Lecture 7 - PCR
Polymerase chain reaction (PCR)
Rapid, repetitive copying of a section of double-stranded DNA located between two regions of a known sequence
Allows for rapid replication of a single copy of DNA
Requires two specific primers that bind to both DNA strands in opposing directions
Method
Heat denaturation at 95 degrees
Annealing at 40-65 degrees → primers bind to complementary sequences
Extension at 72 degrees → DNA polymerase synthesise new strands starting from annealed primers
Repeat steps 1,2,3 for multiple cycles
Double stranded PCR products covering the region we want to amplify are formed exponentially whilst products containing one strand DNA of an undetermined length are formed arithmetically
After 25-35 cycles, a discrete DNA product is formed that can be visualised by Agarose Gel Electrophoresis
Temperature changes in seconds
Heat transfer is very efficient
Heated lid prevents evaporation of sample during high temperature steps
PCR Contains:
Template
DNA polymerase
Primers
Reaction Buffer
dNTPs (ATP, CTP, GTP, TTP)
Template
Wide range of DNA molecules → genomic DNA, plasmids, fragments, phages, previously amplified fragments
Template concentration → sensitive enough to amplify 10 copies of target sequence (from 10-6 μg of DNA)
Depends on template complexity:
Genomic DNA → Use high template concentration
Plasmid DNA → Use low template concentration
Too much template yields non-specific products whilst low template results in low yield
Quality of template → avoid contamination with DNA polymerase inhibitors like proteases, chelating agents, detergents and organic solvents
DNA polymerase
Thermostable → does not denature at high temperatures used during denaturing steps and isolated from thermophilic microorganisms eg. Thermus aquaticus, Pyrococcus furiosus
Proofreading activity:
3’ → 5’ exonuclease activity allows for removal of mismatched nucleotides reducing frequency of erros
Taq polymerase → lacks 3’ → 5’ proofreading, used in routine detection PCR
Pfu polymerase → has 3’ → 5’ proofreading activity, useful when PCR product will be used in cloning or gene expression experiments
Types of ends generated after PCR:
Polymerases without proofreading activity add an extra base to the ends of the PCR product, typically an A
Polymerases with proofreading activity generate blunt-ended PCR products that can be ligated to any blunt ended site on a vector
Primers
Single stranded DNA oligonucleotides that are complementary to regions flanking the sequence of interest
15-30 bases in length, should contain 40-60% GC content

Sense Primer sequence is the same as the Sense strand and is complementary to Anti-Sense strand
Anti-sense primer sequence is the same as Anti-sense strand and is complementary to the Sense strand
Additional Considerations in Primer Design
Avoid sequences that form hairpin loops
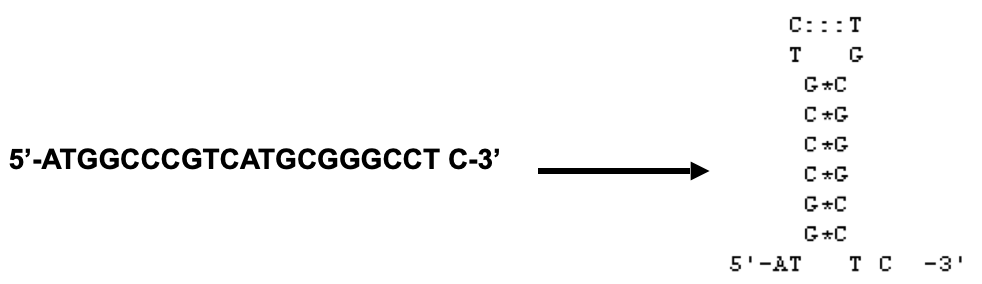
Avoid sequences that bind to each other (primer dimers) or repetitive sequences
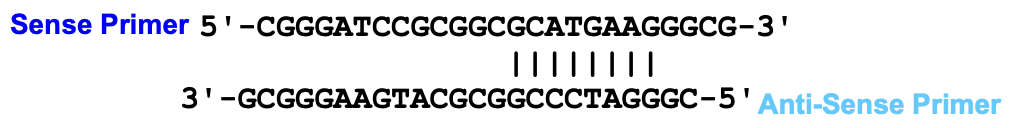
Primer requires a perfect match at the 3’ end so avoid mismatches at the 3’ end of the primer
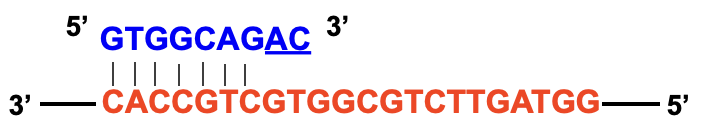
Annealing Temperature
Temperature at which the primer binds in a stable manner to the template
Usually is 2-5 degrees below melting temperature of the primer-template hybrid but it must be determined experimentally
Tm = [(A+T) x 2 + (G+C) x4]
The Tm of both primers should be similar
If primers have different Tm then determine annealing temperature from lower Tm
When annealing temperature is too low, the primer may bind to non-specific templates and the amplification of wrong products can occur
This can also occur when primer concentration is too high
Primer modifications
Primers are incorporated into the PCR product sequence so specific sequences can be added to the PCR product by modifying the 5’ end of the primer
You can add a restriction site to primer so that all new PCR products have restriction sites so they are easy to clone
Cloning PCR products
DNA polymerase lacking proofreading activity (Taq) adds an A overhang to the 3’ end of PCR products
There are commercially available vectors with a 3’ T overhang
Blue-White selection for insert
Convenient restriction sites flanking insert to facilitate further sub-cloning steps
PCR reaction conditions
Standard Components in a PCR Buffer
20mM Tris/HCl → pH 3.8
10mM KCl
2mM MgCl2
K+ and Mg2+ bind to the phosphate backbone (negative charge) and contribute to stabilising the primer annealing to the template
Mg2+ is also needed for DNA polymerase activity and fidelity of replication
If Mg+ concentration is too high non-specific annealing and DNA amplification may occur, leading to amplification of non-specific products
PCR applications
Preparing probes for Hybridisation experiments
Assays for the presence of Infectious agents
Direct Cloning from genomic DNA or cDNA
Quantification of rare or low abundance DNA and RNA
In vitro mutagenesis
DNA cycle sequencing
Genetic Fingerprinting of Forensic samples
Analysis of allelic sequence variations
Prenatal Diagnosis of genetic diseases
PCR and Genetic Disease
PCR allows for rapid genotyping of genetic markers
Used to detect deletion/insertion or point mutations
Microsatellites
Microsatellites → short tandem repeated DNA segments, most are bordered by unique DNA sequences
Huntington Disease:
Caused by dominant mutant allele of HD gene, encoding the protein Huntingtin
The mutation is a CAG trinucleotide repeat in exon 1
Healthy allele contains 10-35 CAG repeats and mutant allele has 36-121 repeats
Variations in the number of tandem repeats results in different sized PCR products that can be detected by gel electrophoresis
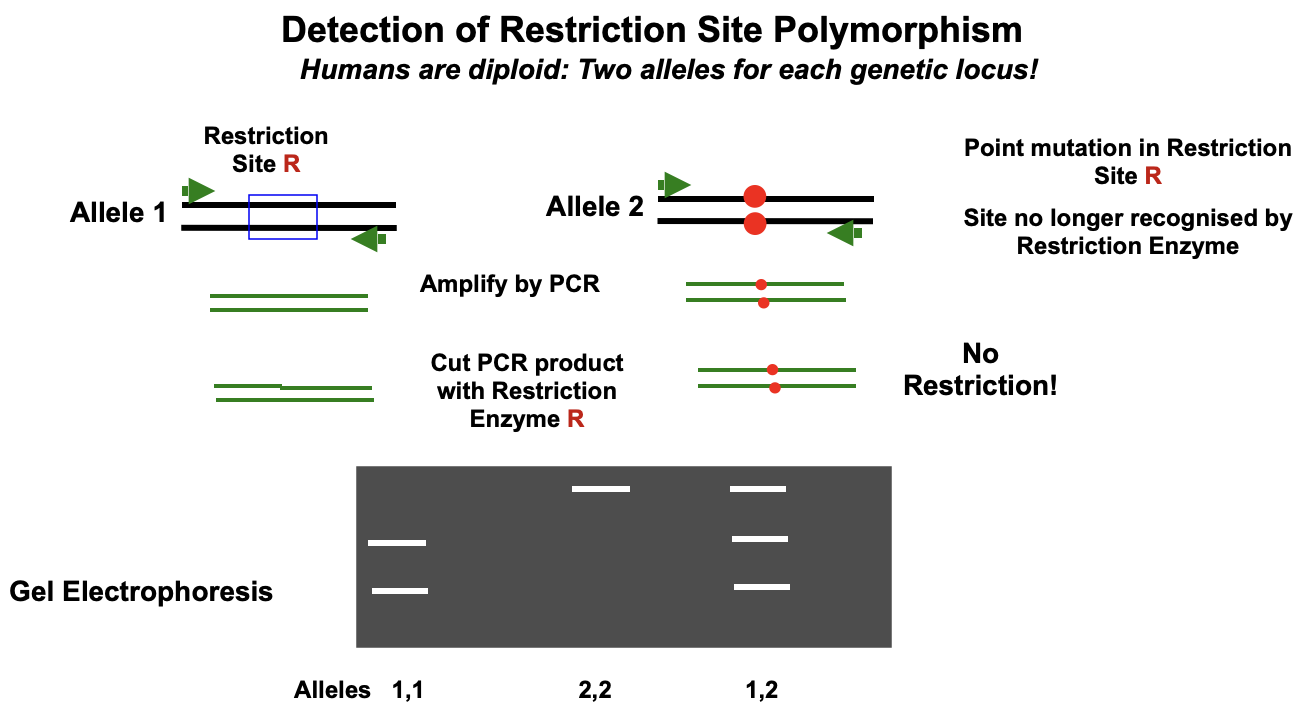
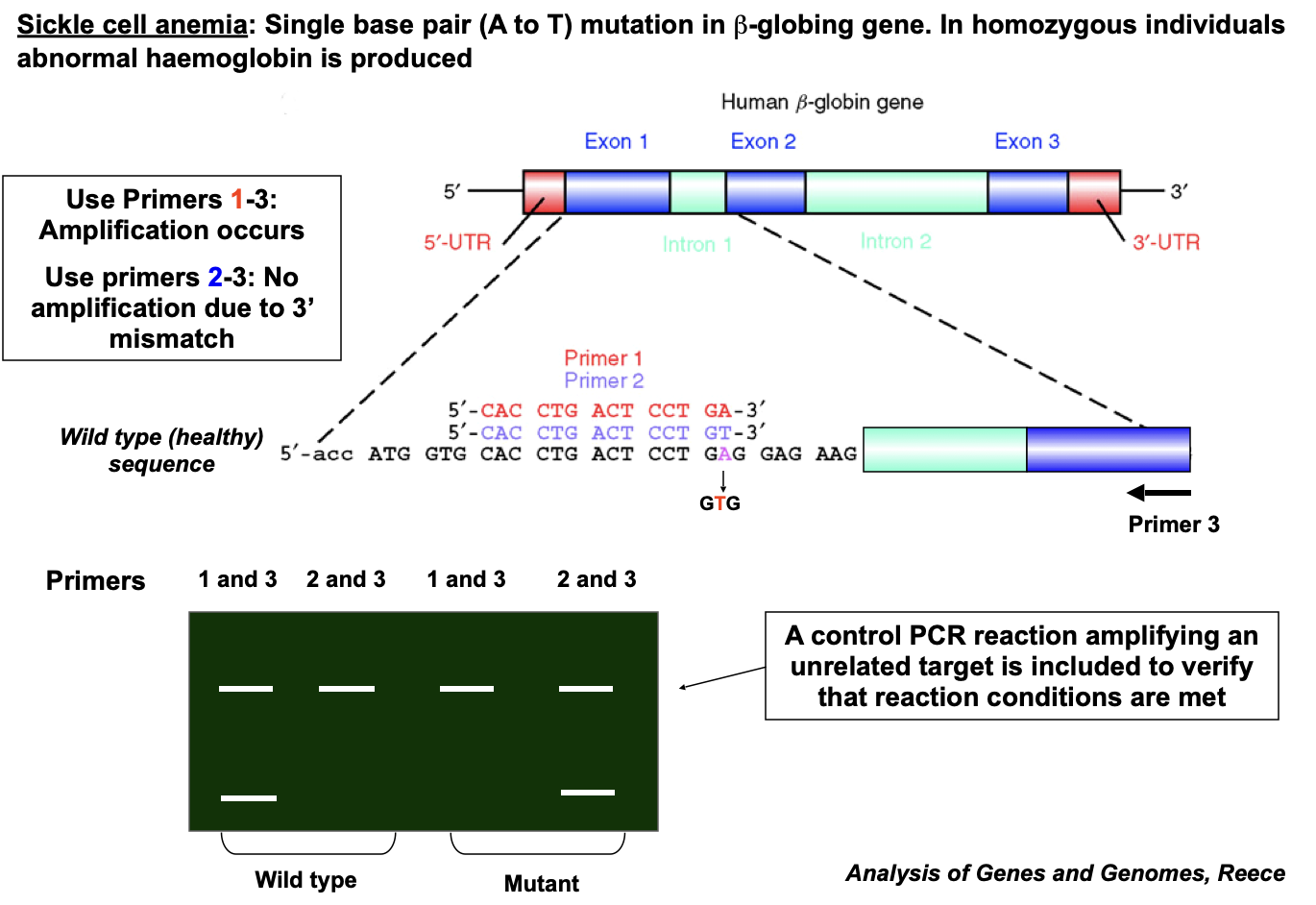
Sickle cell anaemia
Single base pair (A to T) mutation in β-globing gene
In homozygous individuals abnormal haemoglobin is produced
The mutation causing sickle cell anaemia occurs in exon 1
The normal codon is GAG (glutamic acid), while the mutant codon is GTG (valine)
Primer 1 (red) and Primer 2 (blue) are designed to flank the region containing the mutation
PCR amplifies this short region of DNA from the patient’s genome.
Both the normal and mutant alleles can be amplified
After amplification, the sequence is tested for the A→T mutation.
In this example, allele-specific PCR is used:
One primer matches the normal allele (GAG)
Another matches the mutant allele (GTG)
Molecular Epidemiology of Tuberculosis
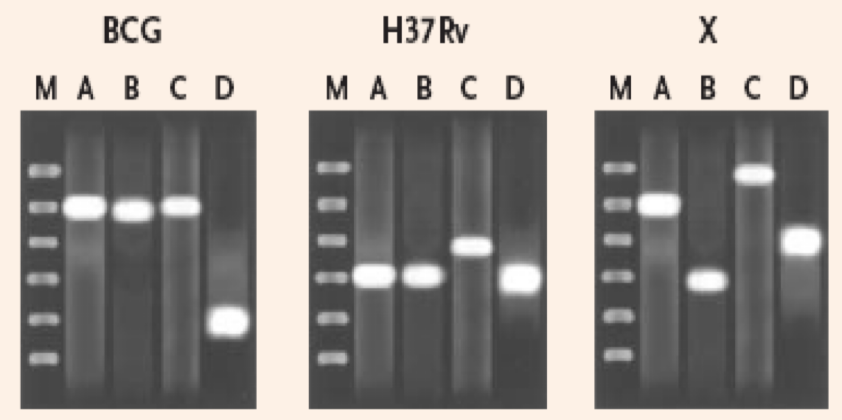
Mycobacterium tuberculosis genome contains several variable-number tandem repeats units called Mycobacterial Interspersed Repetitive Units (MIRUs)
PCR amplification of MIRUs categorises the number and size of each repetitive unit
This permits genotyping comparison of strain X and type strains