Tumor Antigens and Their Identification-class 4
Tumor Antigens: Specific vs Associated
Tumor-specific antigens (TSA), also known as tumor-specific transplantation antigens (TSTA) or tumor rejection antigens (TRA), are exclusively found on the surface of tumor cells. These antigens are not present on any non-tumor cells in the body, making them ideal targets for immunotherapy. TSAs can arise due to various factors, including:
Carcinogen Induction: Exposure to carcinogens can induce mutations leading to the expression of unique antigens.
Viral Infections: When an infecting virus transforms a cell into an immortal state, it may express viral antigens that act as TSAs. For instance, certain viral proteins expressed in tumors caused by viruses like HPV or EBV can serve as TSAs.
Idiotypes of BCR/TCR: In B cell lymphomas or T cell lymphomas, the idiotypes (unique variable regions) of B cell receptors (BCR) or T cell receptors (TCR) can function as TSAs. These idiotypes are specific to the clonal B or T cells of the lymphoma.
Tumor-associated antigens (TAA) are more commonly detected in cancer patients but are not exclusive to tumor cells. They are also found on normal cells under specific conditions:
Onco-fetal Antigens: TAAs can be expressed during fetal life and are normally silenced after birth. However, they may be re-expressed in tumor cells. Examples include carcinoembryonic antigen (CEA) and alpha-fetoprotein (AFP).
Selective Organ Expression: Some TAAs are expressed in specific organs after birth, and their expression may be elevated in tumors originating from these organs.
Low Concentration Expression: TAAs may be present in many cells but at significantly lower concentrations compared to tumor cells. This difference in expression levels can be exploited for targeted therapy.
Immune responses to TAA are often suppressed because the immune system recognizes them as "self" antigens. Overcoming this immune tolerance is a major challenge in cancer immunotherapy.
Desirable Characteristics of an Ideal Cancer Antigen
Universal Expression in Tumors: The antigen should be expressed in a majority of tumors to be effective across a wide patient population.
Tumor-Specific Expression: Expression should be limited to tumor cells to minimize the risk of autoimmunity and off-target effects.
Absent in Adults: The antigen should not be expressed in adults to avoid immune tolerance. Breaking immune tolerance is critical for TAA-targeted therapies.
Essential for Tumor Pathogenesis: The antigen should be crucial for tumor cell survival, ensuring that targeting it will lead to tumor cell death. Antigens derived from mutation drivers are preferred.
Strong Immune Response: The antigen should elicit a robust immune response, with epitopes that strongly stimulate the immune system and are distinct from non-mutated epitopes.
Recognition by T Cells: The antigen should have epitopes recognized by both and T cells to induce a comprehensive immune response. Recognition by lymphocytes is more effective than antibody-only recognition.
Tumor Antigens: Specific or Associated
Tumor antigens are categorized into two main types: Specific and Associated.
Specific (TSA): An antigen exclusively expressed in tumor cells and absent in healthy tissues. These are ideal targets due to their tumor specificity but are relatively rare.
Examples include:
Tumor Viral Antigens: Antigens derived from viruses that infect and transform cells (e.g., HPV E6 and E7 oncoproteins in cervical cancer).
Neoantigens/Mutated Antigens: Unique antigens resulting from somatic mutations in tumor cells, which are highly specific to the tumor.
Associated (TAA): Antigens expressed in tumor cells and also in certain normal tissues, often shared among different individuals and tumors.
These antigens are advantageous for identification and broad applicability but may pose challenges due to potential off-target effects.
Tumor Antigens: Shared or Unique
Unique Antigens: Antigens that appear only in a specific individual, often arising after exposure to carcinogens or UV radiation. These are highly personalized but challenging to target broadly.
Some Neoantigens/Mutated antigens can be considered unique due to their patient-specific nature.
Shared Antigens: Antigens common to multiple patients, facilitating the development of broadly applicable immunotherapies.
An antigen found in a single-digit percentage or less of the population is generally considered unique, while those found in several percentage points are considered shared.
For example, the rare isoform of CML is considered a shared antigen, and led to Asciminib being approved for its use.
Viral antigens, such as those from EBV or HPV, can be both specific (not normally found in the body) and shared (present in multiple individuals with viral-associated cancers).
Tumor Antigens as Target for Immunotherapy
Tumor-Specific Antigens (TSA):
Not expressed in normal tissues, offering excellent target specificity.
Examples:
Mutated Antigens (NeoAntigens, Ras, p53, bcr/abl): Result from mutations in genes like Ras, p53, or the bcr/abl fusion protein in leukemia.
Viral Antigens (HPV, EBV): Derived from oncogenic viruses like HPV (associated with cervical cancer) and EBV (associated with lymphoma).
Associated Antigens (TAA):
Expression can occur in immune-privileged sites or tissues with restricted expression.
Examples:
Cancer Testis Antigens (MAGE family, NY-ESO1): Normally expressed only in the testes and ovaries (immune-privileged sites) but can be re-expressed in tumors.
Differentiation Antigens (MART1, gp100, tyrosinase): Tissue-restricted expression, such as in melanocytes (MART1, gp100, tyrosinase).
Overexpressed Antigens in Tumor (WT1, Her2/neu, hTERT): Antigens overexpressed in tumors compared to normal tissues (WT1, Her2/neu, hTERT).
Target Quality:
Tumor-Specific Ags: Excellent targets due to their specificity.
Associated Ags: Target quality varies depending on tissue type and the extent of expression in normal tissue.
Potential Toxicity/Frequency: As tumor specificity goes up , the potential toxicity of the treatment decreases.
Identification of Tumor Antigens
A – Identification of tumor antigens:
Cloning of antigens identified by cellular reactivity: Isolating and cloning antigens based on their ability to elicit a cellular immune response.
SEREX (serological analysis of gene expression): Identifying antigens by screening tumor cell cDNA expression libraries with patient sera.
Next-generation/mass sequencing, DNA microarray technology: Using high-throughput sequencing and microarray technologies to identify tumor-specific gene expression patterns.
Methods 1 and 2 use cDNA libraries prepared from mRNA extracted from cancerous tissues/cells.
Identification of Tumor Epitopes
B – Identification of cancer MHC/epitopes (relevant to T cells):
If the antigen (Ag) is not known: Mass spectrometry (MS)/chromatography. Biochemical methods elute and fractionate TAA peptides naturally expressed on tumor cells in the context of HLA molecules by chromatography and mass spectrometry. This approach identifies the antigen itself.
If the antigen is known: Reverse immunology (epitope prediction on the basis of known HLA-binding motifs performed by dedicated software and sometimes supported by proteasome-cleavage programs).
Cloning of Antigens Identified by Cellular Reactivity
This process involves isolating T-cells specific for a given tumor without prior knowledge of the antigen.
How to identify the antigen?
Tumor antigens recognized by HLA class I restricted T cells are identified using expression cloning methods.
The appropriate HLA class I restriction element involved with presentation of the T cell epitope must be identified.
This can be done by:
Blocking T cell antigen recognition using antibodies that are specifically reactive with particular HLA alleles.
Assaying a series of tumor cell lines that either do or do not share particular HLA gene products with autologous tumor cells for their ability to stimulate the tumor reactive T cells.
A crucial condition is to have enough T-cells to perform the screens. Need about 2 million cells to identify one antigen!!
Identification of Immune Cellular Reactivity
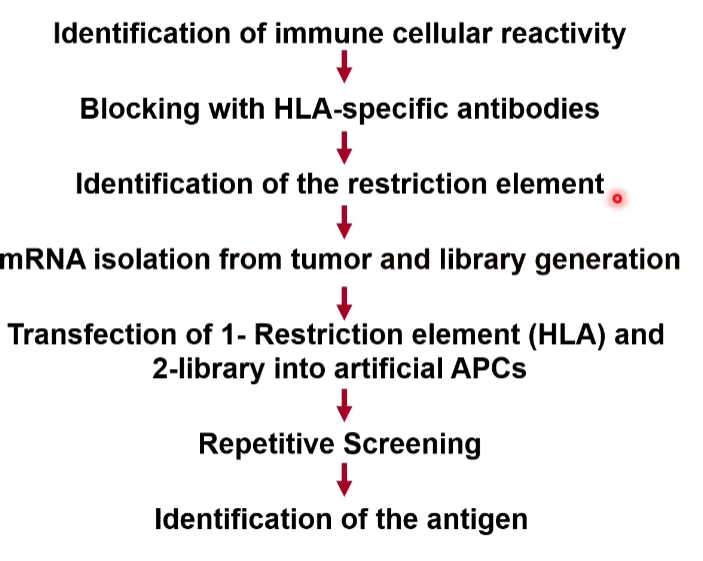
Blocking with HLA-specific antibodies to identify the restriction element.
mRNA isolation from tumor and library generation.
Transfection of 1- Restriction element (HLA) and 2-library into artificial APCs.
Repetitive Screening to isolate specific clones.
Identification of the antigen through sequencing.
SEREX
SEREX (serological analysis of gene expression) involves screening tumor cell cDNA expression libraries with patient sera to identify antigens.
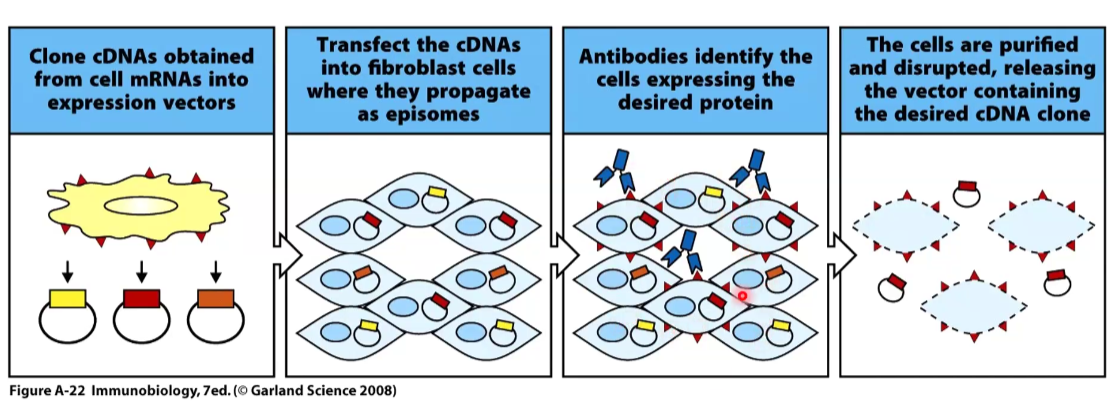
This is less used today because we have next gen sequencing.
This method has identified numerous autoantigens, many of which may not be recognized by tumor-reactive T cells.
However, a small subset of antigens identified via SEREX have been found to be recognized by HLA class I and class II restricted T cells.
High Throughput Approaches
Traditional methods like DNA microarray technology, which compare gene expression profiles in tumor tissues and normal counterparts, have been widely used.
Today, mass sequencing (next-generation sequencing - NGS) is the preferred method due to its higher throughput and sensitivity.
NGS techniques include:
Whole genome sequencing (WGS)
Whole exome sequencing (WES)
RNA-seq on tumors or single cells
Single-Cell Sequencing Strategy Example
scRNA-seq of tumor + adjacent normal tissue: Identifies tumor-specific overexpressed genes.
scDNA-seq or WES of same cells: Detects mutations that may serve as potential neoantigens.
scTCR-seq of tumor-infiltrating lymphocytes (TILs): Analyzes the T cell receptor repertoire to identify expanded or activated clonotypes.
Cross-reference TCR specificity: Matches TCR specificity (e.g., via tetramers or antigen screens) to predicted antigens to validate targets.
One of the examples shown is that you get a line of MSA sequencing to show the differentiation of mutations across different samples, which can indicate how neoantigens vary between tumor subtypes. Usually is is on the level of residues.
Identification of Antigens Via Epitope
T-Cells and MHC Molecules
T-cells recognize small peptides presented by MHC molecules on the surface of cancer cells called epitopes.
This interaction is critical for T-cell activation and effector function.
Determining the sequence of these epitopes is essential for developing targeted immunotherapies.
MS/Biochemical Methods (if the Antigen is Not Known)
This approach involves eluting peptides from MHC class I molecules and fractionating them using reversed-phase HPLC columns.
The fractions are then tested for their ability to sensitize target cells for T cell recognition.
Additional fractionation steps are performed using various columns or buffers to refine the peptide separation.
Peptides associated with T cell recognition are identified based on their elution profile.
The sequence of candidate peptides is determined using tandem mass spectrometry (MS), liquid chromatography (LC), and synthetic peptides are generated and tested for T cell recognition.
MS/Biochemical Methods
Isolation of MHC molecules from cancer cells.
Gentle elution of the peptides to preserve their integrity.
Comparison to peptides isolated from normal cells from the same tissue to identify tumor-specific peptides.
Sequencing of the peptides using Mass Spectrometry to determine their amino acid sequence.
Stimulation of T-cells with synthetic peptides to validate their immunogenicity.
Identification of the antigen and checking its expression in different tumor cells.
Reverse Immunology (if the Ag is Known)
Reverse immunology identifies candidate epitopes from proteins over-expressed in tumors using HLA binding motifs.
The motifs represent consensus sequences derived from:
Direct measurement of MHC-peptide binding affinities.
Identifying common residues in known T cell epitopes.
Analyzing peptides eluted from individual MHC molecules.