Lecture 26: Pedigree Analysis, PCR and DNA Profiling
Learning Outcomes
Analyse a pedigree to determine the pattern of inheritance, identify genotypes and predict progeny ratios.
Explain the steps involved in the polymerase chain reaction (PCR) and give examples of applications of PCR.
Explain the role of primers in determining the specificity of PCR amplification and show where they should be placed to amplify a given region of DNA.
Compare and contrast PCR in a tube with DNA replication in a cell.
Explain how DNA profiling is done and interpret DNA profiling data.
Pedigree Analysis
Definition: Analysis of a family tree to determine the inheritance pattern of a specific trait.
Used to identify genotypes and predict progeny ratios.
Helps determine if a trait is recessive or dominant, and if it's X-linked or autosomal.
Pedigree Conventions
Males are represented by squares, females by circles.
Horizontal lines connect parents, vertical lines connect parents to offspring.
Shaded symbols indicate individuals expressing the trait.
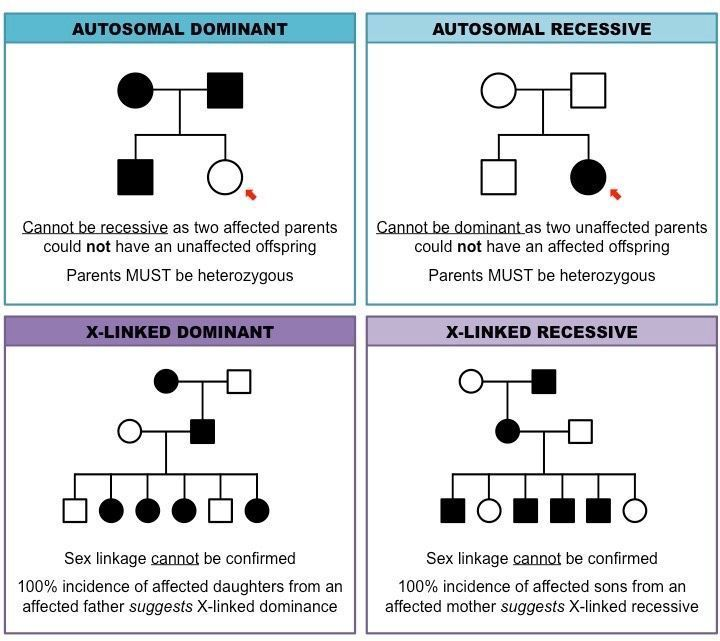
Determining Inheritance Patterns
Autosomal-Not located on a sex chromosome(X,Y)
Autosomal Recessive:
Trait often skips generations.
Affected individuals usually have unaffected parents.
Both parents must carry the recessive allele for a child to be affected.
Autosomal Dominant:
Trait appears in every generation.
Affected individuals have at least one affected parent.
X-linked Recessive:
More males are affected than females.
Trait can skip generations.
Affected males inherit the allele from their mothers.
X-linked Dominant:
Affected males pass the trait to all their daughters but not to their sons.

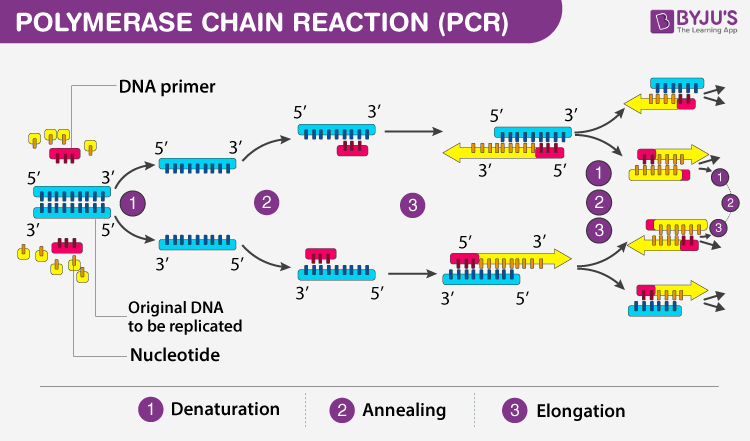
Polymerase Chain Reaction (PCR)
Definition: In vitro DNA replication technique used to amplify a specific DNA sequence.
Steps Involved:
Denaturation: Heating the DNA to 95°C to separate the double helix into single strands.
This step replaces the function of helicase in cellular DNA replication.
Annealing: Cooling the DNA to allow primers to bind to the target sequence.
Extension: Temperature is changes again(slightly increased) so that conditions are ideas for DNA polymerase to extend the primers, synthesizing new DNA strands.
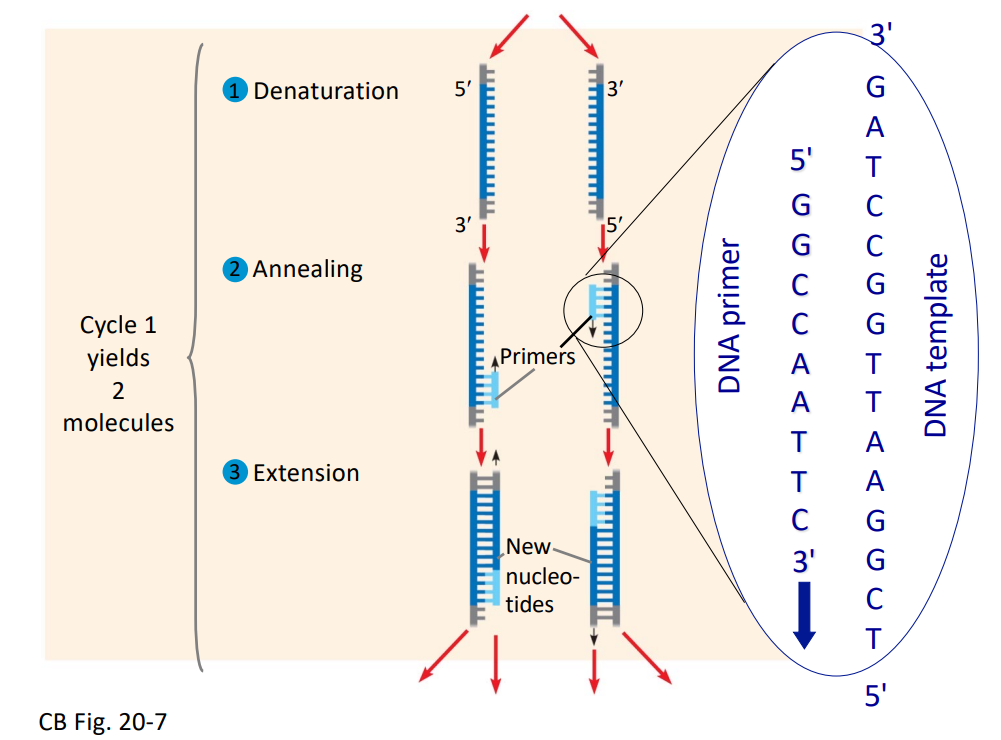
Specificity: Primers determine which DNA sequence is copied.
Primers are designed to flank the target region.
Sensitivity: Requires only a small amount of DNA template.
Cycles: Repeated cycles lead to exponential amplification of the target sequence.
Cycle 1 yields 2 molecules.
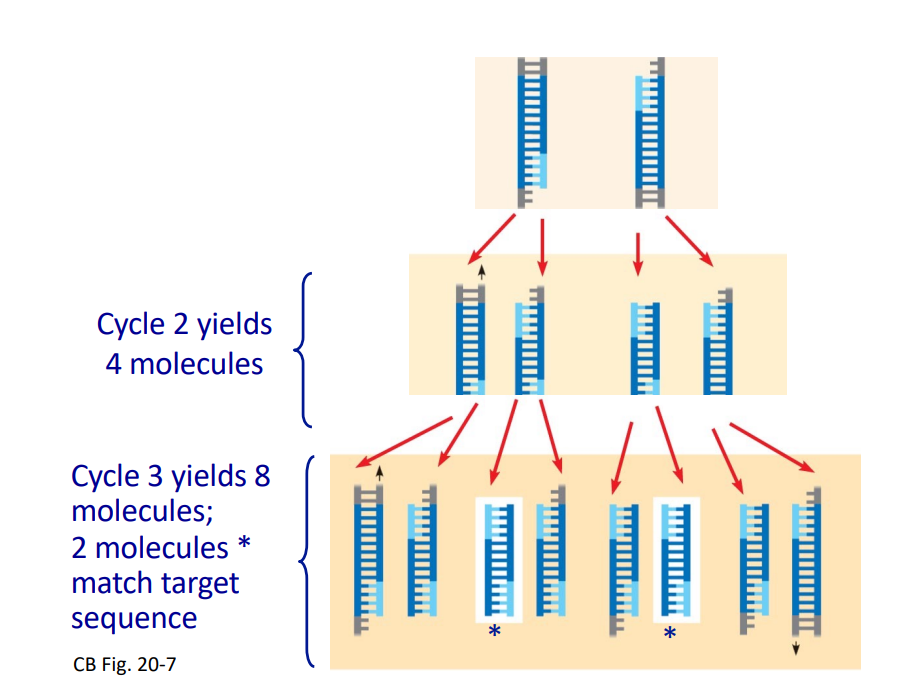
Cycle 2 yields 4 molecules.
Cycle 3 yields 8 molecules, with 2 matching the target sequence.
PCR vs. DNA Replication
PCR (In Vitro):
Occurs in a test tube.
Uses a heat-stable DNA polymerase.
Requires specific primers.
Involves repeated cycles of denaturation, annealing, and extension.
DNA Replication (In Vivo):
Occurs in a cell.
Uses various enzymes like helicase, primase, and DNA polymerase.
Initiated at multiple origins of replication.
Continuous process.
DNA Profiling
Definition: Technique used to identify individuals based on their DNA.
Inherited from parents like any other alleles.
Method: Gel electrophoresis of PCR-amplified Short Tandem Repeat (STR) markers.
Short Tandem Repeats (STRs)
Definition: Short, repetitive DNA sequences (usually 2-5 bp) that are widely used as genetic markers.
Often located in non-coding regions of the genome.
Many different alleles exist at each locus.
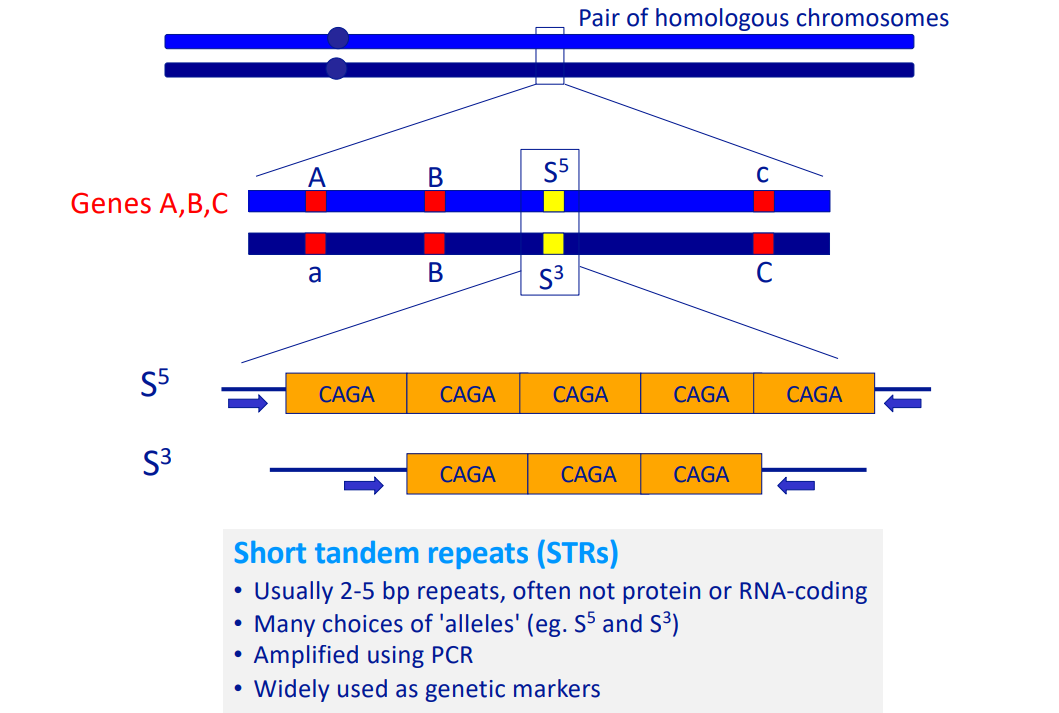
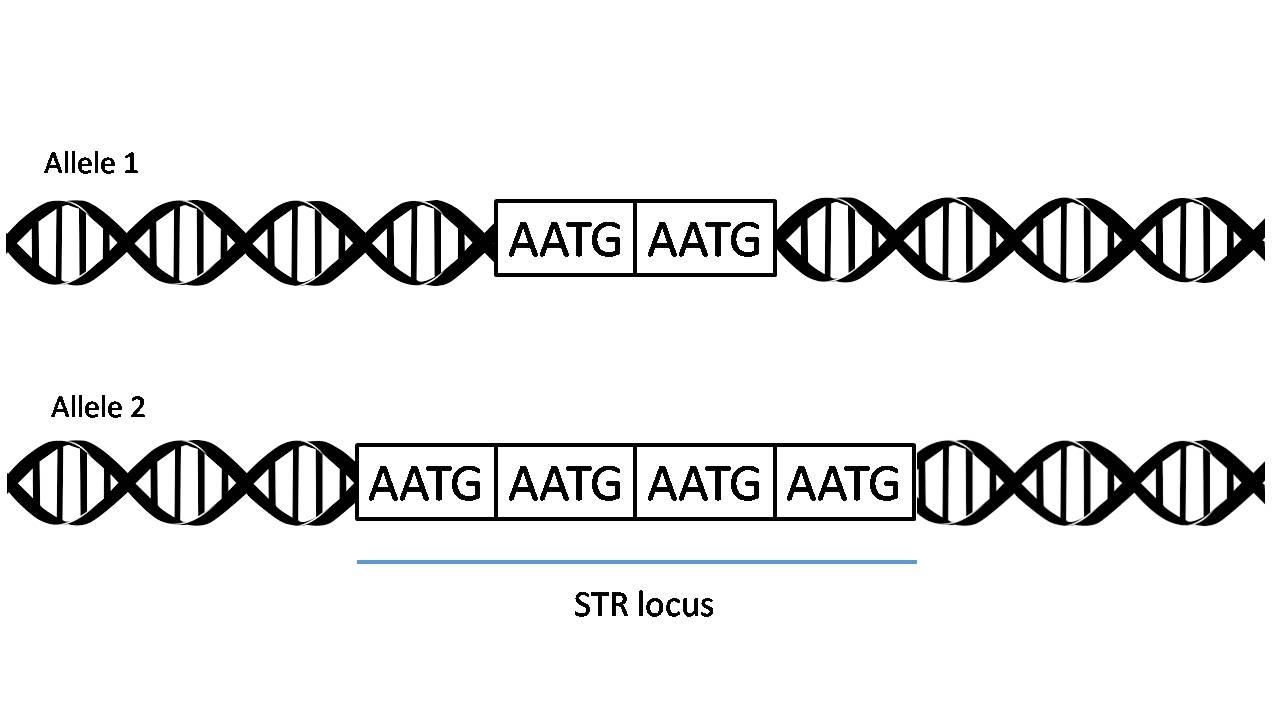
Amplification: Different Short Tandem Repeats loci can be amplified using locus-specific PCR primers to make a DNA profile
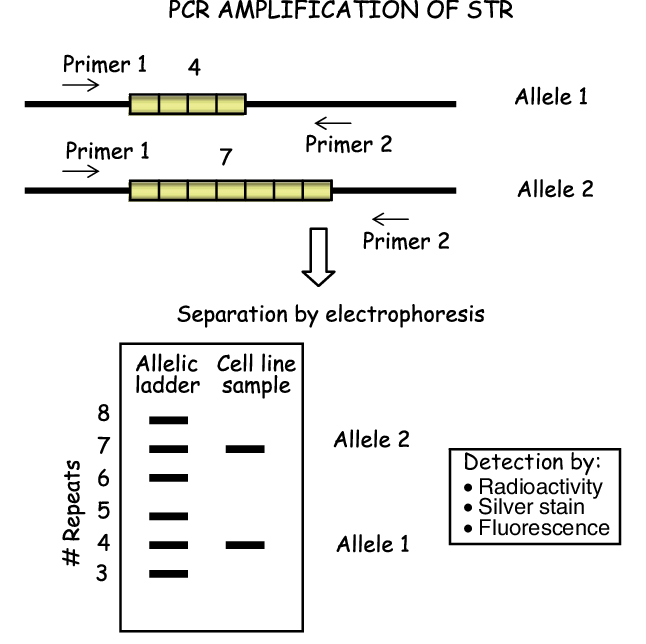
Applications:
Forensics: Based on differences in Short Tandem Repeats profiles between individuals.
Parentage Testing: Based on family similarities in Short Tandem Repeats profiles.
E.g. Determining the father of a child by comparing STR profiles of the mother, child, and potential fathers.
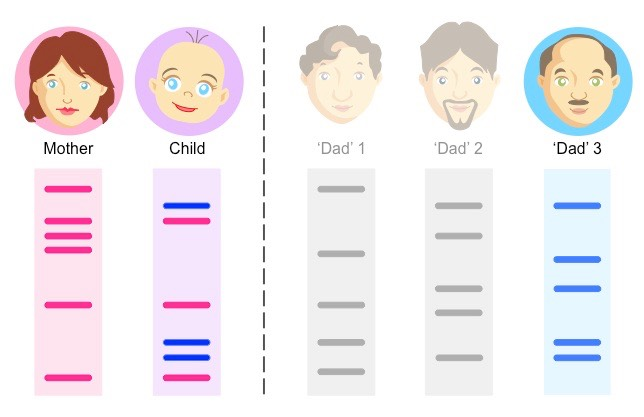
DNA Profile Interpretation
Individuals have different STR alleles at each locus.
Many different STR alleles at each locus, so unrelated people usually differ.
STR alleles are visualized using gel electrophoresis.
By analyzing multiple STR loci, a unique DNA profile can be generated for each individual.