Cycle 3: Energy & Membranes
Basics of thermodynamics (textbook):
kinetic energy — from moving particles
potential energy — “stored” by virtue of position or chemical structure
chemical energy — from chemical reactions
Systems
opens systems — exchange matter & energy (e.g. cells)
closed systems — exchange energy only
isolated — neither
Law 1: energy cannot be created nor destroyed — it can only be transferred from one form to another
Law 2: entropy (“disorder”) of the universe ALWAYS INCREASES
cells fight this law — use energy to build ordered molecules
cells are “islands” of low entropy — energy taken in to replace other things broken down
Levels of Protein Structure
primary (denatured): unfolded, 1 linear chain of amino acids
peptide bonds join amino acids
secondary: 1 chain of amino acids in alpha-helix or beta-barrel form
hydrogen bonds (disturbed by urea!)
tertiary (functional): 1 fully folded 3D chain
4 major interactions between R groups: ionic bonds, H-bonds, hydrophobic interactions, dipole-dipole, disulfide bridges
quaternary (functional): multiple fully folded 3D chains
any other bond — dipole-dipole, disulfide bridges
Lecture 1: Energy & Enzymes
biological molecules are chemically reduced
sugars, fats, amino acids, nucleotides
not complete loss or gain of electrons
methane is highly reduced — lots of free energy
reduction means more hydrogens
the most energetic form of carbon
electrons are not tightly bound to any one atom
no usable energy in carbon dioxide — highly oxidized
oxidization means more C-O bonds (fewer hydrogens)
not as high in Gibb’s free energy
electrons are more tightly bound to O
limits the possibility for chemical reactions
biological molecules are between methane and carbon dioxide
fats are primarily C-H bonds — lots of energy
atom of carbon or hydrogen neither has strong attraction to electrons
neither is holding on to electron very tightly
oxygen has high affinity for electrons
oxygen is highly electronegative
electron is around
oxygen won’t allow you to take electrons from co2
co2 is oxidized
how many c-h bonds vs c-o
more c-h, more free energy (more reduced)
more c-o, less free energy (more oxidized)
Energy
autotroph: carbon starts off oxidized
carbon is brought in as CO2
can’t use CO2 — has no usable energy
uses light energy and drives arrow back towards red (making it more reduced) via photosynthesis
now have carbon molecules with more free energy
starts with something oxidized, uses light energy the make it more reduced
heterotrophs: carbon starts off reduced
takes in a molecules that is both energy and matter
cells are islands of low entropy
cells are thermodynamically open
matter and energy come into the cell
need constant supply, otherwise you die
for an autotroph, matter and energy are distinct arrows
no energy in CO2
energy as light
heterotroph: one arrow — fused together
fat contains carbon and energy (in C-H bond)
constant supply of energy to live
red balls = pool of amino acids/carbon molecules
takes energy to arrange and build those into macromolecules
requires energy to build
energy required for work
everything in our cells break down
entropy: everything breaks down—things tend towards disorder
energy spreads
energy in red balls is spread our over four balls — work comes in to produce one thing (energy is concentrated in that one thing)
entropy is higher in the spread out balls
RNA polymerase may hang around for 10-20 hours before it breaks down
constantly have to transcribe it
if you don’t, cell dies
eat to fight entropy
constantly be bringing food in
constantly having to rebuild the RNA polymerase that breaks down
spews out waste & heat
temperature will quickly rise; everyone spews out heat
free energy (G): energy available to do work
reactions with a negative change in free energy (-delta G) are spontaneous (exergonic)
spontaneity does not dictate the speed of reactions
glucose + 6 oxygen —> 6CO2 + 6H2O
negative free energy system
produce has less energy than reactants
more thermal energy in sugar
more C-H bonds in sugar than there is in CO2
energy is compacted on the left
energy is more spread out on the right
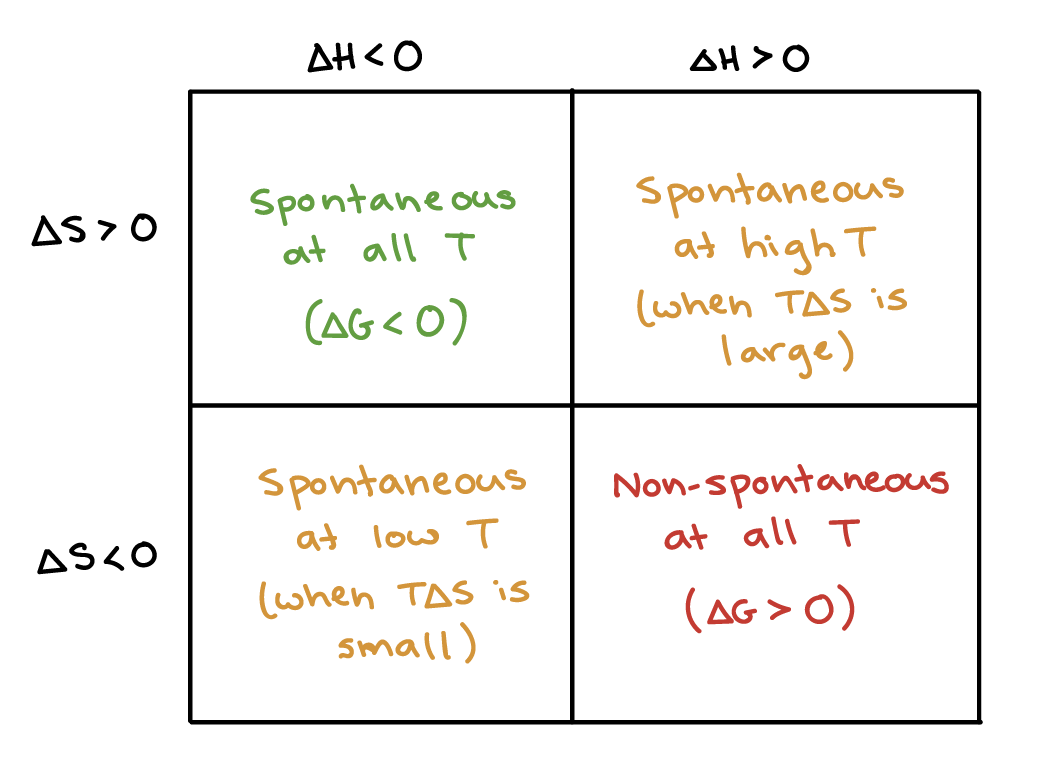
 exergonic reactions can be both exothermic or endothermic
exergonic reactions can be both exothermic or endothermic
e.g. dissolving salts with a positive enthalpy of mixing
cools down (exothermic)
spontaneous; solution has higher entropy
endergonic reactions are endothermic only
metabolism of glucose is exothermic
enthalpy change and entropy change can both change
spontaneous if exothermic & less spread out?
enzymes: primary biological catalysts
other group of biological catalysts: ribozymes
spontaneous reaction has negative delta G (exergonic)
enzymes lower activation energy to speed up the reaction
don’t affect the gibbs free energy
doesn’t tell you how fast it will occur
some are really slow: can take millions of years
movement happens when we have an enzyme
78 million years without an enzyme
20 milliseconds with an enzyme
negative delta G: products have less energy than reactants — spontaneous
reaction can just go
may not go very fast
enzyme can speed this up the rate of the exergonic reaction
positive delta G: products have more energy than reactants — nonspontaneous
enzymes don’t provide free energy
can’t give it more energy
there must be another energy source (light, ATP)
use an enzyme to bring ATP molecule close to B and transfer energy over
transfer is facilitated by an enzyme
brings 2 parties together
Why was the evolution of enzymes critical to life?
chemistry uses high pressure and temperature to speed up reaction
in biology, can’t warm things up
very susceptible to heat damage
enzymes allow us to maintain high rates of catalysis without denaturation
at low temperature
exergonic energy profile: with/without enzyme
energy of products is less than energy of reactants
transition state: bonds start to get strained
barrier is the reason why spontaneous reactions occur slowly
need to reach the activation energy (e sub a)
propane doesn’t expose when it is released because none of the propane molecules reach the activation energy
will combust with oxygen once over ten million years
bring out sparker, provide system with energy to reach transition state
propane is thermodynamically unstable
kinetically stable: doesn’t just explode spontaneously
enzyme lowers the activation energy
activation energy: energy required to get to the activation state and serves as a kinetic barrier
a low activation energy would equate to a faster reaction
rate is proportional to the number of molecules that can get to the transition state
enzymes change the kinetics, but not thermodynamics (delta G is the same)
free energy of start and finish didn’t change
enzymes only work for exergonic reactions
enzymes do not provide energy to a reaction
How do enzymes lower the activation energy of a reaction
These interactions mimic the transition state conformation of the substrate(s)
the rate of reaction is proportional to the number of molecules that can get to the transition state
Precise orientation of 2 substrates
low probability of happening without enzyme
active site of enzyme forces it into weird formation
become incredibly common with enzyme
Charge interactions
Conformational strain
can get it to break
Protein folding and Anfinsen’s dogma
proteins need to fold to be functional
protein folding is spontaneous
what do you need for proteins to fold?
100% active enzyme + urea (chemical denaturant)
urea is polar — can interfere with proper H-bonding that makes up tertiary structure
causes whole enzyme to unfold
urea outcompetes other amino acids for H-bonds
enzyme lost all activity
removed the urea, protein refolded and became 100% active
protein folding needs nothing — spontaneous process
proteins just fold
“Energy Funneling”
there are multiple ways to fold a protein
can fold normally
chaperones and energy are required to bring a transitional or misfolded protein into its active form
as an active protein, it is in its most stable form (low gibb’s free energy)
The pathway of protein folding
sucked down energy funnel
unfolded protein has higher energy than native (active) conformation
native/active conformation = lowest free energy possible
chaperone (e.g. HSPs) use energy to get proteins to unfold and refold correctly
helps them get over the hump
sometimes proteins get caught—get over hump
protein folding is spontaneous and happens at the same time
primary sequence is the only thing that dictates the final conformation
occurs in milliseconds
takes into account secondary structure (intramolecular hydrogen bonding) and hydrophobic effect (non-polar amino acids are buried inside)
Enzyme structure & catalysis
active site: area on an enzyme that binds the substrate
discovered by looking at enzyme’s 3D shape, not primary structure
only functional as tertiary structure
where catalysis occurs
enzymes can bind to more substrates on release of products
different enzymes act at different speeds
catalytic cycle is temperature dependent
substrate binds to enzyme —> induced fit
enzyme can go back and pick up new substrates
some enzymes catalyze 5 molecules of substrate a second
enzymes differ in their catalytic speed
primary sequence and active site
can’t do it
there is no series of amino acids that you can recognize that make up the active site
active site is only apparent when enzyme folds
active site is heavily dependent on its 3D (tertiary structure)
How are enzymes in antarctica vs hot springs different?
Homework — Enzymes & Growth Rate
most organisms on the planet do not maintain a constant body temperature
temperature of the organism is pretty close to the temperature of the environment
streak e. coli and petri plates
slow growth at 22 degrees, fast growth at 37 degrees (optimal growth temperature)
rate at which cells divide is temperature dependent
substrate + enzyme —> substrate-enzyme complex —> enzyme + product
reaction goes faster at higher temperatures
leads to classic growth rate curve
growth rate as a function of temperature
minimum growth rate (growth rate = 0)
reaches optimum (37 degrees)
drops quickly to maximum growth rate (maximum temperature that it can grow at)
why does it have this shape?
enzyme activity = growth rate
A: slow rise
desaturases allow membranes to stay intact
B: steep decline
denaturation
extremophiles: have variant forms of enzymes that are similar to the ones present in us
organisms that have adapted to different temperatures
but adapted to different temperature ranges
due to different tertiary structures; could be stronger or weaker depending on the temperature
too cold: enzyme is too rigid
adapts to have weaker tertiary structure bonds or arrangements
too hot: enzymes too fluid (denatures)
adapts to have stronger tertiary structure bonds or arrangements
organism’s environment dictates enzyme’s optimum temperature
psychrophiles — mesophiles — thermophiles — hyperthermophiles
will have the highest growth rate in optimal temperature
how is the tertiary structure of the enzymes different among the different groups?
although heat can result in the loss of tertiary structure, the active site of many enzymes will remain functional
tertiary structures are caused by different intermolecular forces
thermophiles have stronger IMF’s (ionic bonds, disulfide bridges, etc.)
prevents denaturation at high temperatures
makes it too rigid to be functional at lower temperatures
psychrophiles have weaker IMF’s (van der Waals forces, dipole-dipole)
denaturation at lower temperatures
not too rigid at lower temperatures
hexokinase: found in psychrophiles & thermophiles (does the same thing)
enzymes are different?
LECTURE 2 - Membrane Biology
membranes connect to the nucleus
some ribosomes are attached to the endoplasmic reticulum
secretory pathway
only used by 2 classes of proteins:
proteins that function on the plasma membrane
protein functions in extracellular space (secreted)
not for proteins in other organelles
connected to the golgi through vesicles
golgi connected to the plasma membrane through vesicles
shuttling
growth factors, stomach enzymes, nutrient acquisition
stomach enzymes are in the extracellular space; excreted from stomach cells
TAP media: phosphate, sulfur, ion
transport molecules via integral membrane proteins
can’t get through the membrane
ammonium can’t get through because it’s charged
Proteins are targeted to the ER
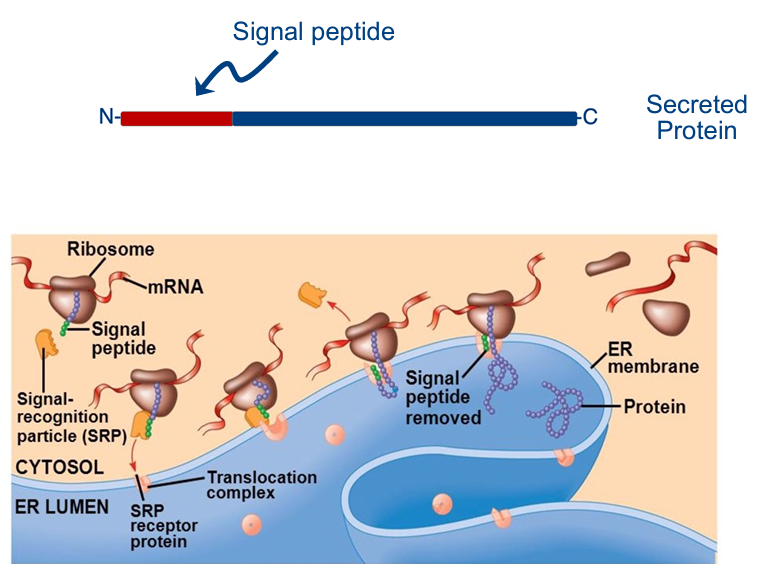 if signal recognition particle (SRP) were unrecognizable to SRP receptor sites on the rough ER
if signal recognition particle (SRP) were unrecognizable to SRP receptor sites on the rough ER
wouldn’t be able to bind to form the translocation complex
would just stop it from entering the ER and being secreted
otherwise functional
if the translated protein is an enzyme, it’s likely the enzyme will still be able to bind substrates
although the enzyme cannot be secreted as it was meant to, it will still be functional
the protein product will still be fully translated
no reason for it to be immediately degraded by a proteasome
protein cannot be translated on the rough endoplasmic reticulum
without a successful SRP-SRP preceptor, the mRNA-ribosome complex cannot enter the ER and proceed with translation there
ribosomal association with mRNA is unrelated to protein targeting
ribosomes will still be able to associate with mRNA and translate a protein
primary sequence has a signal peptide
signal peptide has nothing to do with final function of protein
part of the protein (therefore part of transcription unit, part of gene)
signal peptide: 15-30 amino acids long, sequence is similar in all species
order defines the signal peptide
if give you the primary sequence of protein: can run primary sequence through computer program — can determine (with primary sequence) whether it’s secreted
signal peptide allows the protein to enter the rough ER (via SRP-SRP receptor site interactions) and be secreted
allows it to enter the golgi & leave the cell
every ribosome is free
no ribosome is permanently stuck to the ER
translation gets arrested
signal sequence (highly similar sequence) is recognized by the SRP
SRP: signal-recognition particle that pulls the ribosome to the ER membrane
docks: translocation complex
if mutation to the translocation complex….
if mutation to signal peptide….
SRP couldn’t recognize…
mutation to enzyme that could cut the signal from the protein
once attaches, translation starts up again & finishes
interaction between signal peptide & SRP — brings whole complex to ER
enzyme in lumen of ER that cuts the functional part of the protein from the signal sequence
CAN PROTEINS BE FUNCTIONAL IF IT STILL HAS THE SIGNAL PROTEIN
IF THERE’S A NONFUNCTIONAL SRP/SRP RECEPTOR SITE, PROTEIN CAN’T GET TO ENZYME — CAN’T BE SNIPPED
leaves normal protein — can fold, go to plasma membrane
no other signal sequence
ribosome detaches to become a free, cytosolic ribosome
Membrane Transport
integral membrane proteins: membrane proteins that span the entire lipid bilayer
generally fixed within the lipid bilayer
peripheral membrane proteins: only penetrate the peripheral regions of the lipid bilayer
or peripheral membrane proteins can attach to integral membrane proteins
have some mobility — helps them carry out their biological functions (signaling)
passive transport: follow their concentration gradients, spontaneous, increase in entropy
if there were no membrane, particles would move in this way
the membrane prevents movement of the particle
the passive transporter acts as a hole in the membrane for the particle to move through
active: move against concentration gradients
you add ATP to provide energy to make up for the fact that, without ATP, this reaction would have been endergonic
if there was no membrane, the particle would move in a manner opposite to this
non-spontaneous, decreasing entropy
reaction would have been endergonic
more of one species on one side of membrane than the other
one way to cross = diffusion (high concentration to low concentration)
Simple transport: some things can pass through as if not even there (02, CO2)
no oxygen transporter
no mutation to stop oxygen uptake
if there is a defect in the oxygen transporter… — doesn’t exist
Facilitated transport: for big molecules, polar molecules
e.g. glucose (big & polar)
protein shields what’s being transported from the hydrophobic core
interacting with amino acids that interact with protein core, not hydrophobic interior
active transport — decrease entropy
want to pump even more molecules to high concentration
takes energy
going against the 2nd law
e.g. ATP-binding cassette (ABC) transporter
have hundreds on different membranes
they differ in the transmembrane domain
transmembrane domain: actually has to interact with hydrophobic core — not easy!
confers specificity with regards to what is transported
fold in different ways
pumping ammonium vs. phosphate
ATP binding domain: engine, where ATP binds (where they get the energy to pump solutes into a space of high concentration)
all the same in ABC transporters
driving force of diffusion = entropy
energy becomes spread out
How can a pore/channel be specific for one thing?
e.g. aquaporins
diffusion of water and NOTHING ELSE
individual molecules move from one side to the other
when protein folds, the core is incredibly tight
channel is not a giant hole — very narrow path
very delicate, intimate charge interactions between amino acids that line the pore and the molecule
specificity comes from the SHAPE and CHARGE INTERACTIONS
does protein go through secretory pathway
give primary sequence
amino acids that interact with interior are hydrophobic
Hydropathy plot: membrane protein prediction
x-axis: amino acid number
y-axis: relative hydrophobicity index (hydrophobic is high, hydrophilic is low)
hydrophobic peaks correspond to hydrophobic domain
you can predict if a protein is an integral membrane protein: look at the primary sequence
amino acids that interact with the membrane tend to be hydrophobic
protein sequence is dominated by hydrophobic amino acids
can get a computer program: creates hydropathy index
7 peaks: 7 transmembrane domains in rhodopsin
3 peaks: 3 domains that interact
looking for stretches of primarily hydrophobic
hexokinase: may just have a couple hydrophobic mixed with everything else
interaction with the membrane gives tell-tale signal
it takes about 20 amino acids to traverse a membrane — transmembrane proteins generally have seven domains
Cystic Fibrosis
normal lung physiology — delicate balance of ions and water
chloride pump (CFTR) is an ABC (ATP-binding cassette) transporter
maintains the correct concentration gradients of chloride within the lungs
concentration gradient ensures correct osmosis of water
water needed to keep cilia wet and mobile to prevent lung infections
Chaperones (e.g. HSP90) prevents misfolding CFTR from reaching the membrane
useful in healthy person
causes problems in cystic fibrosis
infections, thick mucus, trouble breathing
heat shock protein 90 recognizes that CFTR is misfolded, due to mutations, and prevents it from reaching the membrane
with no CFTR at the membrane, cells within the lung do not pump Cl- out into the airway environment — water does not follow as well
proteasome = trash bin
causes the airways to become very dry
infections are common: cilia don’t sweep out pathogens stuck in mucus
CFTR: cystic fibrosis transmembrane conductance regulator
caused by mutation to CFTR (an ABC-transporter)
most common inherited disease in Canada among people of European descent
die young, couldn’t breathe
6000 bases, 1480 amino acids — gigantic gene
most common deltaF508 (70% of cases)
ABC transporter doesn’t work
doesn’t have a phenylalanine (F) in position 508
normal does
doesn’t fold like a WT (even though just one change)
slight change in protein conformation — cystic fibrosis
Anfinsen’s Dogma: could show urea and such
don’t need anything for protein folding
what does determining the shape of a protein is the primary sequence
what amino acids you have in what order
unfolded sequence determines it
energetically downhill
genes don’t code fatty acids
epithelial lining — plasma membrane of epithelial cells
CFTR is sitting on plasma membrane of those cells
CFTR is a chloride pump — active transport (bringing it to high concentration)
lining of lung/intestine has lots of chloride
inside of lung has to stay wet — chloride allows for osmotic movement of water from aquaporin
keeps lining wet
cilia + mucus = important
lining of lung is moist to allow for gas exchange
when smoke/bacteria gets into lungs — cough it up
lung clearance — so important
if have cystic fibrosis, doesn’t work — don’t pump chloride, don’t get osmotic movement of water, dried up, cilia can’t move
need lung transplant
because of defect to single protein-coding gene
diffusion of gases is inhibitory
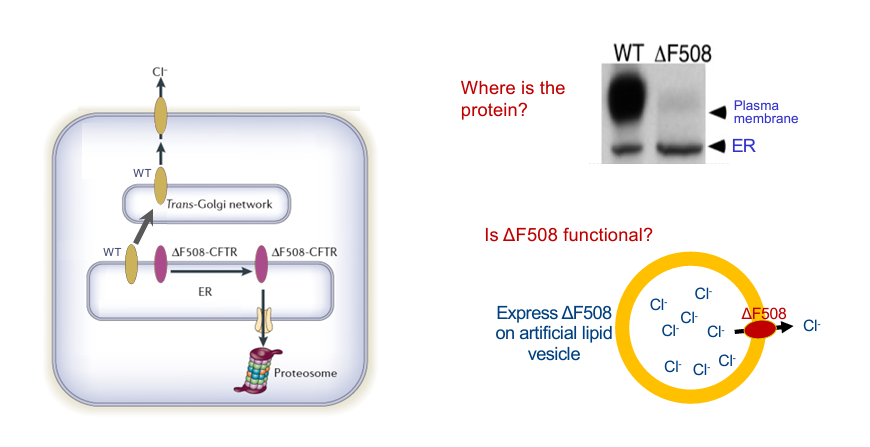
Cellular fate of the F508 form?
don’t worry about word trans
ribosomes attaching to ER
WT goes to epithelium membrane and does it s job
delta F508 never gets to go do its job
get’s tagged
goes through trapdoor
goes to cellular recycling bin: proteosome
delta F508 NEVER LEAVES THE ER
Western blotting (nothing like RNA blot analysis)
looking for expression of single protein
hangs out on plasma membrane, some on ER
isolate protein from CF patient: maybe a little smudge of plasma membrane, most in ER
not very much; degraded very quickly
link protein analysis figure to model on left
Is delta F508 functional?
experiment: artificial lipid membrane
encloses solution
make lipid in solution of chloride
express delta F508
get rid of chloride in extramembrane solution
test if chloride goes out into extramembrane space
can you detect chloride in external environment — YES
add a little ATP
20% as good as WT
one amino acid changes conformation (not as good as WT)
but still partially functional
people have cystic fibrosis because F508 never gets the chance to get to the plasma membrane
don’t need a lot of chloride
if had mutant form in plasma membrane, would be fine (wouldn’t have cystic fibrosis)
ER contains a quality control system
ERQC = quality control system
chaperones (e.g. heat shock protein 90) detect misfolding
if good — go to right
if bad — tagged & degraded (go to left)
what would drugs aim to do?
Lipid saturations and implications
Organisms can adjust membrane fluidity
increased saturation — decreased fluidity (high temp)
decreased saturation — increased fluidity (low temp)
need to maintain proper fluidity regardless of temperature
because electron transport, proteins — need to change shape
cells would die if membrane too rigid
if membrane too fluid, leaky (ions could indiscriminately leak)
living things has a way of adjusting that
stay like Italian olive oil
low temperature: intentionally introduce double bonds (when membrane synthesized via enzymes)
kinks makes the membrane more fluid than if kinks weren’t there
unsaturation
high temperature: don’t want kinks (decreased fluidity)
have saturated fatty acids: no double bonds (linear)
desaturases: enzymes that can adjust membrane fluidity by increasing the degree of unsaturation in membrane fatty acids
can introduce kinks — introduce double bonds
organisms can alter their membrane fluidity at different temperatures by changing the level of desaturase expression
22 degrees is cold for bacteria
high levels of desaturase transcript abundance
high temp: desaturase expression goes away
desB, desD, desA are desaturase genes
how fast can switch on desaturase expression to keep membrane fluid
if adapted to low temperature, lots of desaturase
no enzyme that reverses this
can’t unking fatty acid
can’t do the reverse
rely on the fact the fatty acid biosynthesis always makes them in a saturated form
membrane degrades, replaces
phospholipids: primary constituent of the cell membrane
hydrophilic (glycerol) and hydrophobic (fatty acids)
amphipathic: both h-philic & h-phobic
plasma membrane is a bilayer (2 phospholipids)
has a hydrophilic portion of the outer/inner edges
inside is all hydrophobic
integral membranes are proteins embedded fully in the membrane
nonpolar molecules (any size) can get in
small, polar molecules can get in
Secretory Pathway
how proteins get on the plasma membrane or secreted out of cell
ER —> Golgi —> plasma membrane via vesicles
proteins are synthesized by the RER
vesicles containing synthesized proteins bud off from the ER
reach the golgi, are packaged and modified within the Golgi, then vesicles bud off
vesicles will then travel through the cytoplasm until they reach the cell membrane
will end up either being excreted or a membrane protein
Protein targeting
translation occurs in the cytoplasm
sending protein to membrane
place a signal peptide tag on the protein
indicates final location
cell is able to recognize this tag
cell transports proteins to proper places
when the protein gets translated, the signal peptide is translated as part of the protein
once the protein is translated, the signal peptide is removed (the mature protein no longer has a signal peptide)
basically a set of instructions telling the cell where to send the protein
lots of the processes exist (cytoplasm — nucleus, cytoplasm — mitochondria)
The Secretory Pathway
4-Step process:
RNA translated by ribosome, and a signal sequence pops out as polypeptide
Translation stops: signal receptor protein (SRP) binds to signal sequence
SRP binds to a SRP receptor protein on the ER
translation continues into the ER
SRP-ribosome-polypeptide hybrid moves to the ER
SRP binds to the SRP receptor on the ER membrane (not nuclear or plasma membrane)
translation of the entire protein will continue into the ER
Following translation of protein to be sent to the membrane, the signal sequence is cut
after the entire mRNA is translated, the ribosome assembly leaves, and the signal sequence is cut by an enzyme
protein is now free to be sent to the golgi — and then the plasma membrane
mRNA: has signal sequence
more of a set of instructions for the signal sequence to be encoded
initial protein: signal sequence appears as a polypeptide
this is the tag recognized by the SRP
final/mature protein: signal sequence is excised
pathway to mitochondria
transcribe mRNA in nucleus — begin protein translation in cytoplasm — signal tag recognized — ribosome-mRNA complex brought to that area