21.1 DNA profiling
Every person has a unique combination of DNA in the chromosomes, unless they are identically related.
The human genome
Genome - all the genetic material an organism contains.
For eukaryotes, DNA in the nucleus and the mitochondria combines. Chromosome are made up of hundred of millions of DNA base pairs, but your genes, the 20-25000 regions of DNA that code for proteins, only make up about 2% of your total DNA → they are called exons
Large non-coding regions of DNA that are removed from messenger (m)RNA before it is translated into polypeptide chain → are introns.
Within introns, telomeres, and centromeres there are short sequences of DNA that are repeated several times - satellite DNA.
In region of minisatellite, a sequence of 20-50 base pairs will be repeated from 50 to several hundred times. These occur at more than 1000 locations in the human genome and are known as variable number tandem repeated (VNTRs)
Microsatellite - smaller region of just 2-4 bases repeated only 5-15 times. They are known as short tandem repeats (STRs). These satellites always appear in the same positions on the chromosomes, but the number of repeats of each mini or microsatellite caries between individuals as different lengths of repeats are inherited from both parents.
Only identical twins will have an identical satellite pattern, but the more closely related you are to someone, the more likely you are to have similar patterns.
Patterns in non-coding DNA were discovered by Professor Sir Alec Jeffreys and team at Leicester University in 1984.
DNA profiling / Genetic fingerprinting - producing an image of the patterns in the DNA of an individual, can assist in identification of individuals or familiar relationships.
Producing a DNA profile - 5 main stages
1 - Extracting the DNA
DNA must be extracted from a tissue sample. When it was first discovered relatively large samples were needed however now using a technique called polymerase chain reaction (PCR), the tiniest fragment of tissues can give scientistic enough DNA to develop a profile.
2- Digesting the sample
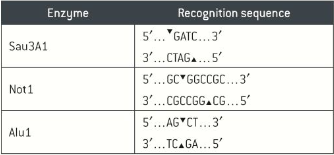
Strands of DNA are cut into small fragments usingi special enzymes called restriction endonucleases. Different restriction endonucleases cut DNA at specific nucleotide sequence, known as restriction site or recognition site. All restriction endonucleases make two cuts, once though each strand of the DNA double helix.
Restriction endonucleases give scientists the ability to cut the DNA strands at defined points in the introns, they use a mixture that leave the repeating units or satellites intact, so the fragments at the end of the process include a mixture of intact mini and microsatellites regions.
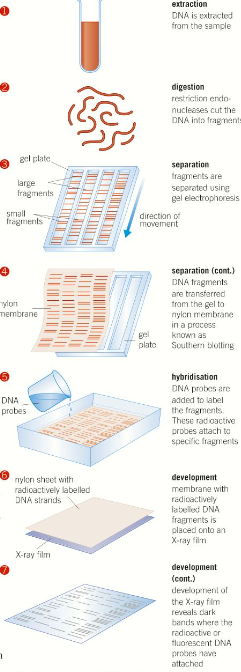
 3 - Separating the DNA fragments
3 - Separating the DNA fragments
The cut fragments of DNA need to be separated to form a clear and recognisable pattern, this is done using electrophoresis, which is a technique that utilises the way charged particles move through a gel medium under the influence of an electric current. The gel is then immersed in alkali in order to separate the DNA double strands into single strands. The single-stranded DNA fragments are then transferred onto a membrane by Southern Blotting.
4 - Hybridisation
Radioactive or fluorescent DNA probes are now added in excess to the DNA fragments on the membrane. DNA probes are short DNA or RNA sequences complementary to a known DNA sequence. They bind to the complementary strands of DNA under particular conditions of pH and temperature. DNA probes identify the microsatellite regions that are more varied than the larger minisatellite regions, the excess probes are washed off.
5 - Seeing the evidence
If radioactive labels were added to the DNA probes, X-ray images are taken of the paper/membrane. If fluorescent labels were added to the DNA probes, the paper/membrane is placed under UV light so the tags glow. → This method is most commonly used today. The fragments give a pattern of bars - the DNA profile - which is unique to every individual except identical siblings.
Separation of nucleic acid fragments by electrophoresis
DNA fragments are put into well in agarose gel strips, which also contain a buffering solution to maintain a constant pH. In one of more wells, DNA fragments of known length are used to provide a reference for fragment sizing.
When an electric current is passed through the electrophoresis plate, the DNA fragments in the wells at the cathode move through the gel towards the positive anode at the other end. This is due to the negatively charged phosphate groups in the DNA fragments. The rate of movement depends on the mass or length of the DNA fragments - the gel has a mesh-like structure that resists the movement of molecules. Smaller fragments can move through the gel mesh more easily than larger fragments causing them to move further over a period of time.
When the fastest smallest fragments reach the anode end of the gel, the electric current is switched off.
The gel is then placed in alkaline buffer solution to denature the DNA fragments, the two DNA strands of each fragment separate, exposing the bases.
In a technique called Southern Blotting, these strands are transferred to a nitrocellulose paper or a nylon membrane, which is placed over the gel. The membrane is covered with several sheets of dry absorbent paper, drawing the alkaline solution containing the DNA through the membrane by capillary action.
The single-stranded fragments of DNA are transferred to the membrane, as they are unable to pass through it. They are transferred in precisely the same relative position as they had on the gel, they are then fixed into place using UV light of heated at 80’c.
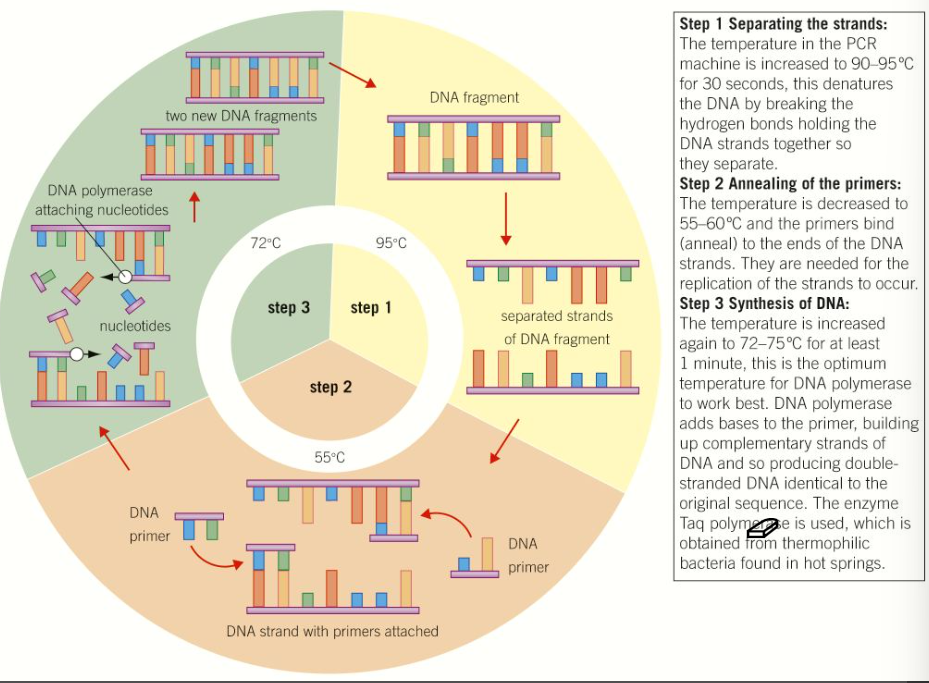
Polymerase Chain Reaction (PCR)
DNA profiling is often used in solving crimes and only very tiny amounts of DNA may be available, the PCR is a version of the natural process by which DNA is replicated and allows scientists to produce a lot of DNA from a tiny sample.
An excess of 4 nucleotide bases A,T,C and G (in form of deoxynucleoside triphosphates), small primer DNA sequences, and enzyme DNA polymerase are placed in a vial that is placed in PCR machine that is carefully controlled and changes rapidly at programmed intervals, triggering different stages of the process.
Can be repeated many times by the PCR machine, about 30 repeats will give 1 billion copies of the original sample.
Uses of DNA profiling
Forensic science, especially criminal investigations, where DNA can be obtained from blood, semen, saliva, hair roots, and skin cells.
DNA profile can then be compared to a sample taken of the suspect or using criminal DNA database.
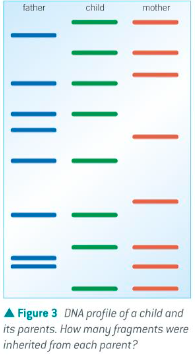
Prove paternity of a child
Immigration cases to prove or disprove family relationships
Identifying the species which an organism belongs to
Demonstrate the evolutionary relationships between different species.
Identifying individuals who are risk of developing particular diseases.
Certain non-coding microsatellites, or repeating patterns that they make have been found to be associated with an increased risk/ incidence of particular diseases, including various cancers and heart diseases.
Specific genes markers can be identified and observed in DNA profiles.
Pitfalls of profiling
May be mistaken DNA identity which occurred in 2000. Raymond Easton was in advanced stage of Parkinson’s disease and was arrested for a burglary that happened over 200 miles away, based solely on DNA evidence.