Protein synthesis
Transcription
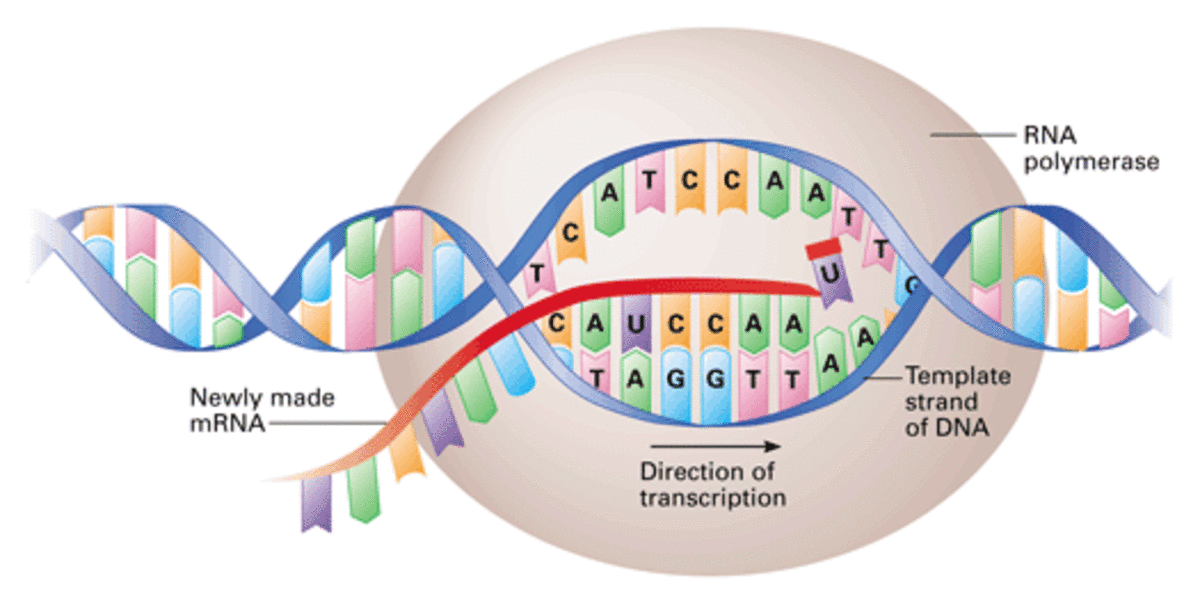
Steps in transcription:
RNA polymerase binds to the promoter at the start of the gene, helped by transcription factors.
A helicase unzips the 2 DNA strands by breaking the hydrogen bonds between the bases.
Activated RNA nucleotides bind via hydrogen bonds to the exposed bases on the template stands forming complementary base pairs.
RNA polymerase catalyses the condensation reaction that joins the incoming RNA nucleotide to the growing RNA chain, forming a phosphodiester bond.
Two phosphates are released as pyrophosphate and hydrolysed to provide energy for the reaction.
RNA polymerase stops transcription when it reaches a termination signal.
mRNA is released from the template strand and leaves the nucleus through a nuclear pore.
Copying sections of DNA into RNA before translation is advantageous as:
mRNA is a small molecule that can leave the nucleus via a nuclear pore.
Many copies of mRNA can be made and then translated to synthesise many polypeptides.
There are more steps that allows for more regulation.
Translation
Translation is split into 3 stages: initiation, elongation, and termination
Initiation: Initiator tRNA carrying methionine binds to AUG start codon and ribosome assembles.
Elongation: The next tRNA, carrying a specific activated amino acid, binds via its anticodon to the next exposed mRNA codon, forming hydrogen bonds between complementary bases. Ribosome catalyses the condensation reaction that joins adjacent amino acids by a peptide bond. Ribosome moves one codon along mRNA and empty tRNA is ejected.
Termination: Translation continues until a stop codon is reached. Since there is no corresponding tRNA, a protein release factor binds instead. The polypeptide is released and the ribosome dissociates.
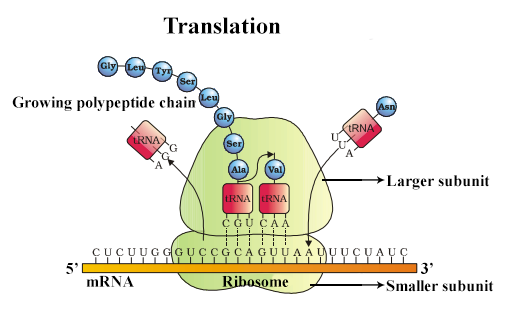
ATP is needed to attach a specific amino acid to its corresponding tRNA with the aid of an enzyme. When the bond between the tRNA and amino acid is broken, this provides the energy for peptide bond formation.
Role of RNA in protein synthesis
Messenger RNA (mRNA) | Single stranded RNA copy of a gene synthesised in the nucleus. Leaves nucleus through a nuclear pore and assembles with a ribosome in the cytoplasm. Its length can vary depending on how long the gene is. Sequence of bases determines the sequence of amino acids in the polypeptide. Short lifespan before broken down. |
Transfer RNA (tRNA) | Small, L-shaped molecule that functions as an adaptor to bring activated amino acids to the ribosome and align them in the correct order along the mRNA. Activated amino acids attach at one end. Three bases (anticodon) at other end binds to the next mRNA codon exposed in ribosome, by complementary base pairing. |
Ribosomal RNA (rRNA) | Transcribed in the nucleolus and assembles with proteins to form a ribosome. Catalyses peptide bond formation. |
Folding and modification of polypeptide chains
Polypeptides that are released from the ribosomes must be correctly folded and modified to form the functional protein e.g. enzyme. Chaperone proteins help the polypeptide to correctly fold into its final specific 3D shape (tertiary structure), which is held in place by bonds between the amino acid R groups including hydrogen bonds, ionic bonds and covalent disulfide bridges. All genes start with the initiation codon AUG that codes for methionine. This first methionine is removed by the enzyme methionine amino peptidase as the polypeptide is translated. Any necessary prosthetic groups are later added, and the polypeptide chains link to form a quaternary structure in the Golgi body.