09-Biochemistry-Lecture9 | Week 5 - Lecture 2
Transcription: DNA-dependent synthesis of RNA
RNA Metabolism Overview
Overview of RNA Functions
Well-Understood Roles of Ribonucleic acids:
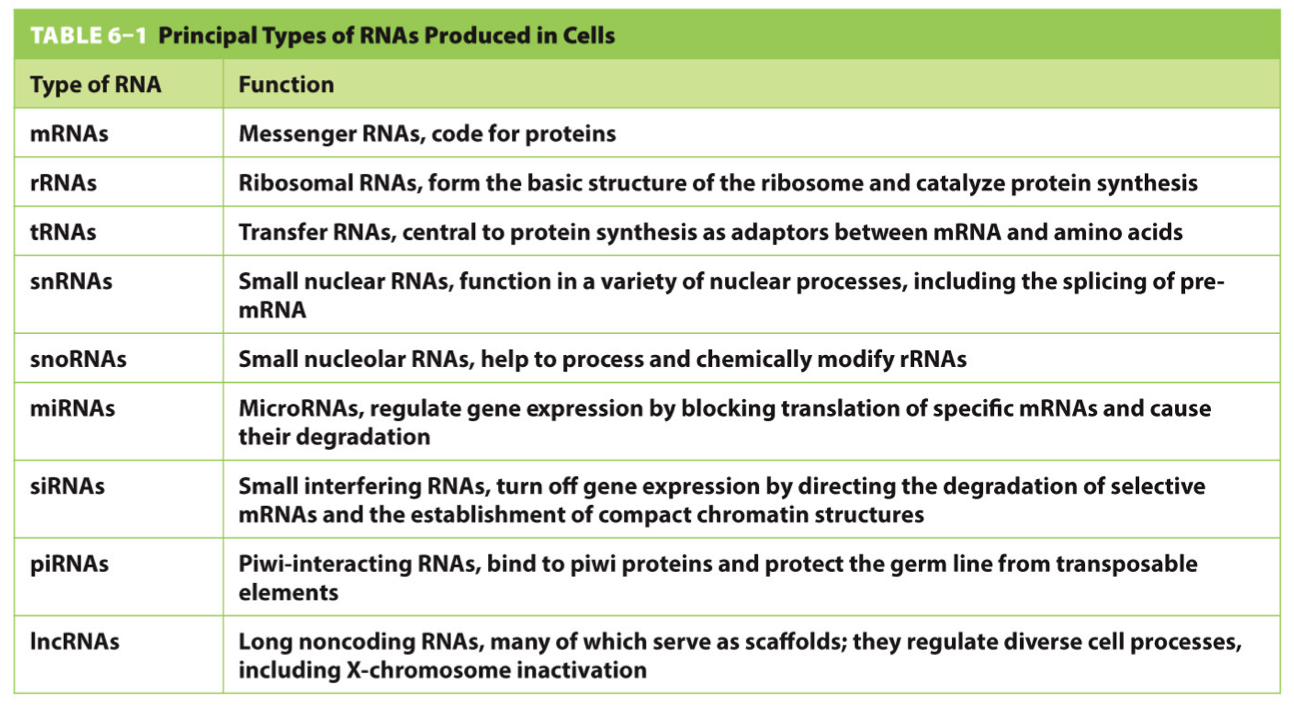
Messenger RNAs (mRNAs): encode amino acid sequences of polypeptides.
Transfer RNAs (tRNAs): match specific amino acids to mRNA codons during protein synthesis.
Ribosomal RNAs (rRNAs): structural and catalytic components of ribosomes.
Less-Understood Functions:
MicroRNA: regulates gene expression by binding to specific sequences.
Long noncoding RNAs (lncRNAs): involved in cell differentiation and identity maintenance.
RNAs act as genomic material in RNA viruses.
Principal Types of RNAs
Overview of RNA Metabolism
Synthesis:
Ribonucleic acids are synthesized using DNA templates in transcription.
Transcription is tightly regulated to control protein concentrations.
RNA Structure:
Single-stranded RNA can fold into 3D structures with specific functions.
Some RNAs function as catalysts (ribozymes).
Post-transcriptional Processing: Most eukaryotic ribonucleic acids are processed after synthesis
Elimination of introns and joining of exons.
Poly-adenylation at the 3’ end and capping the 5’ end.
Transcription in E. Coli: DNA-Dependent Synthesis of RNA
Features:
Nucleoside triphosphates are added to the 3’ end of the RNA strand.
The growing chain is complementary to the template strand in DNA
RNA synthesis is catalyzed by RNA polymerase over a segment of about 35 bp of DNA.
Transcription by RNA Polymerase in E.coli:
Catalytic Mechanism:
Similar to DNA polymerase, involves two Mg2+ ions facilitating RNA synthesis.
One Mg2+ aids in the attack of the 3’-hydroxyl group, the other stabilizes the reaction state by facilitating the displacement of PPI.
First Steps in Transcription:
RNA polymerase binds to the promoter sequence to initiate transcription without a primer.
The growing RNA base-pairs with the DNA template and an unwinding of the DNA duplex forms a transcription bubble.
RNA Polymerase generates positive supercoils ahead, later relieved by topoisomerases
The Transcription Bubble:
For synthesis of an RNA strand complementary to one of two DNA strands in a double helix, the DNA is transiently unwound.
About 17 bp of DNA are unwound during transcription.
RNA polymerase facilitates RNA synthesis as it moves along the template strand.
The DNA is unwound ahead and rewound behind as RNA is transcribed.
Movement of RNA Polymerase Along The DNA Creates Supercoils:
Movement of an RNA polymerase creates positive supercoils ahead of it and negative supercoils behind.
Topoisomerases rapidly eliminate the positive supercoils and regulate the level of negative supercoiling
The RNA polymerase is in close contact with the DNA ahead of the transcription bubble as well as with the separated DNA strands and the RNA within and immediately behind the bubble.
A channel in the protein funnels new NTPs to the polymerase active site.
The polymerase footprint encompasses about 35 bp of DNA during elongation.
Terminology:
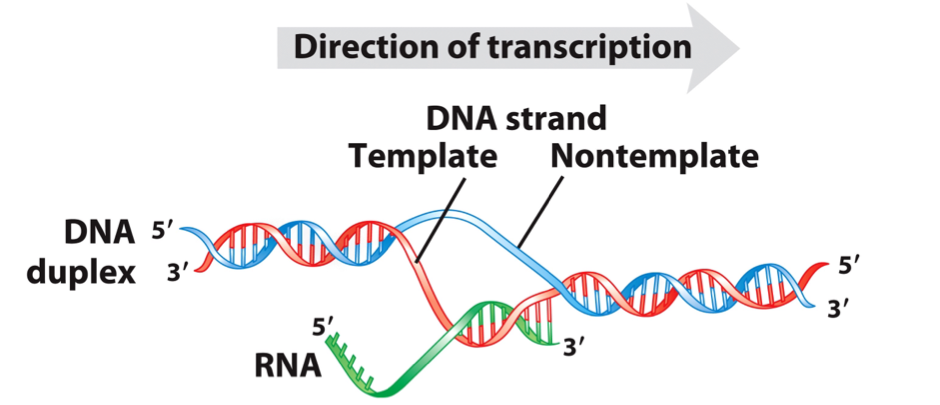
DNA Template Strand: serves as a template for RNA synthesis.
DNA Coding Strand: non-template strand with identical sequence to the RNA transcript except for T replaced by U.

Template and non-template (coding) DNA strands.
The two complementary strands of DNA are defined by their function in transcription.
The RNA transcript is synthesized on the template strand and is identical in sequence to the non-template strand, or coding strand.
Both Strands May Encode For Proteins
Both DNA strands can encode proteins: Coding information may be present on either strand, referred to as the "top" and "bottom" strands.
Adenovirus genome encodes proteins from both double-stranded DNA strands
Adenovirus has a linear genome: Its genome is a double-stranded DNA molecule of 36,000 base pairs.
The direction of transcription is determined by the orientation of the promoter at the beginning of each gene.
Most proteins are encoded by the top strand, which is oriented 5’ to 3’ from left to right.
The bottom strand serves as a template for these transcripts.
Some proteins are encoded by the bottom strand, which is transcribed in the opposite direction using the top strand as a template.
RNA Polymerase Is A Large Enzyme with No Proofreading Capability
RNA polymerase holoenzyme consists of five core subunits (α₂ββ’ω) plus a σ (sigma) subunit
RNA polymerase lacks 3’ → 5’ exonuclease activity, meaning it cannot proofread, leading to a high error rate (1 in 10⁴–10⁵ nucleotides).
RNA polymerase binds to promoter regions to initiate transcription.
Structure of Bacterial RNA Polymerase:
The crab claw shape formed by subunits, with different subunits responsible for various functions, including binding and catalysis.
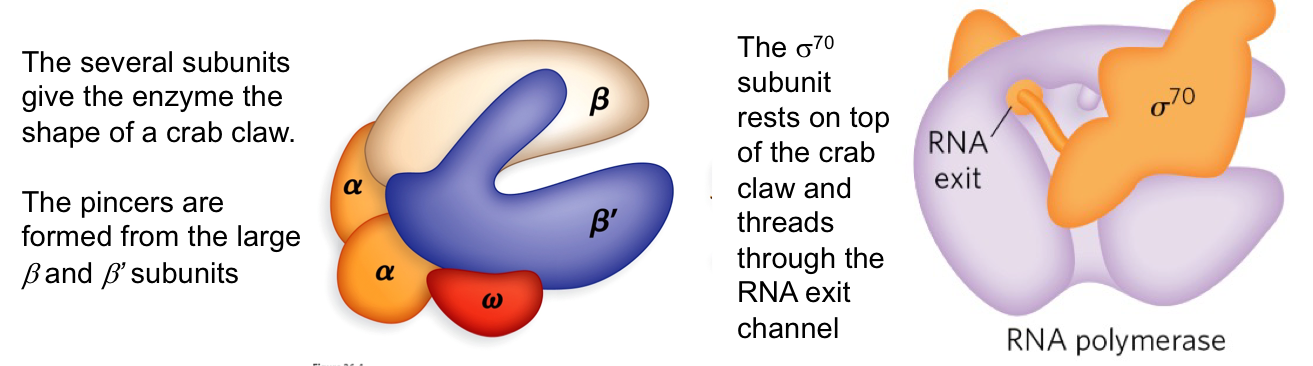
Two α subunits are involved in assembly and binding to upstream promoter elements.
The β subunit is the main catalytic subunit, responsible for polymerization activity.
The β’ subunit is responsible for DNA binding.
The σ⁷⁰ subunit directs the enzyme to the promoter (important for transcription initiation).
The ω subunit helps protect RNA polymerase from denaturation.
Promoters in E. Coli That Bind The Same RNA Polymerase Have Common Features:
A-T rich sequences (Upstream Promoter Elements or UP elements) are located between -40 and -60 and bind the α subunit of RNA polymerase, helping to promote strand separation.
UP elements regulate RNA polymerase binding efficiency, affecting gene expression levels.
Two consensus sequences are essential for σ subunit binding:
-10 sequence (TATAAT) & -35 sequence (TTGACA)
These sequences are collectively called TATA sequences.
Some E. coli Promoters Recognized By an RNA Polymerase Holoenzyme Containing σ⁷⁰
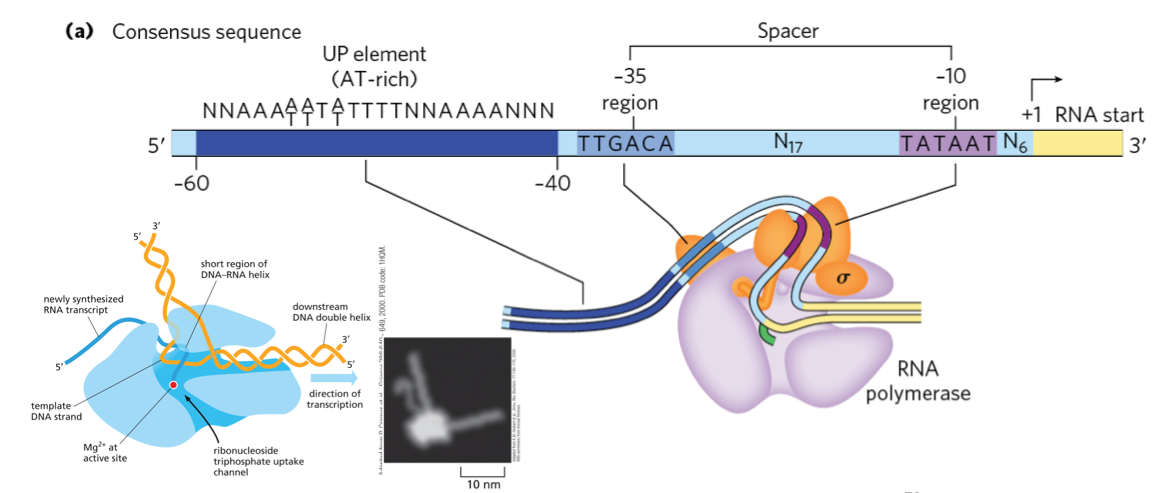
The consensus sequence for E. coli promoters recognized by σ⁷⁰ is shown on the non-template strand, read in the 5’ → 3’ direction.
Sequences vary among promoters, but the -10 (TATAAT) and -35 (TTGACA) regions show significant similarities across different promoters.
The UP element (AT-rich region between -40 and -60) is not present in all E. coli promoters but enhances transcription when present.
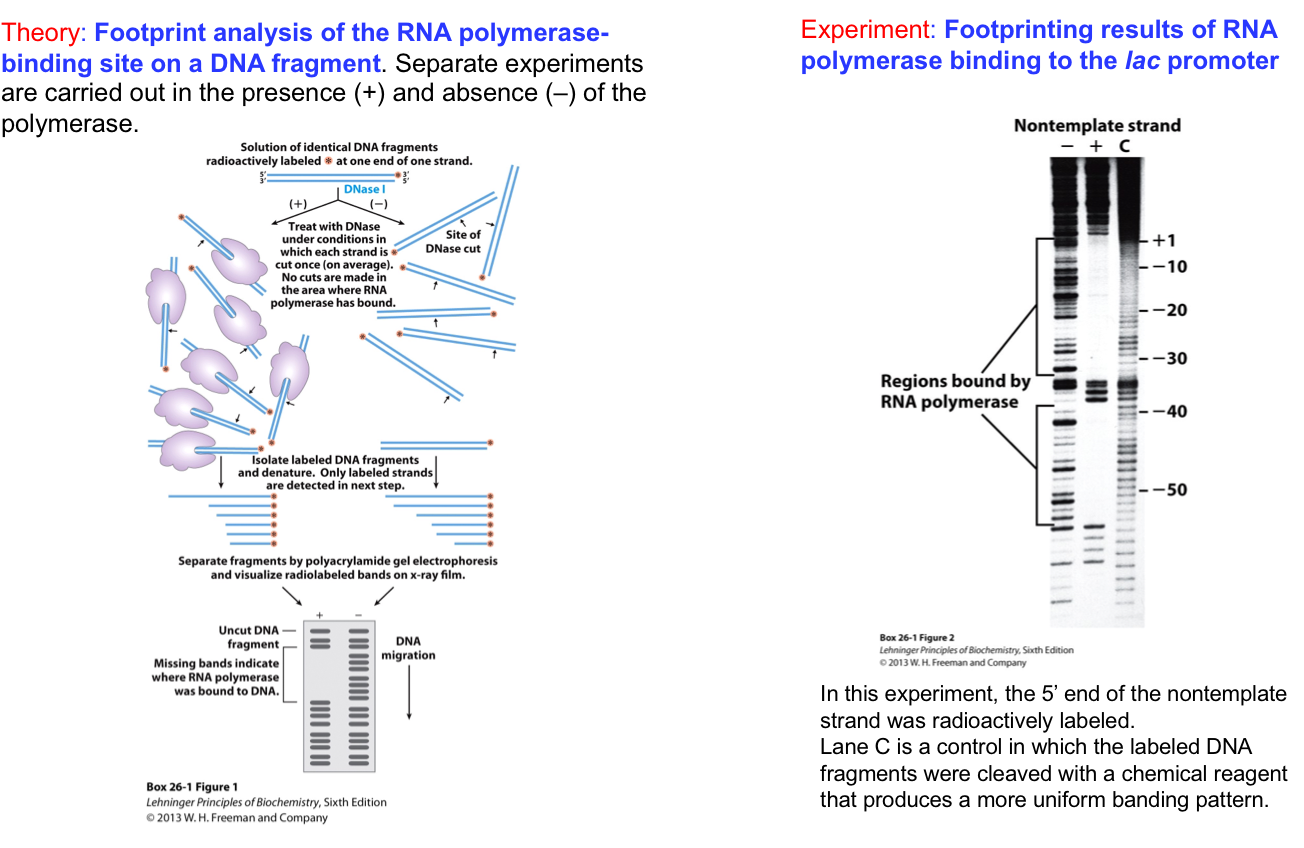
RNA Polymerase Leaves Its Footprint on a Promoter
Premise: DNA bound by RNA polymerase (or any protein) is protected from chemical or enzymatic cleavage at the binding site.
Experimental steps to detect RNA polymerase binding (DNA footprinting assay):
Isolate a DNA fragment suspected to contain a binding site.
Radiolabel the DNA for visualization.
Bind the protein (RNA polymerase) to the DNA in one test tube; keep another as a "naked DNA" control.
Treat both samples with a cleavage agent (chemical or enzymatic).
Run gel electrophoresis to separate the fragments and visualize bands on X-ray film or an imager plate.
Footprinting Techniques
A technique used to identify RNA polymerase binding sites on DNA by comparing protected and unprotected fragments after cleavage.
Transcription Initiation and Elongation Steps:
Transcription initiation begins with RNA polymerase (RNA Pol) binding to the promoter in the presence of σ factor.
This forms a closed complex, where DNA is not yet unwound.
An open complex forms, leading to:
DNA unwinding from ~ -10 to +2 (the transcription bubble).
RNA polymerase moves away from the promoter, transitioning to elongation:
The σ factor is replaced by NusA, which helps in transcription elongation.
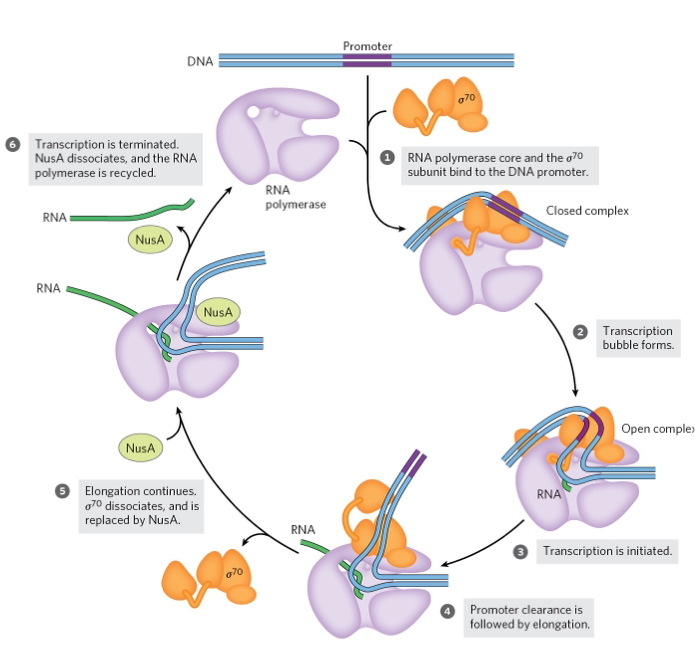
Transcription Initiation & Elongation in E. coli
Transcription Initiation in E. coli
Initiation of transcription occurs in two phases: binding and initiation.
In the binding phase, RNA polymerase interacts with the promoter, forming a closed complex where DNA is bound but not unwound.
An open complex forms when a 12–15 bp region of DNA unwinds from -10 to +2 or +3.
The initiation phase includes transcription initiation and promoter clearance (steps 1–4).
Elongation Phase
Once elongation starts, the σ⁷⁰ subunit is released and replaced by NusA.
RNA polymerase clears the promoter and is now committed to RNA elongation.
Termination Phase
When transcription is complete, the RNA is released, NusA dissociates, and RNA polymerase detaches from DNA (step 6).
A new σ subunit binds to RNA polymerase, allowing the cycle to restart.
Transcription Termination in E. Coli
Types:
ρ-independent: Forming a hairpin structure in the mRNA causing polymerase to pause.
ρ-dependent: Requires ρ protein to dissociate mRNA from RNA polymerase.
Termination of Transcription in E. coli
ρ-Independent Termination (Intrinsic Termination)
RNA polymerase pauses at specific DNA sequences, some acting as terminators.
Two possible outcomes:
RNA polymerase bypasses the site and continues transcription.
A conformational change (isomerization) occurs, leading to termination.
Formation of a hairpin structure in the RNA transcript disrupts:
The RNA-DNA hybrid.
Interactions between RNA and RNA polymerase.
Both of the above.
The RNA-DNA hybrid at the 3' end is unstable, causing RNA to dissociate, leading to termination.
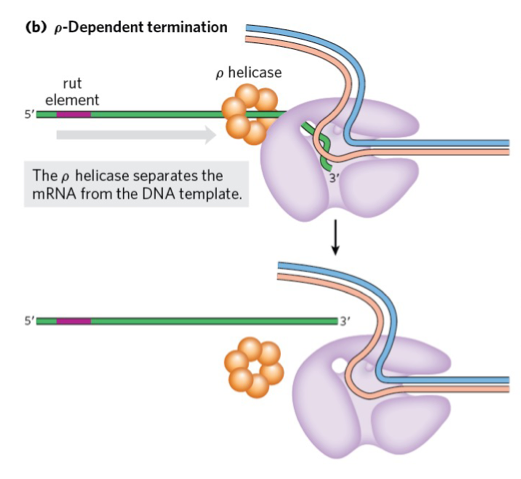
ρ-Dependent Termination
RNA transcripts with a rut (Rho utilization) element recruit Rho (ρ) helicase.
ρ helicase moves along RNA (5’ → 3’) and separates RNA from the polymerase, stopping transcription.
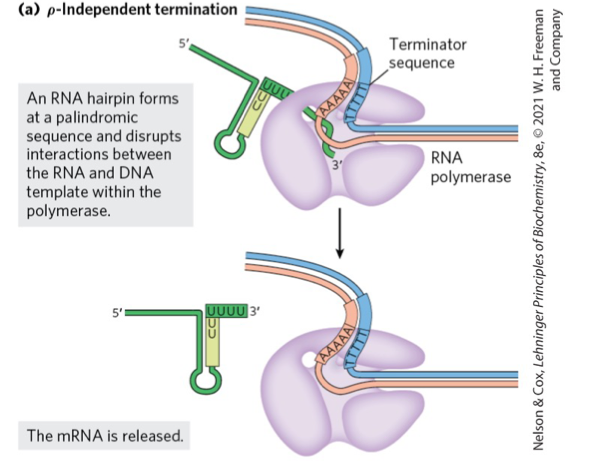
RNA hairpin Formation in ρ-Independent Termination of Transcription
A termination signal at the 3' end of mRNA forms a hairpin structure.
This structure is created by inverted repeat sequences followed by a U-rich region.
The hairpin causes RNA polymerase to pause, leading to dissociation and termination of transcription.
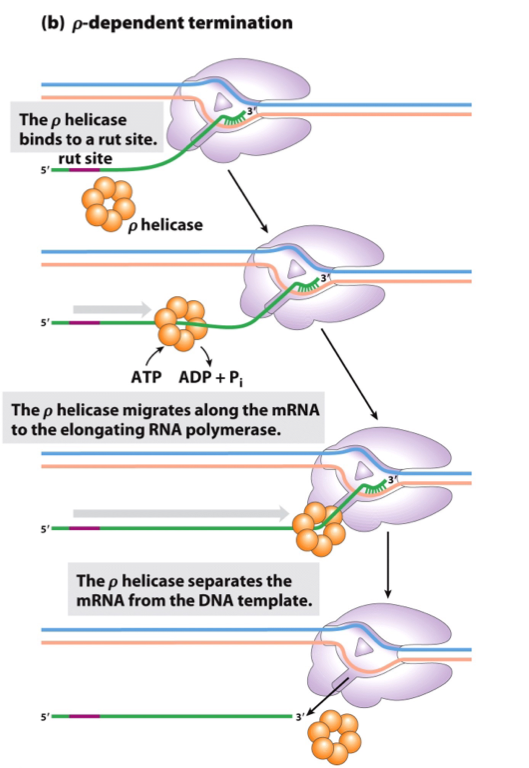
ρ-Dependent Termination of Transcription
RNAs with a rut site recruit the ρ helicase (ρ factor).
ρ helicase moves along mRNA (5’ → 3’) using ATP.
It unwinds the RNA-DNA hybrid
Regulation of Transcription
Transcription is energy-intensive and regulated at initiation via promoter affinity and regulatory proteins.
Summary of RNA Polymerase Function
Stages of Initiation, Elongation, and Termination completely processive without requiring a primer.
RNA Polymerase:
Searches DNA for initiation sites (promoters)
Unwinds a short stretch of dsDNA
Selects the correct NTPs
Does not require a primer to initiate RNA synthesis
Elongation proceeds in 5’ to 3’ direction
Is completely processive
Detects termination signals
Its activity can be modified by activator and repressor proteins
Transcription in Eukaryotes: DNA-Dependent Synthesis of RNA
Eukaryotes Contain Several Distinct RNA Polymerases
RNA polymerase I synthesizes pre-ribosomal RNA
RNA polymerase II is responsible for synthesis of mRNA
Very fast (500–1000 nucleotides/sec)
Specifically inhibited by mushroom toxin a-amanitin
Can recognize thousands of promoters
RNA polymerase III makes tRNAs and some small RNA products
Plants appear to have RNA polymerase IV that is responsible for the synthesis of small interfering RNAs
Mitochondria have their own RNA polymerase
Features of Some Promoters Recognized By Eukaryotic RNA Polymerase II
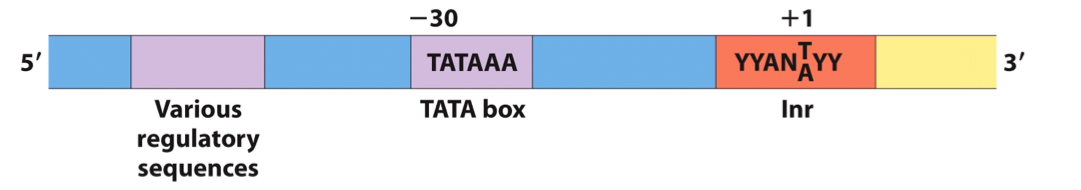
TATA box is the main assembly point for preinitiation complexes.
DNA unwinds at the initiator (Inr) sequence.
Transcription start site is near the Inr sequence.
Additional regulatory sequences exist upstream and downstream of the TATA box.
Promoter sequences are more complex and variable compared to prokaryotic promoters.
Eukaryotic mRNA Trancription Involves Many Proteins
Eukaryotic mRNA transcription relies on protein-protein contacts
RNA Polymerase II:
Large complex of 12 subunits
Has a carboxy-terminal domain with highly conserved repeats
Initiation, Elongation, and Termination of Eukaryotic Transcription:
Initiation:
RNA Polymerase assembles at the promoter.
Initiated by TATA box-Binding Protein (TBP) → binds to the promoter.
TBP is part of TFIID (along with TFIIB, TFIIA, TFIIF, TFIIE, and TFIIH).
TFIIH unwinds DNA (helicase activity).
TFIIH phosphorylates RNA Pol II at CTD, enabling transcription.
Elongation: Elongation factors enhance processivity and regulate post-translational modifications.
Termination: RNA Polymerase II is dephosphorylated to terminate transcription.
Regulation: Complex interactions of transcription machinery regulate transcription.
Transcription At RNA Polymerase II Promoters
Stepwise assembly of transcription factors:
TBP binds to the promoter, along with TFIIA and TFIIB.
TFIIF and RNA Polymerase II join.
TFIIE and TFIIH complete the closed complex.
DNA unwinding at the initiator (Inr) sequence.
TFIIH phosphorylates RNA Pol II, leading to promoter escape.
Elongation factors stabilize RNA polymerase.
Termination involves dephosphorylation and RNA Pol II recycling.
Function of TFIIH In Repair:
Transcription-coupled DNA repair: Actively transcribed genes are preferentially repaired.
TFIIH is involved in nucleotide-excision repair (NER):
Recruits the NER complex at damaged sites.
TFIIH defects cause genetic repair diseases, such as Xeroderma Pigmentosum.
Inhibitors of RNA Polymerases
Certain compounds can selectively inhibit prokaryotic and eukaryotic RNA polymerases, affecting transcription processes.
Actinomycin D & Acridine:
Intercalate into DNA, preventing transcription in prokaryotes and eukaryotes.
Rifampicin:
Binds to the β-subunit of bacterial RNA polymerases (affects only prokaryotes).
α-Amanitin (from Amanita phalloides):
Blocks Pol II & Pol III in eukaryotes (toxic to predators of the mushroom).
Actinoycin D & Acridine, inhibit DNA Transcription By Intercalating in DNA, then Bending It
Intercalation between successive G≡C base pairs in DNA.
Binds to the minor groove, distorting DNA structure.
Inhibits transcription by bending DNA.
The DNA is bent as a result of the actinomycin binding
