RECOMBINANT DNA TECHNOLOGY
involves the transfer of fragments of DNA from one organism or species to another.

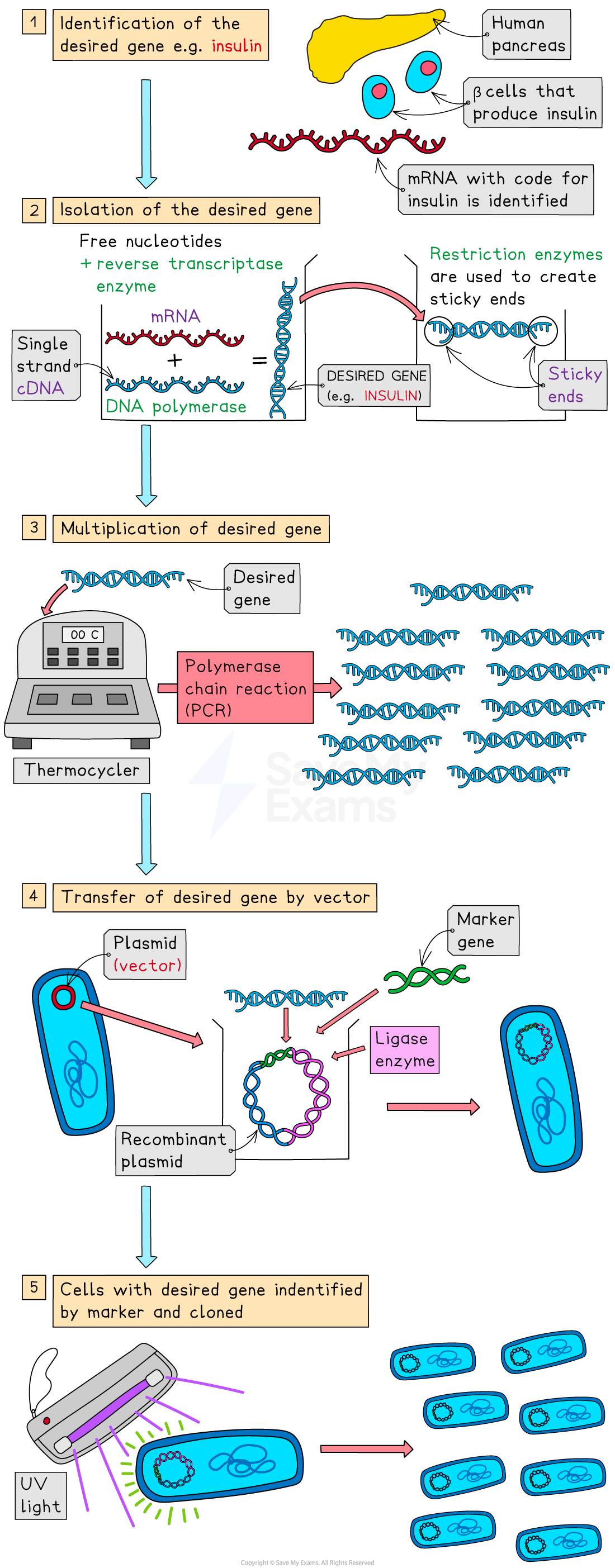
restriction endonucleases recognise specific palindrome sequences and will cut DNA at these places
gene machine
the sequence that is required is designed
the first nucleotide in the sequence is fixed to support
nucleotides are added step by step, in the correct order, in a cycle of processes that includes adding protecting groups.
IN VITRO AND IN VIVO CLONING
IN VIVO
insertion - DNA fragments is inserted into a host
transformation - host cell takes up a plasmid and the DNA
identification - shows which cells have been transformed
INSERTION -
uses a vector to transfer the DNA fragment into the host.
vector - plasmid found in bacteria
plasmid - circular DNA found in bacteria
step one - the vector is isolated
step two - the vector DNA is cut open using the same ‘restriction endonuclease’ that was used to isolate the DNA fragment containing the target gene
step three - this produces ‘sticky ends’ of the vector DNA which are complementary to the sticky ends of the DNA fragment containing the gene.
step four - the vector DNA and the DNA fragment are mixed together with the DNA ligase. DNA ligase joins the sticky ends together. this process is called ‘ligation’
IN VITRO
step 2 - annealing the primers
cooled to between 50 and 60c
hydrogen bonds are reformed
oligonucleotides bind to template DNA
oligonucleotides are called primers as enzymes in step 3 must attach to one of them before it can start to work
free DNA nucleotide are also added
step 3 - synthesis of new DNA