What Is The Formula For Calculating The Tm In Pcr
Page 4:
Biotechnology company discovered a 13-amino acid polypeptide that stimulates milk production in cows
DNA sequence with start and stop codons provided
Primers needed to amplify the 42-base pair region
Designate the 5' and 3' ends of the primers
Page 5:
Sequences of the two 20-nucleotide primers provided
Primers written in 5' to 3' direction
Tm (melting temperature) calculated for each primer
The formula for calculating the melting temperature (Tm) in PCR is typically estimated using the Wallace Rule of Thumb:
Tm = 4(G + C) + 2(A + T)
Sequences of the two 20-nucleotide primers provided
Tm (melting temperature) calculated for each primer
The criteria for primers in PCR (Polymerase Chain Reaction) are as follows:
Length: Typically 18-30 nucleotides long.
GC Content: Ideally 40-60% to ensure stable binding.
Melting Temperature (Tm): Similar Tm values for both primers to promote annealing.
Specificity: Primers should be specific to the target DNA sequence to avoid non-specific amplification.
Absence of Self-Complementarity: Primers should not have significant self-complementarity to prevent hairpin formation.
Absence of Primer-Dimer Formation: Primers should not have significant complementarity to each other to avoid primer-dimer formation.
Avoidance of Repeats: Primers should not contain repetitive sequences to prevent non-specific amplification.
3' End Stability: Primers should have stable 3' ends to promote efficient extension by DNA polymerase.
Avoidance of G/C Clamp: Primers should not end with G or C to prevent non-specific binding.
Avoidance of Secondary Structures: Primers should not have significant secondary structures to ensure efficient amplification.
Page 7
Sequences of the primers needed for directional sub-cloning provided
HindIII and EcoRI restriction enzyme sites used
Additional nucleotides added to the 5' ends of the primers for improved digestion with enzymes
Page 8
Amplified DNA segment after digestion with HindIII and EcoRI
Top strand: 5’-AGCTTATGGGTACTTGTGAGAAGAGCACTGGGTTCACTGCGTGTTAAG-3’
Bottom strand: 3’-ATACCCATGAACACTCTTCTCGTGACCCAAGTGACGCACAATTCTTAA-5’
5’ and 3’ ends designated
Review begins here
Page 9
Definition of a gene
Gene as a unit of heredity
Gene in the context of functional and structural aspects
Components of a gene
Enhancers, promoter, terminator, open reading frame
Genes can encode different products through alternative splicing (from a single locus)
RNA processing is required for functional RNA production

Page 10
Properties of genetic material
Replication for accurate transmission of genetic information
Gene expression to control phenotype
Mutability to allow for variation
Anlage = Gene (Factors)
Page 11
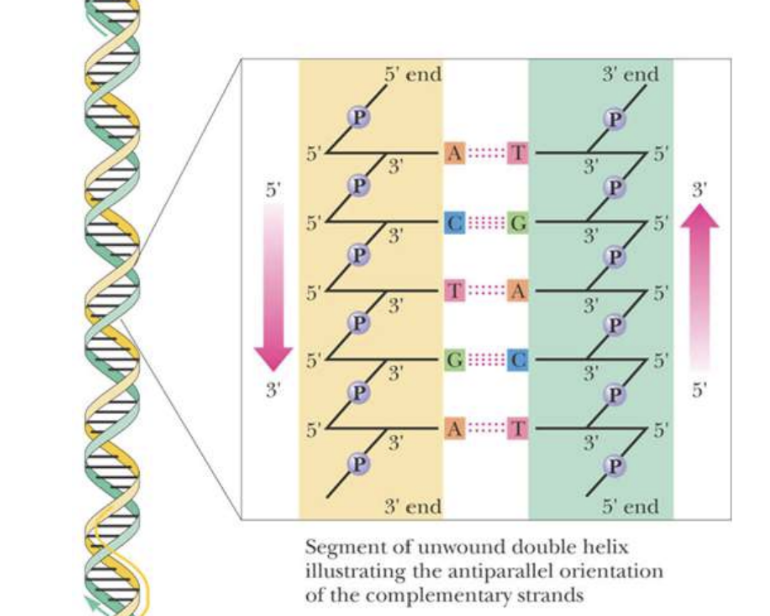
Structure of DNA double helix
Anti-parallel orientation of complementary strands
Strands held together by interchain H-bonds
Complementary base pairing leads to specific association of the two chains
Page 12
Nucleic acid hybridization
Based on base-pair complementarity
Can be DNA-DNA, RNA-RNA, or DNA-RNA duplexes
Various techniques utilize hybridization
Page 13
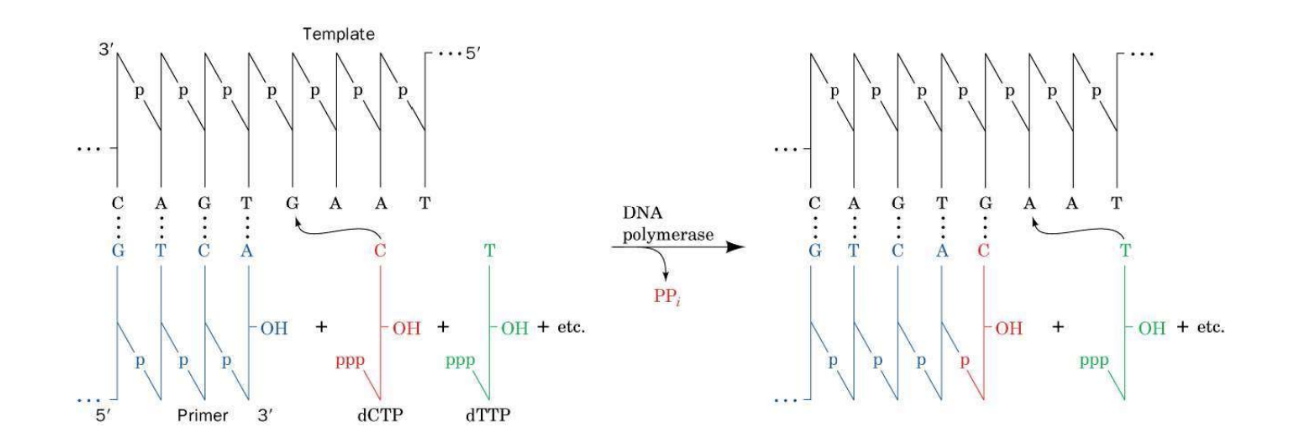
DNA synthesis by DNA polymerases
Replication occurs in a 5’ to 3’ direction
Deoxyribonucleotides added to the 3’ end of a growing chain
Template strand used for synthesis
Page 14
Polymerase Chain Reaction (PCR)
Amplification of DNA sequence in vitro
Ingredients: DNA template, primers (oligonucleotides), deoxyribonucleotides, DNA polymerase (Thermostable, Taq Polymerase), buffer
Taq Polymerase is a commonly used thermostable polymerase, was discovered in organisms which lived in extremely harsh conditions
Page 15
Steps of PCR in a thermocycler
Denature DNA strands at high temperature (92-95˚C)
Allow primers to anneal at lower temperature, lower temp (50-60˚C)
Use thermostable DNA polymerase to replicate DNA at optimal temperature (72˚C)
Cycle of steps repeated multiple times, typically 30 times
Page 16
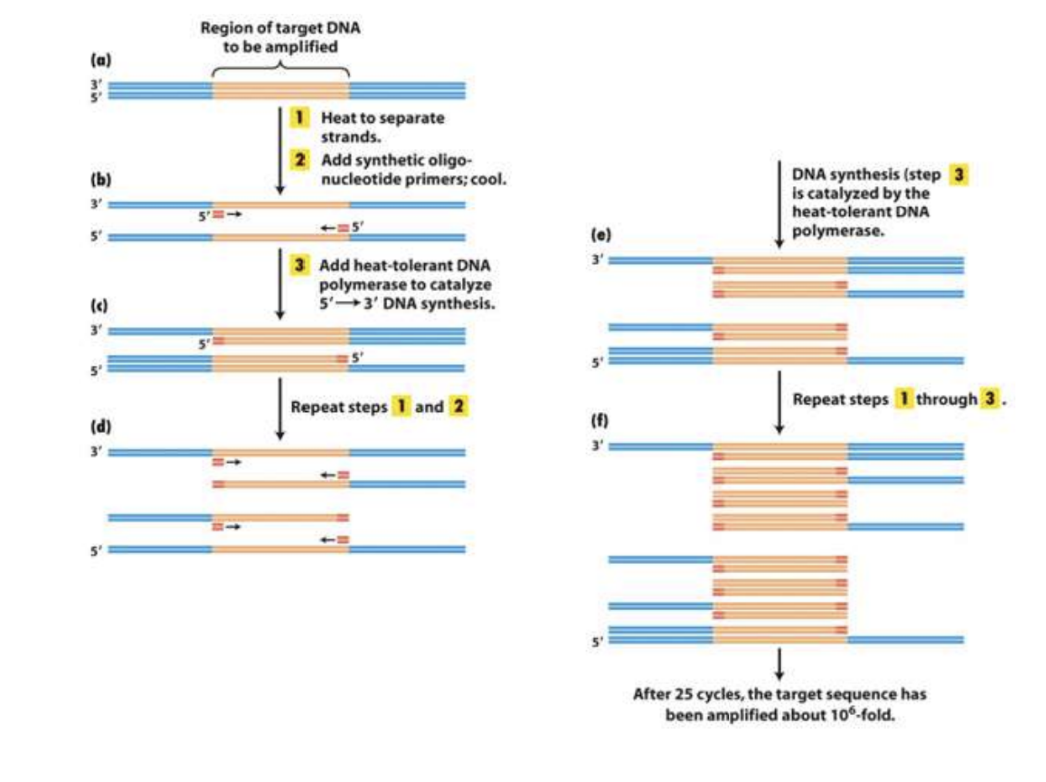
PCR cycles and DNA amplification
Illustration of the first few cycles of PCR
Heat denaturation, primer annealing, DNA synthesis
Amplification of target sequence after multiple cycles
Page 17
Primer design for PCR
Properties of primers: length (18-24 bp), G/C content (40-60%), melting temperature (Tm) (50-65˚C, anneling temp will be 3-6˚C lower)
3’-end critical for specificity
Considerations for primer pairs, Tm withing 5˚C of eachother, they should not have complementary pairs (primer dimer)
Page 18
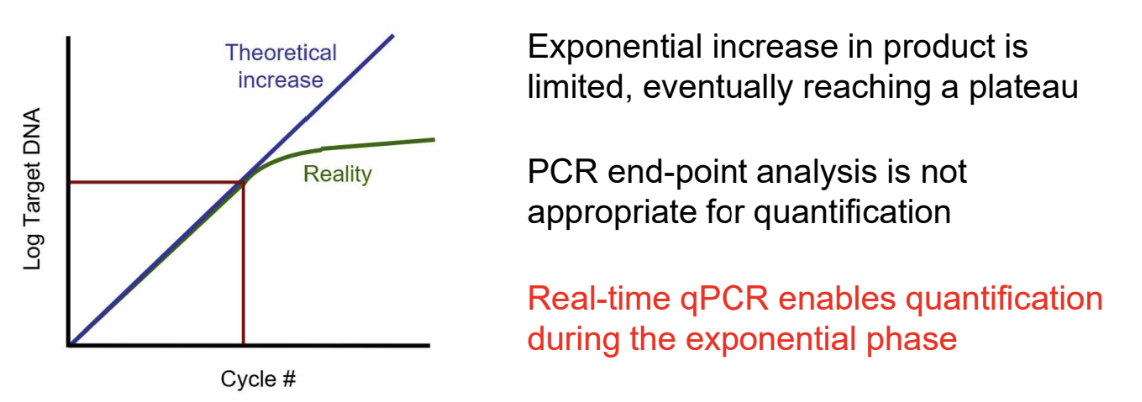
Real-Time Quantitative PCR (qPCR)
Fluorescence-based detection of amplification products in real time
Measures input quantity of nucleic acid during exponential phase
Contrasted with traditional PCR using endpoint analysis
Page 19
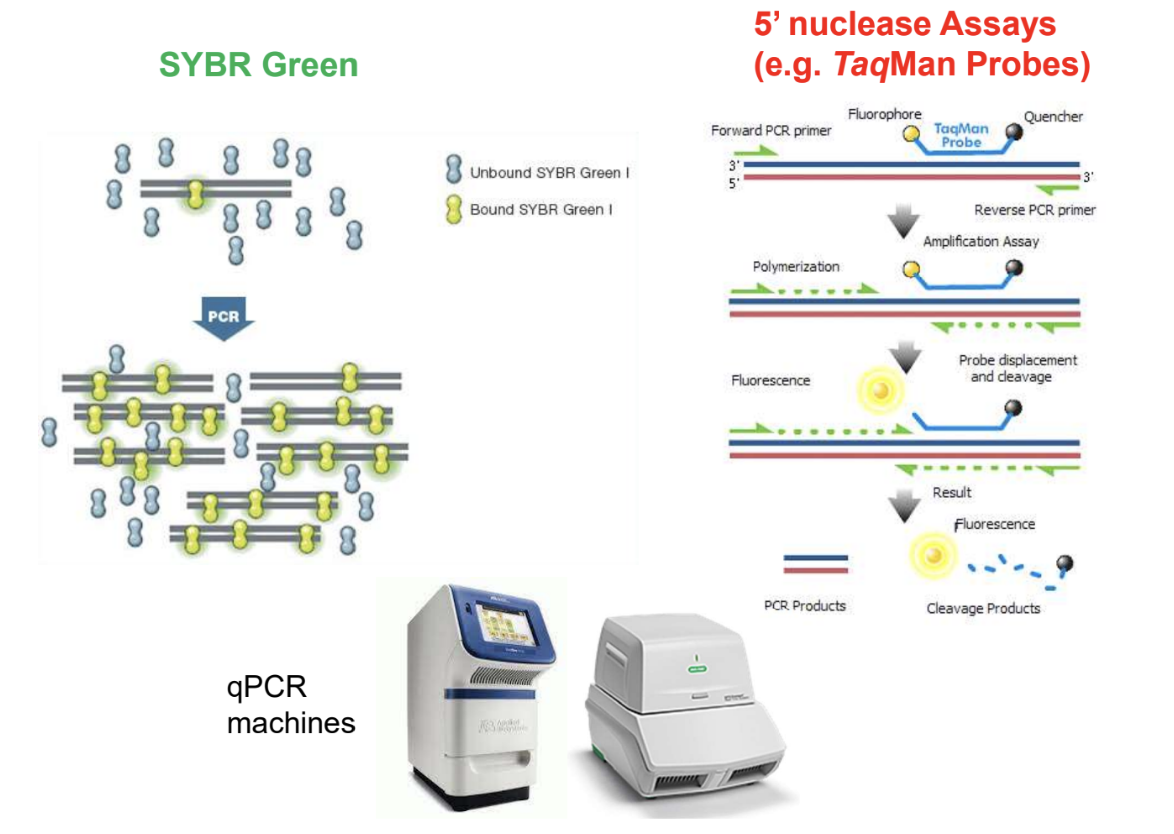
qPCR fluorescence detection methods
5' nuclease assays and SYBR Green
TaqMan Probes and PCR primers
Fluorescence and cleavage for detection
Page 20
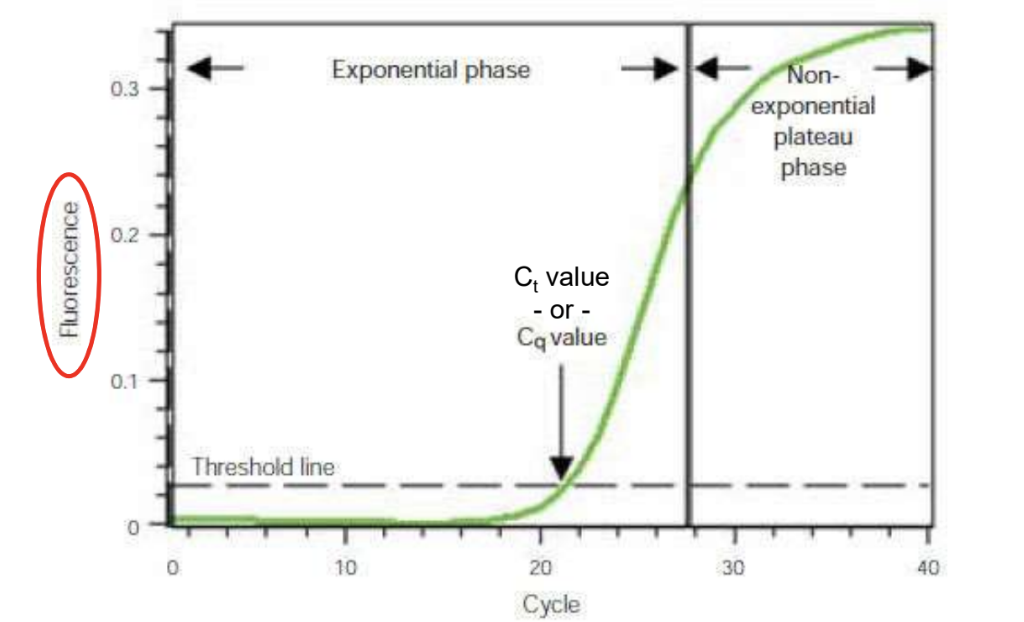
Ct value in qPCR
Ct value determined by the amount of template present
High starting template gives low Ct value, small amount gives high Ct value
Page 21
Chromatin and chromosomes
Chromatin as DNA-protein complex in the nucleus
Chromosomes as separate pieces of chromatin during cell division
Composition of chromatin: DNA, histones, non-histone proteins, RNA
DNA interaction with proteins allows compaction into the nucleus
Page 22
Histones in chromatin
Histones as major protein component of chromatin
Five types of histones: H1, H2A, H2B, H3, H4
Positively charged amino acids in histones
Conservation of H2-H4 proteins across species (cows & grass) - diverged 1.3 billion YAG
Post-translational modifications of histones (methylation, acetylation and phosphorylation)
Page 23:
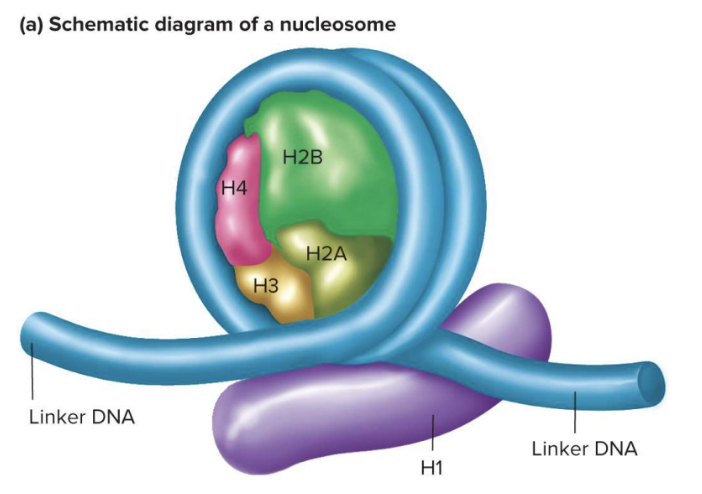
Nucleosome Core:
Octamer of two each of histones H2A, H2B, H3, and H4
~160 bp of DNA wrap twice around core of eight histones
Positive charges at the N-termini of histones attract the negative charges of DNA phosphates
40 bp of linker DNA connects adjacent nucleosomes
Histone H1 associates with linker DNA as it enters and leaves the nucleosome core
Page 24:
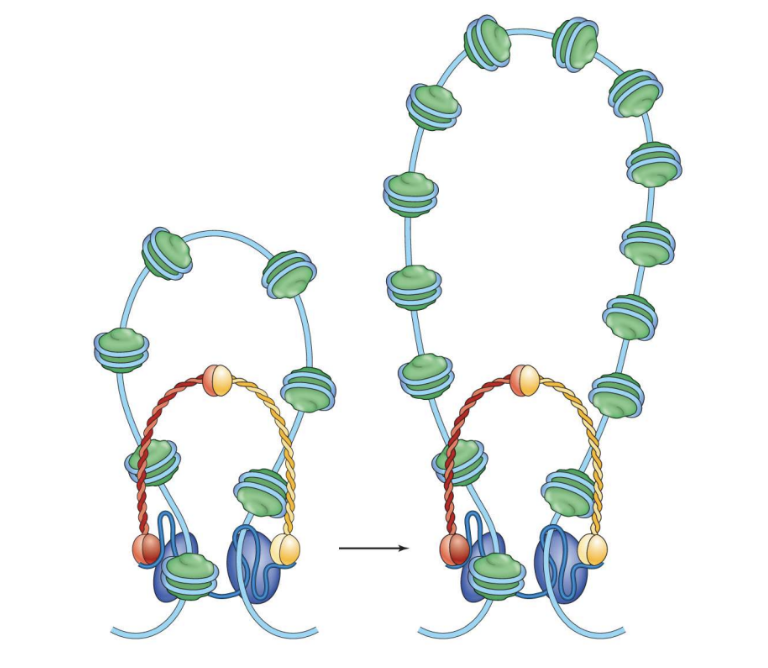
Radial Loop-Scaffold Model for Higher Levels of Compaction:
Loops of DNA formed by protein complexes called condensins
Five protein subunits constitute condensin
are rings; form arounf nucleosome studded DNA, chromosomes passes through the ring twice to form loop
Loops can contain 30 – 400 kbp of DNA
Page 25:
Summary:
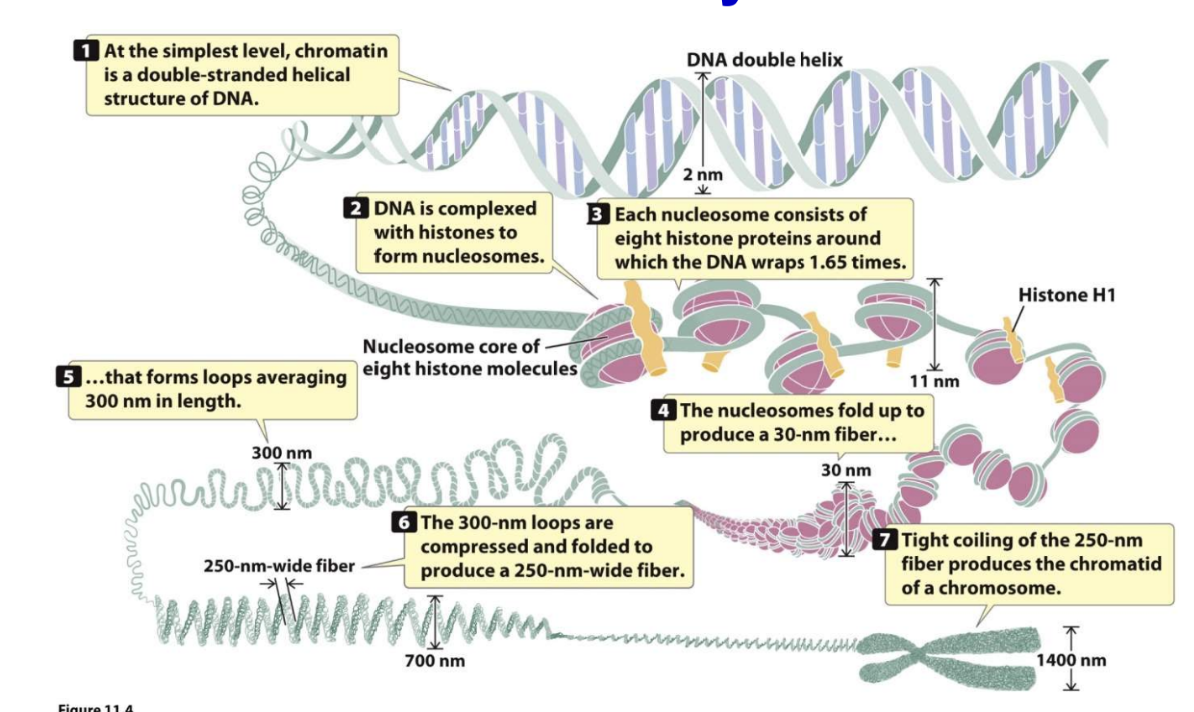
Chromatin DNA double helix is a double-stranded helical structure of DNA
DNA is complexed with histones to form nucleosomes
Each nucleosome consists of eight histone proteins around which the DNA wraps 1.65 times
Histone H1 associates with nucleosome core
Nucleosomes fold up to produce a 30-nm fiber
300-nm loops are compressed and folded to produce a 250-nm-wide fiber
Tight coiling of the 250-nm-wide fiber produces the chromatid of a chromosome
Page 26:
Chromosomal Packaging and Function:
Heterochromatin is highly condensed and usually transcriptionally inactive
Darkly stained regions of chromosomes
Constitutive heterochromatin is condensed in all cells
Facultative heterochromatin is condensed in only some cells and relaxed in other cells
Euchromatin is relaxed and usually transcriptionally active
Lightly stained regions of chromosomes
Page 27:
Fluorescence In Situ Hybridization (FISH):
Hybridization between metaphase chromosomes and a labeled DNA sequence
Probe could also be complementary RNA molecule
FISH involves denaturation of chromosomes
Page 28:
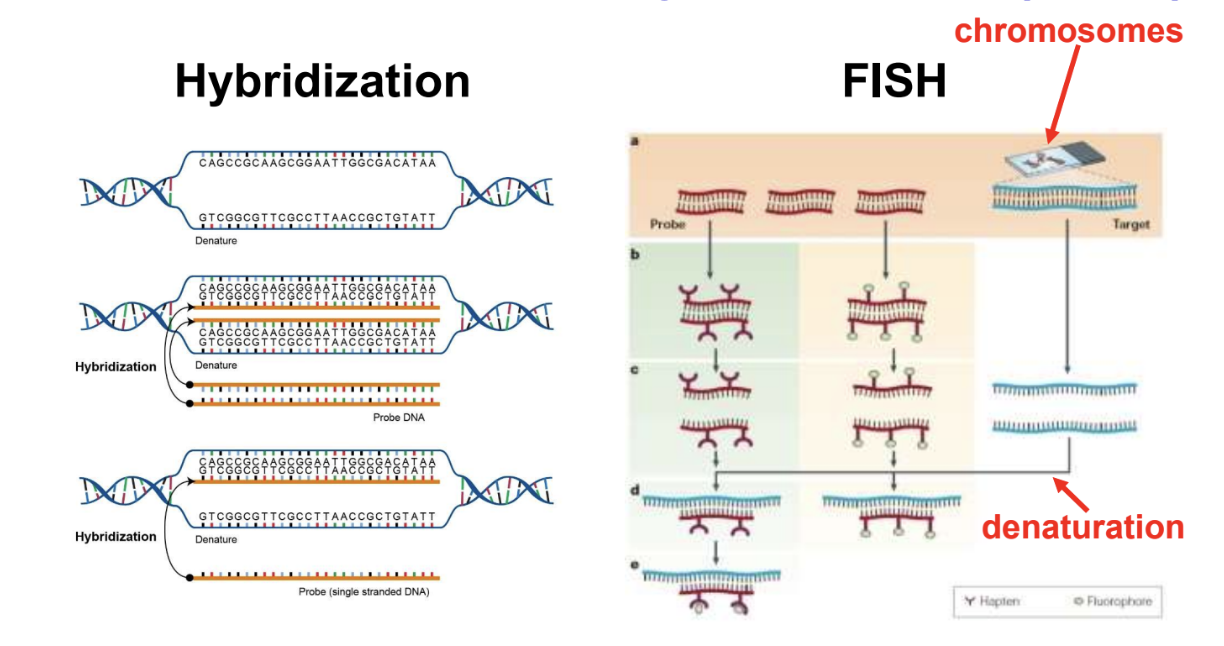
Fluorescent In Situ Hybridization (FISH) is Used to Characterize Genomes as Visualized on Chromosomes:
Depends on hybridization between metaphase chromosomes and a labeled DNA sequence
Chromosomes are spread on a glass slide and denatured to make them single stranded
A DNA sequence is labeled with a fluorescent tag to make a probe
The probe hybridizes to chromosomes at complementary regions
Page 29:
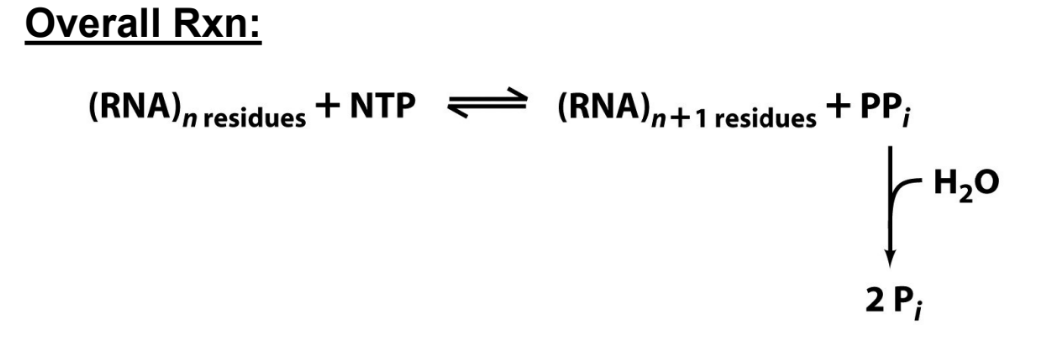
DNA-dependent RNA Polymerases (RNAPs):
Synthesize RNA from DNA templates (transcription)
DNA template determines which base is added (complementary base pairing)
Synthesis proceeds in a 5’ to 3’ direction
Targeted to specific genomic DNA sequences (genes)
No primer is needed to initiate RNA synthesis
Page 30:
Some Essential Terminology:
Nontemplate (RNA-like) strand and template strand
RNA synthesis is complementary and antiparallel to the template strand
New nucleotides are added to the 3'-OH group of the template strand
Transcription proceeds in a 5' to 3' direction
Page 31:
Overview of Transcription:
Transcription involves the unwinding and rewinding of DNA
RNA polymerase synthesizes RNA from the template strand of the gene
Transcription proceeds in a 5' to 3' direction
Page 32:
Generalized Components of a Gene Encoding a Protein:
Promoter, mRNA, gene stop, 5’-UTR, 3’-UTR, terminator, open reading frame
Enhancers are DNA sequences that influence the level of transcription and are often far away from the promoter sequence
Page 33:
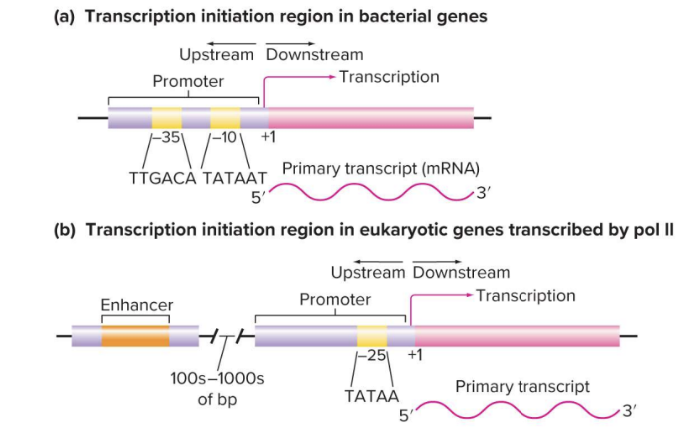
Transcriptional Initiation Varies Between Prokaryotes and Eukaryotes:
Eukaryotic genes often have enhancers that can be far away from the promoter and are required for efficient transcription
Page 34:
Multiple Eukaryotic RNA Polymerases:
Three distinct RNA polymerases in nuclei of eukaryotes: RNAP I, RNAP II, RNAP III
RNAP I synthesizes precursors of most rRNA
RNAP II synthesizes mRNA precursors, snoRNAs, some miRNAs, and some snRNAs
RNAP III synthesizes small RNAs such as 5S rRNA, tRNAs, some miRNAs, and some snRNAs
Page 35:
Eukaryotic Promoters:
Eukaryotic RNAPs require General Transcription Factors (GTFs) to recognize promoters and recruit RNAP to the transcription start site
Eukaryotic promoters are more complex and diverse than prokaryotic promoters
Three core eukaryotic RNAPs recognize different types of promoters
Page 36:
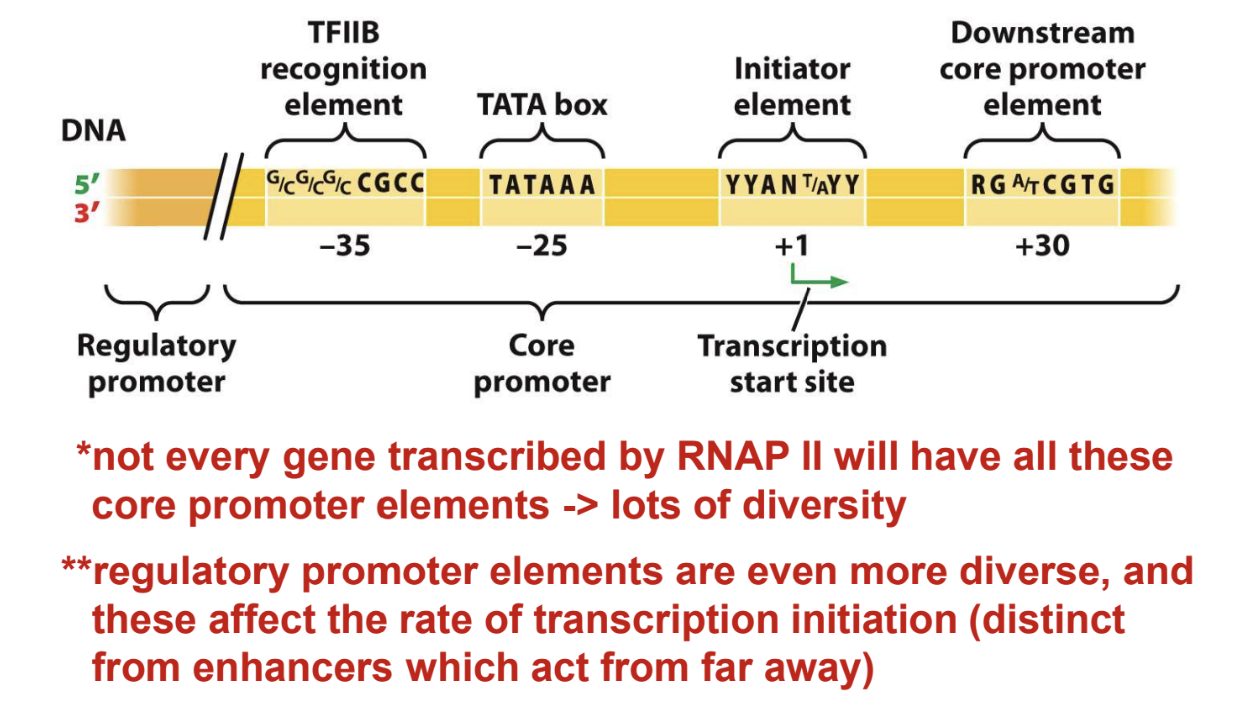
RNA Polymerase II Promoters:
Promoters are complex and diverse
Core promoter elements and regulatory promoter elements affect the rate of transcription initiation
Page 37:
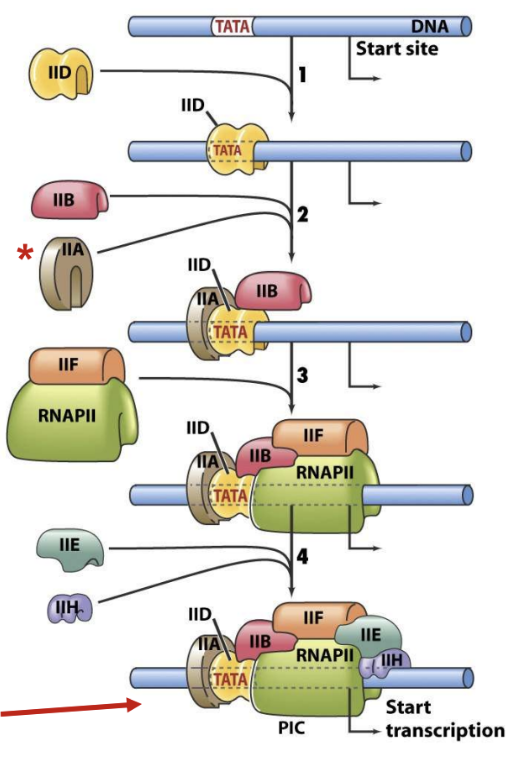
Assembly of GTFs and Transcription Initiation:
TFIIH induces open complex formation for RNA synthesis to begin
TBP binds TATA box
TFIIB determines start site
Preinitiation complex is formed
Page 38:
TATA-binding protein (TBP):
TBP recognizes TATA-box and induces a large bend in DNA
TBP is a universal eukaryotic transcription factor required for initiation by RNAP I, RNAP II, and RNAP III
SL1 = TBP + TAFs for RNAP I
TFIID = TBP + TAFs for RNAP II
TFIIIB = TBP + TAFs for RNAP III
Page 39:
Elongation and Termination of RNAP II Transcription:
After initiation, RNAP II shifts to elongation mode
Phosphorylation of CTD of Rpb1 subunit of RNAP II
Termination is imprecise and the transcript is processed
Page 40:
C-terminal Domain of RNAP II RPB1 (or β’–like) Subunit
CTD has repeats of the consensus sequence Tyr-Ser-Pro-Thr-Ser-Pro-Ser (YSPTSPS)
52 repeats in mammals, 26 repeats in yeast
Ser residues subject to reversible phosphorylation
RNAP II initiates only when CTD is unphosphorylated
RNAP II elongates after CTD becomes phosphorylated
Phosphorylated CTD provides binding sites for many proteins
Page 41:
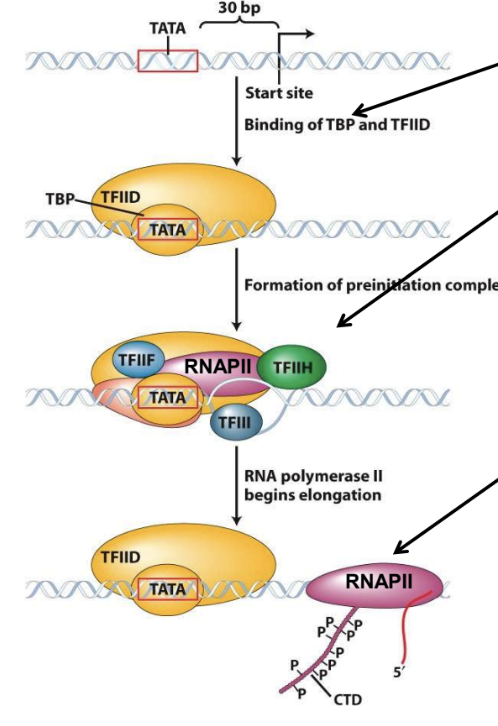
Transcription by Eukaryotic RNAP II
Phosphorylation of carboxyl terminal domain (CTD) of RNAP subunit to begin elongation
Termination is imprecise (transcript is processed)
Binding of RNAP II holoenzyme
Binding of TBP (as part of TFIID)
Page 42:
RNA Processing Prokaryotes vs. Eukaryotes
Prokaryotes:
mRNAs often encode more than one protein (polycistronic)
transcription/translation are coupled
Eukaryotes
transcription/translation are spatially separated
mRNAs generally encode only one protein (monocistronic), or various related proteins due to alternative RNA splicing
most mRNAs translated without further modification
mRNAs undergo extensive modification while still in nucleus
primary transcripts generally not functional
Page 43:
Processing of Eukaryotic mRNAs Produces Mature Transcripts
Addition of ‘cap’ to 5’-end of nascent transcript
Transcribed bases capping enzyme adds “backward G”
Methylated cap – not transcribed
Triphosphate bridge
poly(A) tail added to 3’-end
RNA splicing – coding information is fragmented into exons
Internal modifications, e.g. methylation of some A residues, N6-methyladenosine (m6A)
Page 44:
Processing of Eukaryotic mRNAs Produces Mature Transcripts
poly(A) tail added to 3’-end
RNA splicing – coding information is fragmented into exons
Internal modifications, e.g. methylation of some A residues, N6-methyladenosine (m6A)
Page 45:
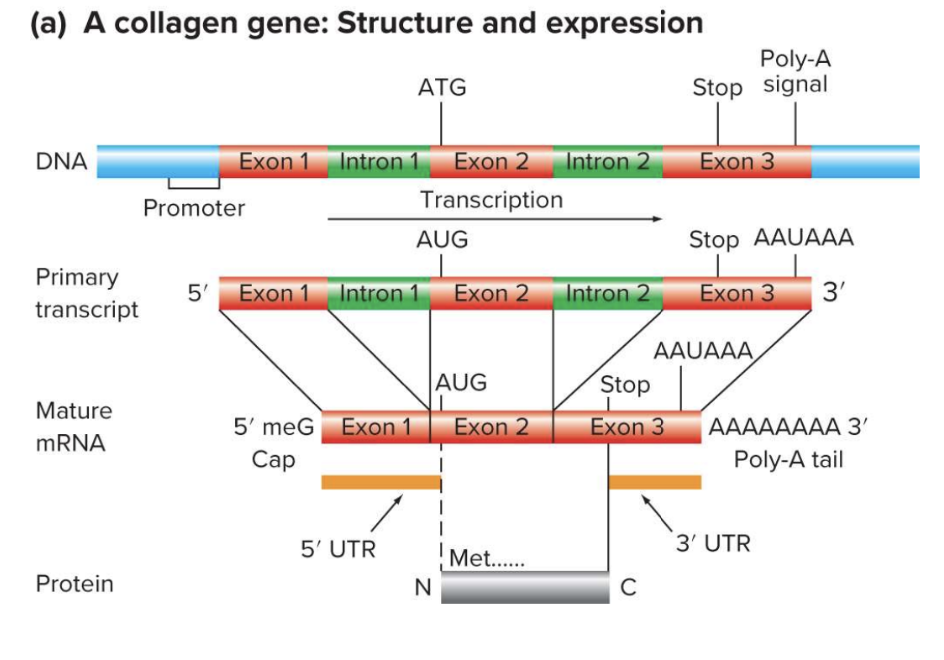
Structure and Expression of a Typical Eukaryotic Gene (a) A collagen gene: Structure and expression
Poly-A
ATG
Stop signal
Page 46:
Alternative Splicing Produces Different mRNAs from the Same Primary Transcript
Alternative splicing uses different 3' cleavage and combinations of polyadenylation exons.
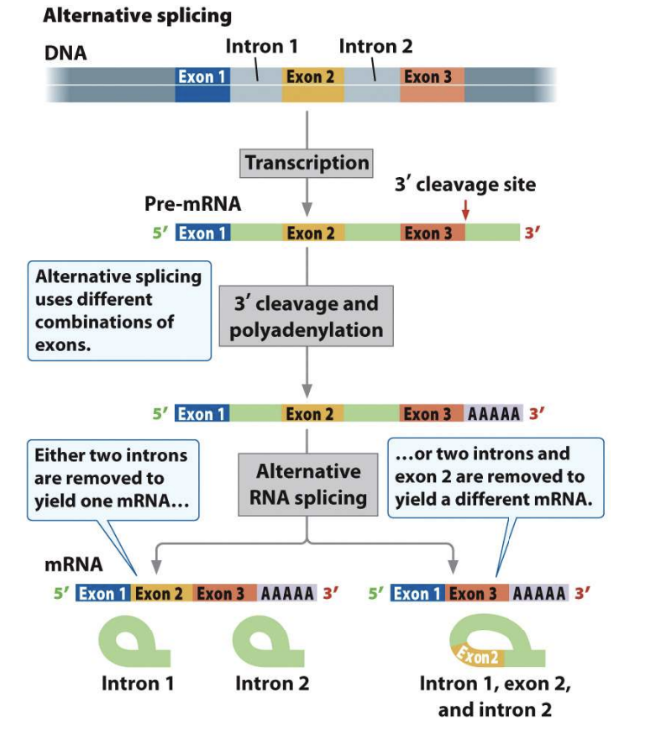
Page 47:
Nature of the Genetic Code
Triplet code
Non-overlapping
Continuous (no punctuation)
Degenerate (most amino acids are specified by more than one codon)
Read continuously from a fixed starting point in mRNA
Page 48:
Reading Frames
Initiating AUG codon (start codon) sets reading frame
DNA duplex has 6 potential reading frames (AUG -> ATG)
An mRNA has 3 potential reading frames but generally only one used
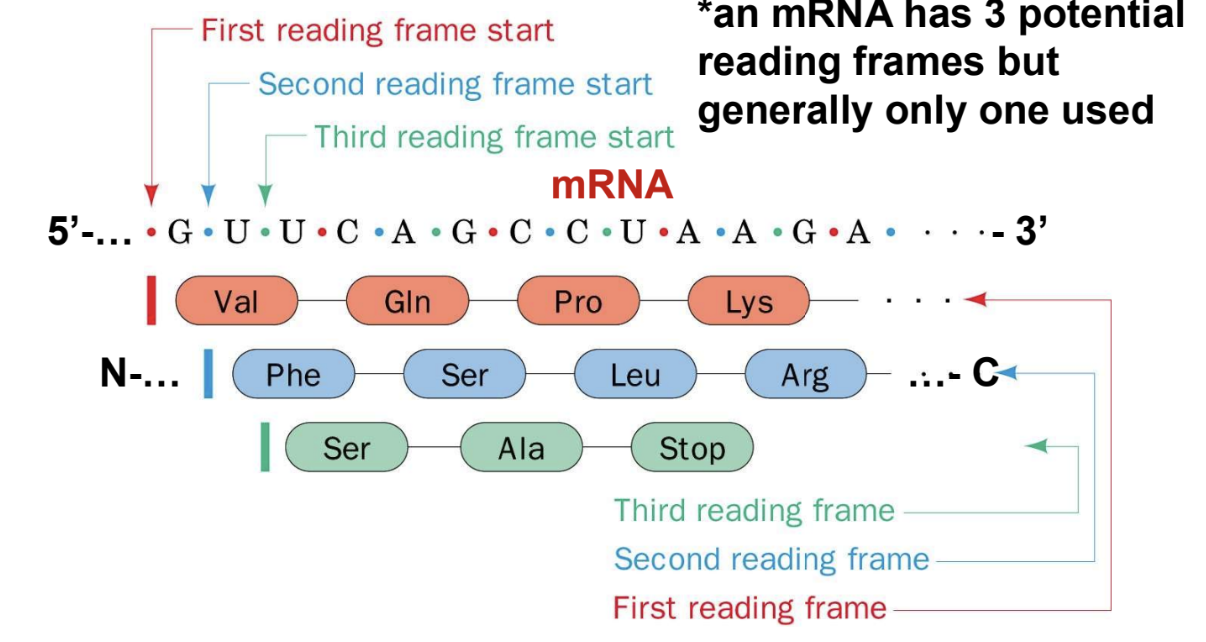
Page 49:
Eukaryotic mRNAs
7methyl-GTP cap is essential for mRNA binding by eukaryotic ribosomes and enhances the stability of mRNAs by preventing degradation by 5’-exonucleases
poly(A) tail enhances both stability and translational efficiency of eukaryotic mRNAs
There is NO Shine-Dalgarno sequence at the 5’-end of eukaryotic mRNAs
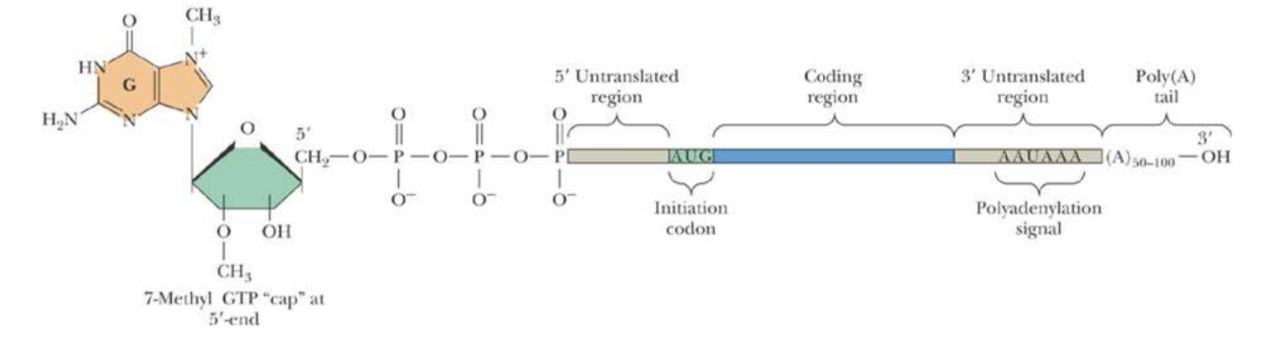
Page 50:
Initiation Phase in Eukaryotes
Small ribosomal subunit binds to 5' cap, then scans the mRNA for the first AUG codon
Initiator tRNA carries Met
CAP-independent translation can occur on some eukaryotic mRNAs, sometimes in response to special conditions
Page 51:
Base Substitutions, Insertions and Deletions
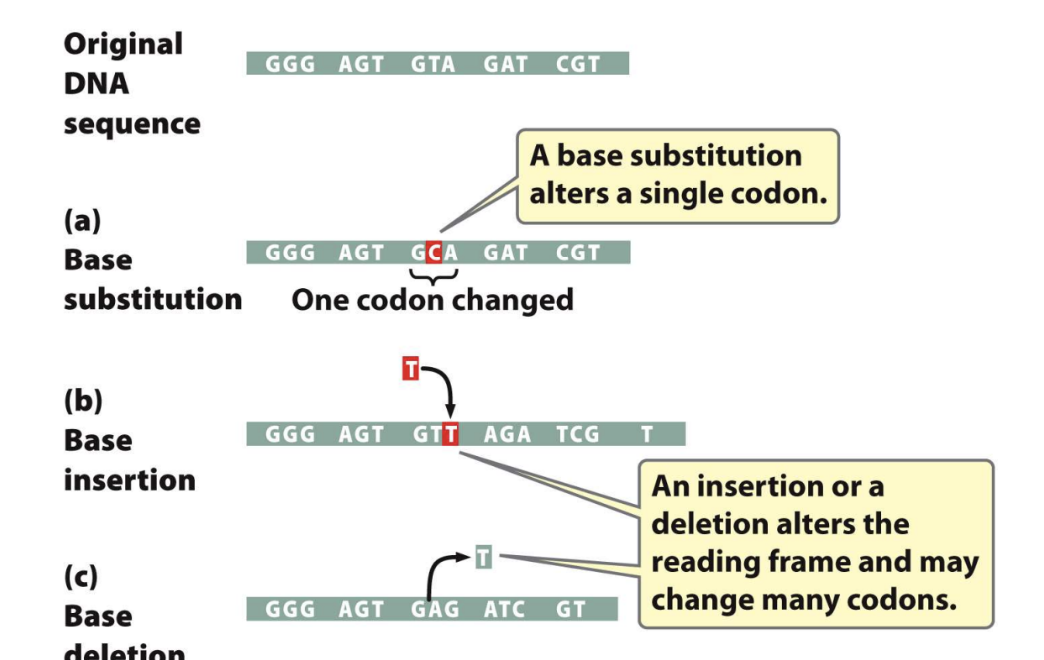
Page 52:
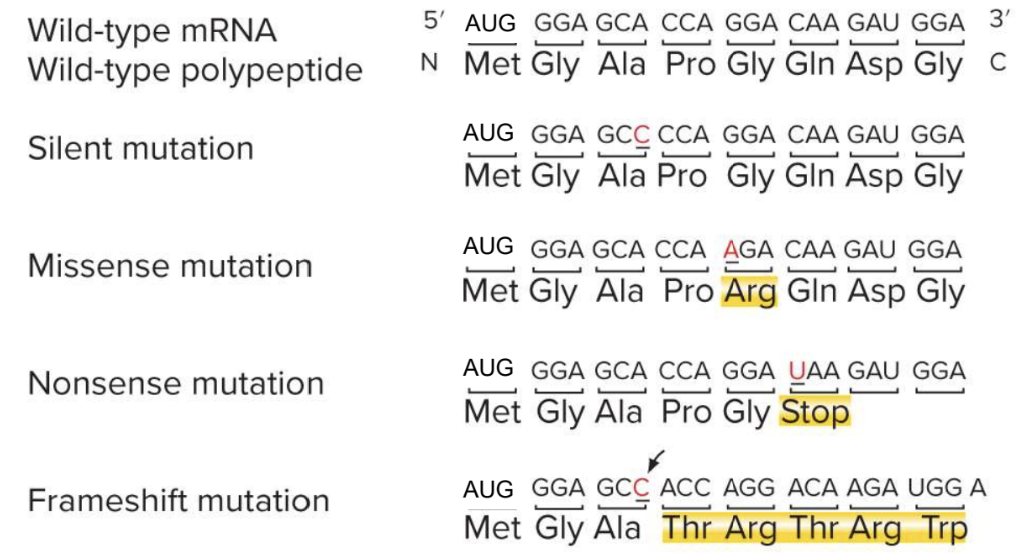
Types of Mutations in the Coding Sequence of Genes
Silent mutation
Missense mutation
Nonsense mutation
Frameshift mutation
Page 53:
Mutations in the Coding Sequence of a Gene can Alter the Gene Product
Silent mutations do not alter the amino acid sequence
Missense mutations replace one amino acid with another
conservative - chemical properties of mutant are similar to the orginal
nonconservative - chemical properties of mutant are different than the original
Nonsense mutations change codon that encodes an amino acid to a stop codon - result in producrion of a truncated protein
Frameshift mutations result from insertion or deletion of nucleotides with the coding region - No framsshift if multiples of threee are inserted or deleted
Page 55:
Point Mutations in Noncoding Regions Can Affect Gene Expression
Mutations in Regulatory and Other Noncoding Sequences:
Binding sites for RNA Polymerase (and associated factors)
Binding sites for regulatory transcription factors
Ribosome binding sites
Polyadenylation signal
5’- and 3’-splice sites, and also branch site
Other sites that regulate translation
Sites that influence RNA stability
Page 57:
Protein-DNA Interactions
Two Types of Interaction:
Non-specific: interactions mainly between functional groups on protein and phospho-ribose backbone of DNA
Specific: recognize specific sequences of nucleotides (base contacts) as well as non-specific portions of DNA
transcription factors
Page 58:
Mapping Protein-DNA complexes
DNA ‘footprinting’ using DNaseI
Create a ladder of DNA fragments by increasing amounts of protein
Page 59:
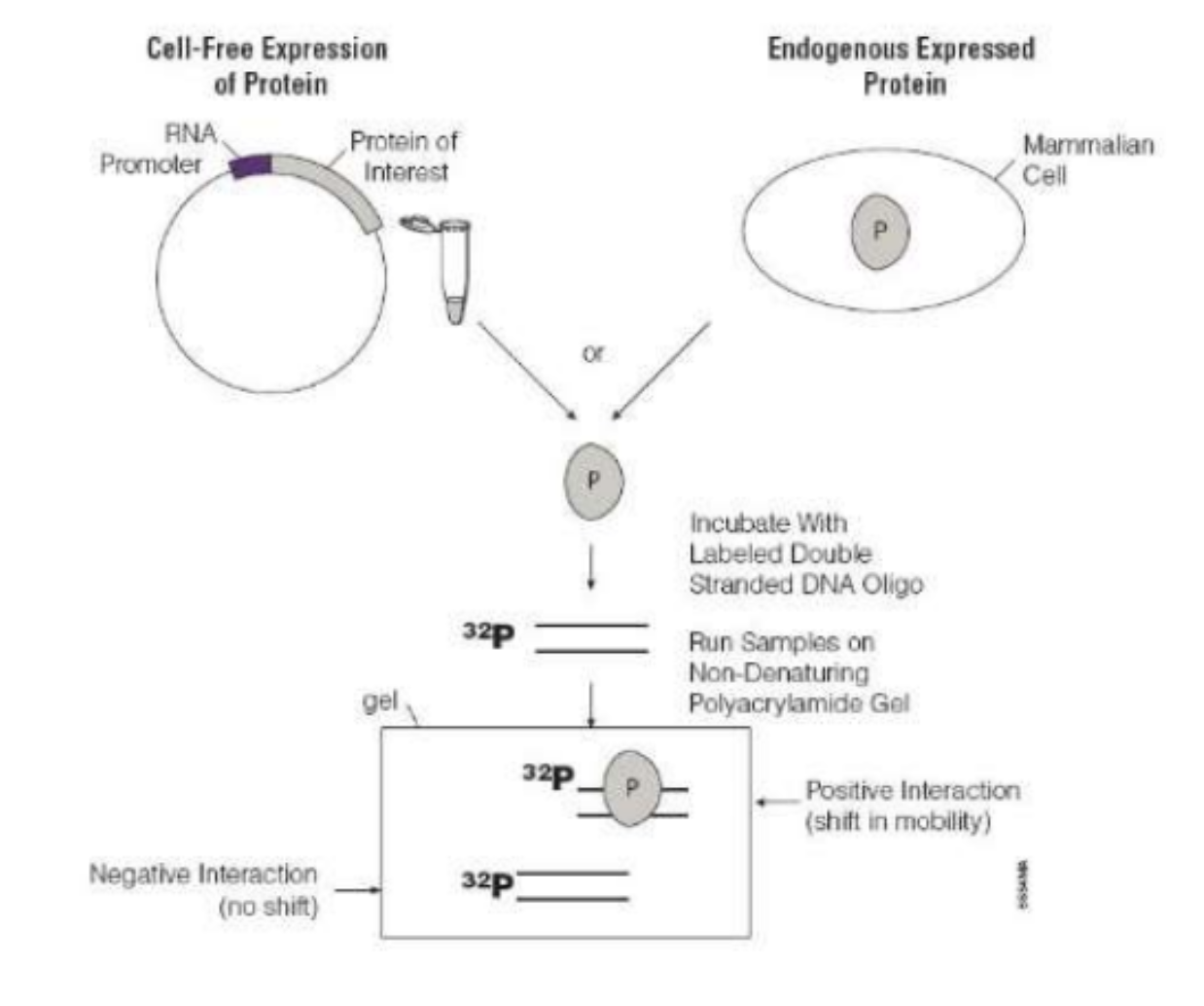
Electrophoretic Mobility Shift Assay (EMSA)
Cell-Free Expression
Endogenous Expressed of Protein
Protein RNA Protein of Mammalian Promoter Interest
Cell P or P Incubate With Labeled Double Stranded DNA Oligo 32P
Run Samples on Non-Denaturing gel Polyacrylamide Gel 32P
Positive Interaction (shift in mobility)
Negative Interaction 32P (no shift)
Page 60:
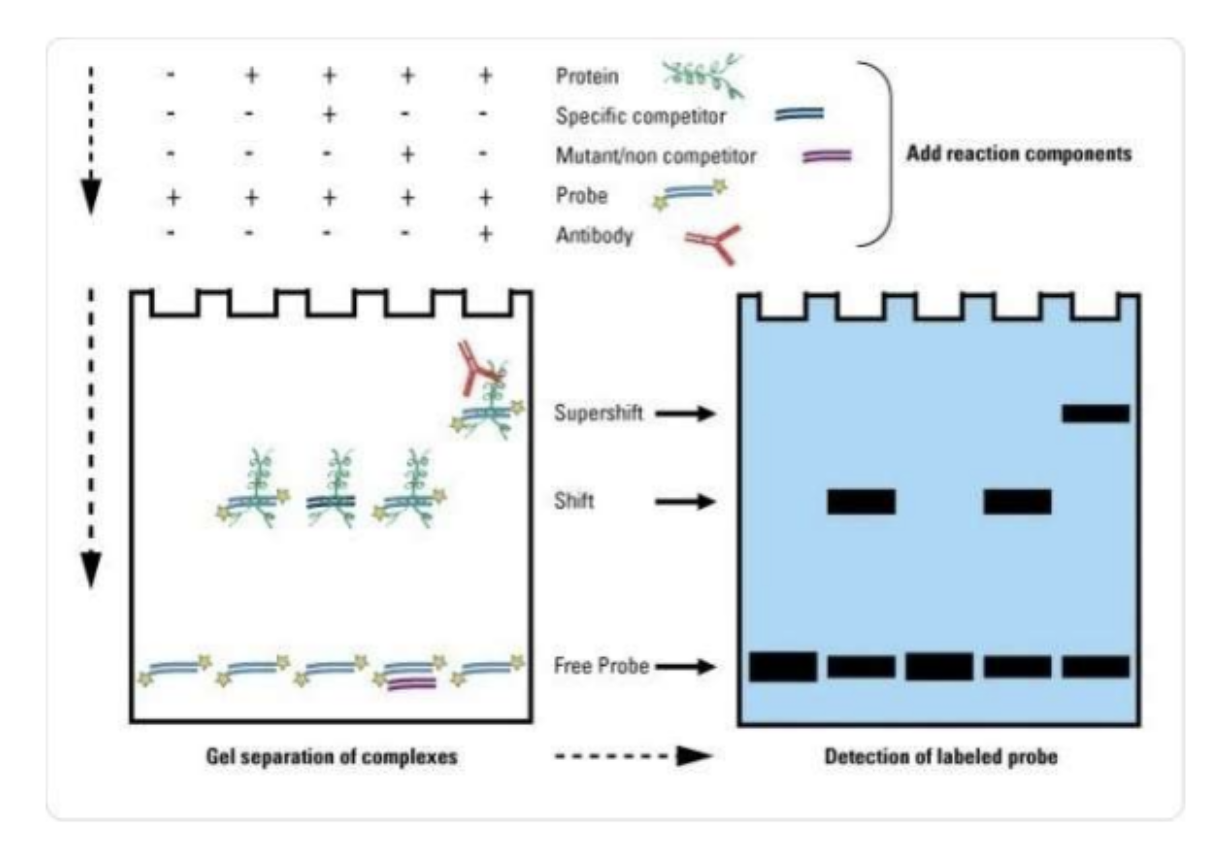
Electrophoretic Mobility Shift Assay (EMSA)
Protein
Specific competitor
Mutant/non competitor
Add reaction components
Probe
Antibody Supershift
Shift 25 Free Probe
Gel separation of complexes
Detection of labeled probe
Page 61:
Chromosomal Packaging and Function
Heterochromatin – highly condensed, usually inactive transcriptionally
Darkly stained regions of chromosomes
Constitutive – condensed in all cells (for example most of the Y chromosome)
Facultative – condensed in only some cells and relaxed in other cells (for example position effect variegation, X chromosome in female mammals)
Euchromatin – relaxed, usually active transcriptionally
Lightly stained regions of chromosomes
Page 62:
DNase I Hypersensitive Sites Within Chromatin (‘in vivo’ hypersensitivity)
DNase I hypersensitive sites: more open chromatin configuration site, upstream of the transcription start site
Relaxation (opening) of the chromatin structure allows transcription factors access to binding sites on the DNA
Page 63:
Chromatin Remodeling and Histone Modifications
Activation of eukaryotic transcription is dependent on two things:
Interaction of RNAPs with the promoter and transcription regulatory proteins
Relief of repression imposed by chromatin structure (nucleosomes)
Page 64:
Chromatin Remodeling Complexes
Chromatin-remodeling complexes (huge) mediate ATP-dependent conformational changes that, for example, peel 50 bp of DNA from edge of nucleosomes
This allows DNA-binding proteins (RNAP II, GTFs, other transcription factors) to access the DNA
Examples of chromatin-remodeling complexes: SWI/SNF, RSC (remodels the structure of chromatin)
Impose “fluid’ state on chromatin that maintains its DNA’s overall packaging but transiently exposes individual sequences to interacting factors
Page 65:
The Four Core Histone Tails Can Be Modified With Chemical Groups
Histone tails extend out from nucleosome, are platforms for modification
Enzymes can add chemical groups (acetyl groups, methyl groups, phosphate groups, ubiquitin, etc.)
Modified tails can alter nucleosomes and bind chromatin modifier proteins
Modifications in N-terminal tails of histones: Lys, Arg, Ser, Thr and His residues
Page 66:
Histone Tail Modifications Alter Chromatin Structure - Acetylation
Histone acetyltransferases add acetyl groups to histone tails
Prevents close packing of nucleosomes
Favors expression of genes in euchromatin
The process is reversed by histone deacetylases
Page 67:
Histone Tail Modifications Alter Chromatin Structure - Methylation
Histone methyltransferases add methyl groups to histone tails
Effect depends on specific amino acid modified (activation or repression)
Adding methyl group to H3 lysine 9 favors heterochromatin formation
The process is reversed by histone demethylases
Page 68:
Histone Writers, Erasers and Readers
Histone ‘Writers’: acetyltransferases, methyltransferes, kinases, ubiquitinases
Histone ‘Erasers’: deacetylases, demethylases, phosphatases, de-ubiquitinases
Histone ‘Readers’: post-translational modifications of histone N-terminal tails are recognized by proteins (‘readers’) that exert function on gene expression (e.g. bromodomain and chromodomain proteins)
Page 69:
Histone Modifications Affect Transcription
Histone acetyl transferases (HATs) acetylate histone tails; many transcription factor co-activators are HATs
Histone acetylation opens the chromatin – favours gene expression
Histone methyltransferases (HMTases) can activate or repress transcription; some HMTases are coactivators and others are corepressors
Histone acetylation and methylation are dynamic – modifications can be taken off rapidly by histone deacetylases or histone demethylases
Page 70:
DNA Methylation
Another change in chromatin structure associated with transcription is methylation of cytosine residues, which occurs most commonly when adjacent to guanine nucleotides (‘CpG’ methylation). This is distinct from histone methylation!
Heavily methylated DNA is associated with repression of transcription in eukaryotes, whereas transcriptionally active DNA is usually unmethylated
So called ‘CpG’ islands are found near transcription start sites (promoters) -> methyl groups are removed before initiation of transcription
An association exists between DNA methylation and histone deacetylation, both of which repress transcription
Page 71:
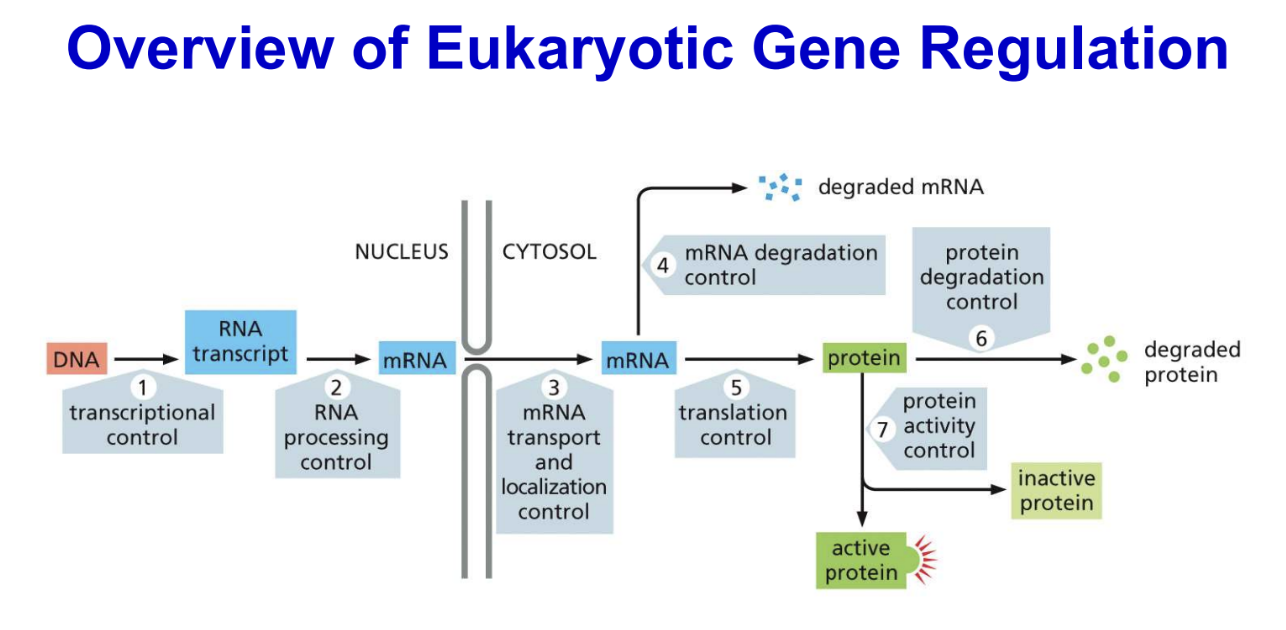
Overview of ‘Epigenetic’ Regulation of Gene Expression
Some of these changes in chromatin state, which are not changes in DNA sequence itself, can be passed down during cell division and even sometimes passed to future generations (epigenetics)
Page 72:
Chromatin Immunoprecipitation (X-ChIP) Assay
X-ChIP = crosslinked ChIP
Know the order of these steps and the reason for each step
Page 73:
Major Cis-Acting Regulatory Elements
Core Promoter – DNA sequence that is usually directly adjacent to the gene. Bound by General Transcription Factors (GTFs)
Often have a TATA box:
Allow basal level of transcription (unregulated)
TATA A
Regulatory Promoter – Other more gene-specific transcription factors bind nearby at the regulatory promoter
Page 74:
Promoters: Core + Regulatory
Promoters contain ‘consensus’ sequences that are mixed and matched in different combinations and in difference promoters.
Different transcription factors bind to each consensus sequence, so each promoter responds to a unique combination of transcription factors.
Page 75:
Major Cis-Acting Regulatory Elements
Enhancers – DNA sequence that can be far away from gene
Augment or repress the basal level of transcription
May be located either 5’ or 3’ to the transcription start site
Still function when moved to different positions or orientations relative to promoter
Page 76:
Reporter Genes Identify Enhancers in Eukaryotes
Enhancers can be identified by:
Constructing a recombinant DNA molecule that has a putative enhancer sequence fused to a promoter + reporter gene such as the green fluorescent protein (GFP), luciferase (LUC) or β-glucuronidase (GUS)
Generating a transgenic organism or cell line that has the recombinant DNA in its genome or transiently present. Or LUC or GUS reporter genes, for example
Page 77:
Deletion Constructs to Identify Important Enhancer cis-Acting DNA Elements
Ozeki et al., 2001, Biochem J.
Foot A +28, therefore includes core promoter
Page 78:
Further Finer-Scale Mutations to Identify Important DNA regulatory sequences (cis elements)
Ozeki et al., 2001, Biochem J.
A + B, enhancer core promoter (-95 to +28)
Page 79:
Proteins Act in Trans to Control Transcription Initiation
Transcription factors
Sequence specific DNA binding proteins
Bind to promoters and enhancers
Recruit other proteins to influence transcription
Three types: basal factors, activators, repressors
Page 80:
Mediator Cryo-EM Structure of RNAP II/Mediator: Human Mediator:
Mediator is a complex of more than 20 proteins
Mediator doesn’t bind DNA directly – bridges RNA pol II at the promoter and activator or repressor proteins at the enhancer
Binds the unphosphorylated form of Pol II but not the phosphorylated form. Phosphorylation of the CTD causes dissociation of Pol II from the mediator to enable initiation of transcription (switch to elongation mode).
Page 81:
Transcription Factors Bind to Sites on DNA and Regulate Transcription
Coactivators: mediate interactions between DNA-binding transcription activators and Mediator. Coactivators do not bind DNA directly themselves. Some Coactivators also have histone acetyltransferase activity to locally further open the chromatin for GTFs / RNAP.
Mechanisms of Activator Effects on Transcription
Activators are responsible for variation in transcription levels of different genes
Activators stimulate recruitment of basal factors and RNA pol II to promoters
Activators recruit coactivators to open chromatin structure
Domains Within Activators
Activator proteins have at least two functional domains
DNA binding domain binds to specific enhancer
Activation domain binds to other proteins (basal factors, coactivators, mediator)
Dimerization domain allows interaction with other proteins
Repressor Proteins Suppress Transcription Initiation by Recruiting Corepressors
Corepressors prevent RNA pol II complex from binding the promoter
Corepressors modify histones to close chromatin structure
Corepressors do not bind DNA directly, often recruit histone deacetyltransferases
Repressor Proteins Can Act Through Competition With an Activator Protein
Indirect repressor interferes with the function of an activator
Competition due to overlapping binding sites
Repressor binds to activation domain (quenching)
Repressor can bind to activator and keep it in the cytoplasm
Repressor can bind to activator and prevent homodimerization
Cell-Type Specific Transcription is Achieved by Changes in Transcription Factors
Function of trans-acting proteins changes through allosteric interactions, modification (e.g. phosphorylation), and transcription factor cascades
Complex Regulatory Regions Enable Fine-Tuning of Gene Expression
~2000 genes encode transcriptional regulatory proteins in humans
Each regulatory protein can act on many genes
Each enhancer has binding sites with varying affinities for activators and repressors
How Does an Enhancer Know Which Genes to Regulate?
Insulators are sequences located between an enhancer and a promoter that block access to the promoter
Example: Galactose Metabolism in Yeast through GAL4 Transcription Factor
GAL4 binds to the UASG site and controls the transcription of genes involved in galactose metabolism
UAS (Upstream Activation Sequence) in yeast is a type of enhancer acting at a distance
Small RNAs Regulate mRNA Stability and Translation
Specialized RNAs (miRNAs, siRNAs, piRNAs) prevent expression of specific genes through complementary base pairing
Small RNAs are found in eukaryotes and regulate gene expression, defense against viruses, suppression of transposons, and modification of chromatin structure
RNA Interference (RNAi)
RNAi is a conserved biological response to double-stranded RNA that mediates resistance to endogenous parasitic and exogenous pathogenic nucleic acids
siRNAs and miRNAs are involved in RNAi
siRNAs degrade mRNA, inhibit transcription, and modify chromatin
miRNAs degrade mRNA, inhibit translation, and modify chromatin
Mechanism of siRNA Production and Function
dsRNAs are processed by Dicer to produce short ~22 bp siRNAs
siRNAs form RNA-protein complex with Argonaute proteins and interfere with gene expression or destroy viral mRNAs
siRNAs are useful experimental tools for selectively knocking down expression of target genes
Primary Transcripts Containing miRNA
Most miRNAs are transcribed by RNA polymerase II, some by RNA polymerase III
Primary transcripts have double-stranded stem loops
miRNAs can be processed from introns of protein-coding transcripts
miRNA Processing
Drosha excises stem-loop from primary miRNA to generate pre-miRNA
Dicer processes pre-miRNA to a mature duplex miRNA
One strand is incorporated into miRNA-induced silencing complex (RISC)
Two Ways That miRNAs Can Down-Regulate Expression of Target Genes
When complementarity is perfect, target mRNA is degraded
When complementarity is imperfect, translation of mRNA target is repressed
miRNA binding sites are usually in the 3’-UTR of the target mRNA
microRNAs
Thousands of novel miRNAs in plants, worms, flies, and mammals
Each miRNA can potentially base pair with sequences on hundreds of different target mRNAs
miRNAs play important roles in diseases and disorders, including cancer
Mechanisms Regulating mRNA Translation
Control of translation often occurs at initiation
Small subunit of ribosome recognizes a complex structure built around the 5’ cap of the mRNA
eIFG protein in this complex binds to poly-A binding protein (PABP) at the poly-A tail to circularize mRNA
Regulating mRNA Translation in Response to Nutrients
4E-BP1 binds to initiation factor eIF4E and blocks initiation
Presence of nutrients and growth factors leads to phosphorylation of 4E-BP1
Translational Control Through poly-A Tail Length
Longer poly-A tails bind PABP more efficiently, leading to more efficient formation of translation initiation complex
Artificial Transformation of Bacteria (e.g. E. coli)
E. coli can be transformed artificially by exposing bacteria to a salt solution and applying heat shock or exposing bacteria to electric current in the presence of foreign DNA
Plasmids: Selection of E. coli Transformants by Using an Antibiotic Resistance Gene
Antibiotic selection is key to obtain only bacteria that have taken up the plasmid
Transformants will grow on the selection media
Restriction Enzymes (Endonucleases)
Restriction enzymes recognize and cleave specific DNA sequences
Recognition sites are typically 4-8 bp in length and palindromic
Restriction enzymes are produced by bacteria for protection against viruses
Restriction enzymes create sticky or cohesive ends with 5’- or 3’- overhangs
Page 104: Restriction Mapping
Three portions of cloned DNA are divided and cut with different restriction enzymes (EcoRI, BamHI, EcoRI and BamHI).
The digested samples are loaded into a gel along with size markers.
The gel results show the presence of BamHI sites at 7 kb and 5.5 kb, and EcoRI sites at 14 kb, 5 kb, and 4.5 kb.
A restriction map can be created based on the presence of these sites.
Page 105: DNA Methylation and Restriction Enzymes
DNA methylation can affect the ability of a restriction enzyme to digest DNA.
Mammalian CpG methylation and Dam and Dcm methylation in prokaryotes can impact restriction enzyme activity.
Page 106: Cloning Fragments of DNA
Molecular cloning is used to purify a specific DNA fragment and make multiple copies of it.
The process involves inserting DNA fragments into cloning vectors and then transferring the recombinant DNA into living cells.
The group of replicated DNA molecules is called a DNA clone.
Page 107: Cloning Vectors: Plasmids
Plasmids are commonly used as cloning vectors.
They have an origin of replication for DNA replication in host cells.
Plasmids are small and easy to handle.
They have unique restriction sites for cloning DNA fragments.
They also have selectable markers to verify the presence of foreign DNA.
Page 108: pUC19 Plasmid: A Typical Cloning Vector
The pUC19 plasmid has a "multiple cloning site" (MCS) or polylinker, which contains unique cutting sites for different restriction enzymes.
Page 109: Expression Vector
In gene cloning, the goal is not just to replicate the gene but also to produce the protein it encodes.
Expression vectors allow for the transcription and translation of the inserted coding sequence.
The genetic code is the same between humans and E. coli, allowing for the expression of human genes in E. coli.
Page 110: Various Cloning Vectors
Bacterial plasmids like pUC19, pBR322, and pBluescript II are commonly used cloning vectors.
Bacteriophages like lambda phage vectors and cosmids can also be used.
Yeast artificial chromosomes (YACs) and bacterial artificial chromosomes (BACs) are used for larger DNA inserts.
Shuttle/binary vectors are species-specific, such as plant vectors like binary Ti plasmids and yeast shuttle vectors.
Page 111: Creating Recombinant DNA Molecules With Plasmid Vectors
DNA ligase is used to seal the phosphodiester backbones between the vector and inserted fragment.
Linearized plasmids are often dephosphorylated at the 5'-ends to reduce self-ligation.
Page 112: Host Cells Take Up and Amplify Recombinant DNA
Transformation is the process by which a cell or organism takes up foreign DNA.
In E. coli, only a small percentage of cells will be transformed with a plasmid.
Cells with the plasmid will grow on media with ampicillin.
Each cell with the plasmid will produce a colony on an agar plate, resulting in millions of identical plasmids in the colony.
Page 113: Construction of a Recombinant DNA Molecule
Recombinant DNA molecules can be created by combining a vector and an insert, such as an EcoRI-digested DNA fragment.
The insert can come from another plasmid, a PCR product, chemically synthesized DNA, etc.
Page 114: Directional Cloning
Directional cloning involves using two different restriction enzymes to ensure the insert ligates into the recipient plasmid in only one direction.
This is achieved by using compatible overhangs between the insert and plasmid.
Page 115: Directional Cloning: 'Subcloning' Example
An example of directional cloning is shown using the T7 promoter, EcoRI, NotI, CMV promoter, XhoI, and Neo insert.
The digested donor insert is ligated into the recipient plasmid in a specific orientation.
Page 116: Directional Cloning: A Detailed Look
The donor plasmid and recipient plasmid are digested with restriction enzymes.
The digested insert and plasmid are combined and ligated together.
The resulting recombinant DNA molecule has the insert in a specific orientation.
Page 117: Cloning with PCR-Amplified Inserts
PCR-amplified inserts are often used in cloning.
Some restriction enzymes do not cut efficiently at the end of a linear piece, so adding 3-6 bases upstream of the restriction site can improve cutting efficiency.
Page 118: Verification of Recombinant DNA Construct
Several methods can be used to verify the presence and integrity of the insert in the plasmid, such as colony PCR, diagnostic restriction digests, and sequencing.
Page 119: DNA Libraries
A DNA library is a collection of cloned fragments representing the genes of a specific organism or the mRNAs from an organism.
Genomic libraries are prepared by isolating total DNA, digesting it into fragments, and cloning them into a vector.
cDNA libraries are constructed by synthesizing cDNA from mRNA templates using reverse transcriptase.
Page 120: Genomic DNA Library
Genomic DNA can be digested with restriction enzymes to create fragments.
These fragments can be cloned into plasmids or phages to create a genomic library.
Page 121: Screening a Genomic DNA Library
Multiple master plates are created for each library.
Probes, which are partial DNA sequences, are used to hybridize to the locus (gene) of interest.
Page 122: Converting mRNA Transcripts to cDNA
mRNA can be isolated from other RNAs in eukaryotic cells based on their poly A tails.
Oligo-dT is used to hybridize to the poly A tails and initiate the synthesis of cDNA.
Page 123: Converting RNA Transcripts to cDNA
Reverse transcriptase is used to synthesize cDNA from mRNA templates.
The cDNA synthesis is initiated using oligo-dT and the appropriate nucleotides.
Page 124: Creating the Second DNA Strand Complementary to the First cDNA Strand
mRNA is digested with RNase
This breaks down the mRNA into smaller fragments.
The 3' end of cDNA folds back and acts as a primer for 2nd strand synthesis
The cDNA strand forms a loop structure, allowing it to serve as a starting point for the synthesis of the second cDNA strand.
The first cDNA strand acts as a template for the synthesis of the second cDNA strand
Using DNA polymerase and dNTPs, the second cDNA strand is synthesized based on the template provided by the first cDNA strand.
Double-stranded cDNA can be cloned into a plasmid
The resulting double-stranded cDNA can be inserted into a plasmid for further manipulation and analysis.
cDNA library can be screened to identify individual clones of interest
Similar to screening a genomic library, probes corresponding to the sequence of interest can be used to identify specific cDNA clones.
Page 125: A Comparison of Genomic and cDNA Libraries
Genomic library represents all regions of DNA equally, including introns
A genomic library contains DNA fragments that represent the entire genome, including both coding and non-coding regions.
cDNA library includes only exons from part of the genome that was transcribed for translation in cells
In contrast, a cDNA library contains DNA fragments that are derived from the mRNA molecules present in cells. It only includes the coding regions (exons) of the genome.
Page 126: cDNAs and Alternative Splicing
Alternative splicing can produce different proteins from a single gene
Alternative splicing is a process where different combinations of exons are included or excluded during mRNA processing, leading