DNA Manipulation
- Restriction enzymes are enzymes that cut DNA at specific sequences, called restriction sites. They are used to cut DNA into fragments that can be analyzed, manipulated, or inserted into other DNA molecules.
- Sticky cuts leave overhangs that are complementary to another strand, while blunt cuts produce fragments with a straight end.
- Ligase is like glue that seals the gaps that polymerase makes by making new bonds.
- Prokaryotic DNA is circular therefore one cut produces one fragment. # cuts = # fragments
- Eukaryotic DNA is linear therefore one cut produces two fragments (think about cutting a straight line). # cuts = # fragments + 1
\
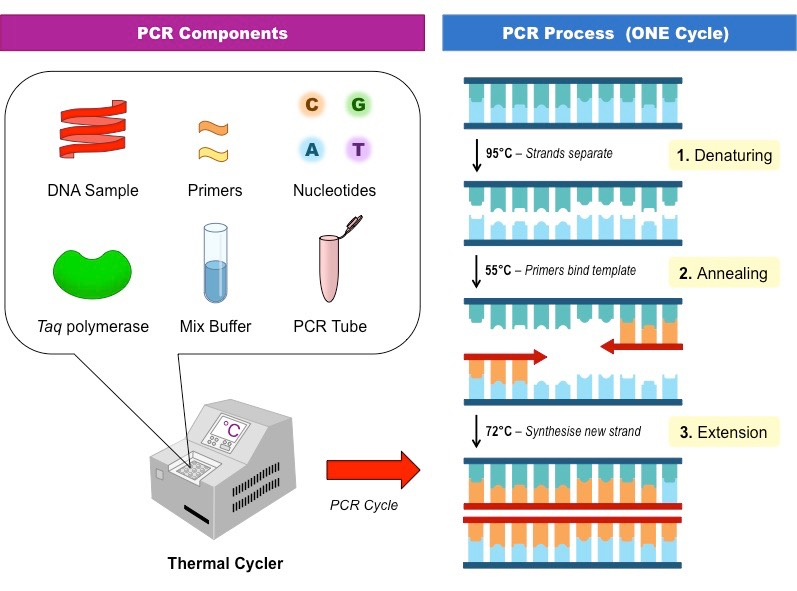
PCR
- PCR (polymerase chain reaction) is a technique used to amplify a specific segment of DNA, making millions of copies in a relatively short amount of time. * The stages of PCR are: denaturation (at 94-98°C), annealing (at 50-65°C), and extension (at 72°C). In the denaturation stage, the DNA is heated to separate the two strands. In the annealing stage, primers bind to complementary sequences on each strand of the DNA. In the extension stage, a DNA polymerase adds nucleotides to the 3' end of the primers, extending the new strands.
- Primers are short DNA sequences that bind to specific regions of the template DNA during PCR. Their function is to provide a starting point for the DNA polymerase to begin synthesizing new DNA strands.
- Polymerase is an enzyme that synthesizes new DNA strands by adding nucleotides to the 3' end of a primer.
- The equipment used to perform PCR typically includes a thermal cycler, pipettes, and PCR tubes or plates.
- The components needed to perform PCR include DNA template, primers, DNA polymerase, free nucleotides, and a buffer solution.
Gels & Electrophoresis
- Gel electrophoresis is a technique used to separate and analyze DNA fragments based on their size and charge.
- The ingredients to make a gel include agarose powder, and buffer solution (not water).
- Different % of agarose gels are used to separate DNA fragments of different sizes. Higher % agarose gels are used to separate smaller fragments, while lower % agarose gels are used to separate larger fragments.
- ==Polyacrylamide== gels are used to separate smaller DNA/RNA fragments, or proteins while ==agarose== gels are used to separate larger DNA fragments.
- The running buffer provides the ions needed to carry the electrical current during electrophoresis, as well as maintaining the pH and preventing the gel from overheating.
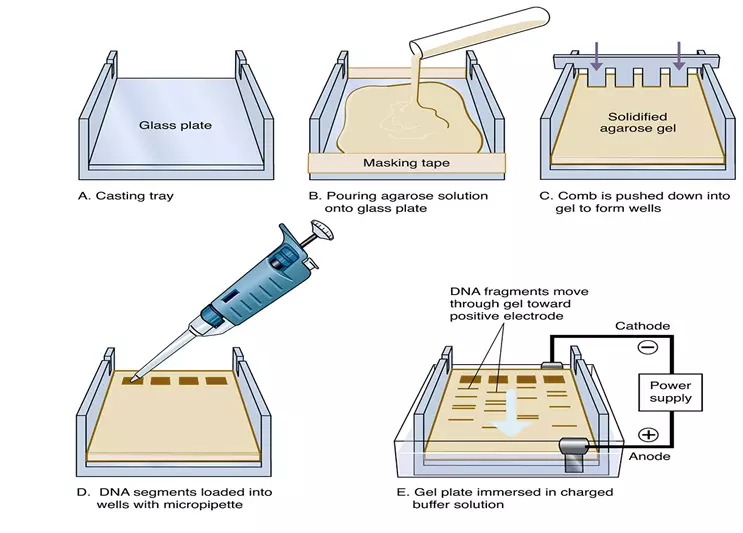
- To set up an electrophoresis chamber, the gel is placed in a casting tray with a comb at one end. The casting tray is then placed in the electrophoresis chamber with buffer solution added to cover the gel. The samples are loaded into wells at the opposite end of the gel from the comb.
- Loading dye is used to help visualize the samples during electrophoresis and to ensure that they sink into the wells. It contains a tracking dye and a dense solution that makes the sample more visible.
- DNA has a negative charge due to its phosphate groups, and it migrates towards the positive electrode during electrophoresis.
- The DNA ladder is a mixture of DNA fragments of known sizes that is used as a standard for determining the size of unknown. It is used as a ==reference!==
- We stain gels to visualize the DNA fragments. Methylene blue is a dye that binds to DNA, making the fragments more visible.
- Electrophoresis safety includes wearing gloves, avoiding contact with the buffer solution, and turning off the power supply before opening the electrophoresis chamber.
Mapping & Tools
- To read a gel, you can determine the number and size of DNA fragments based on their position in the gel relative to the DNA ladder. The cuts can be identified by comparing the fragment patterns to the known recognition sequences of the restriction enzymes used.
- A plasmid map is a diagram of the plasmid showing the location of different DNA sequences and the location of restriction sites as well as base pair sizes.
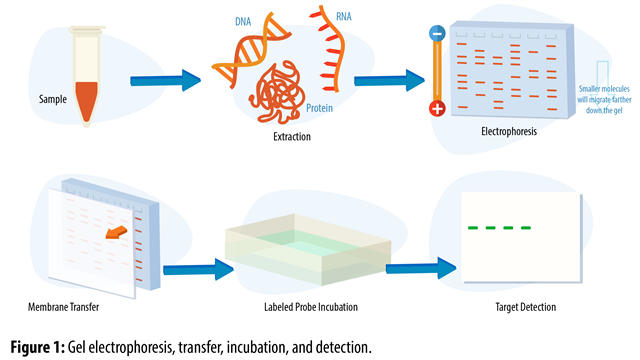
- Southern blotting is a technique used to detect specific DNA sequences in a complex mixture of DNA fragments. The fragments are separated by gel electrophoresis and ==transferred to a membrane.== The membrane is then probed with a ==labeled DNA probe== that hybridizes (attaches) to the target sequence. These are concrete results.
- Probes are single-stranded DNA or RNA molecules that are labeled with a radioactive or fluorescent marker. ==They are used to detect specific DNA sequences== (specifically in Southern blotting by hybridizing) to complementary sequences on the target DNA.
- A semilog graph is used to estimate the size of DNA fragments based on their migration distance in a gel. The graph plots the fragment size against its distance in the gel. The size of an unknown fragment can be estimated by measuring its migration distance and using the graph to estimate the corresponding size.
- Short tandem repeats (STRs) are repetitive DNA sequences that are found in specific locations in the genome. They are used in forensic analysis to identify individuals based on their unique STR profiles.