses in Molecular Biology
Cloning Eukaryotic Genes: mRNA can be converted to DNA by reverse transcriptase for cloning into bacterial vectors. The introns must be spliced out since bacteria cannot perform splicing.
Gene Expression Analysis: Leverage known sequences in mRNA to prime reverse transcription, allowing the quantification of gene expression via measuring the resultant DNA amounts. For broad analysis, a poly-T primer matching the poly-A tails of eukaryotic mRNA can be used.
Next-Generation DNA Sequencing Techniques
Overview
Enables the production of short DNA sequences from millions of DNA fragments simultaneously, delivering massive amounts of sequence data, equivalent to about 200 human genomes in a single run.
Methodology: Shotgun Sequencing
The technique involves shearing DNA into small fragments and can also work effectively on cDNA.
DNA Fragment Treatment for Efficient Ligation
Steps for End Repair
Example provided with sequence treatment to address fragmented DNA with frayed ends before proceeding to blunt-end ligation:
Treat DNA with Klenow fragment to generate blunt-ended DNA fragments.
A-Overhangs Addition
Following blunt-end creation, the next step is to ligate an adaptor molecule. Due to the poor efficiency of blunt-ended fragment ligation, employing Klenow exonuclease along with dATP to create overhangs for compatible ligation.
Ligating Adaptors
The addition of adaptors is crucial for setting up amplification in subsequent amplification steps utilizing PCR techniques.
Other Modification Enzymes
Enzymatic Functions
DNA Methylases/Methyltransferases: Enzymes that add or remove methyl (CH₃) groups to DNA sequences, which can protect against or enable digestion by restriction enzymes, impacting gene expression and stability.
Topoisomerases: Similar to DNA ligase but instead can join DNA molecules with corresponding one-nucleotide overhangs. They can also be fused to vectors to facilitate complex manipulations, further enhancing DNA tandem joining capabilities.
Transposases: Enzymes that facilitate the movement of transposons, allowing segments of DNA to insert and excise themselves within the genome, providing an alternative to traditional DNA shear techniques for constructing libraries.
Transposons in Genome Libraries
Historically, shotgun genome libraries required random shearing of DNA, posing challenges due to inefficiency and expense. Transposons represent a solution, as they can replicate and jump between genomes, providing a more effective means of manipulating DNA without random destructive processes.
Use of Transposases in Applications
By pooling engineered transposons, transposases, and target DNA, integration occurs randomly, and afterward, transposases can remove integrated DNA segments, allowing targeted fragmentation for various experimental needs.